|
Size: 36392
Comment:
|
Size: 34705
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 13: | Line 13: |
| In Brainstorm we offer two approaches for computing time-frquency decompositions: the first one is based on the convolution of the signal with series of complex Morlet wavelets, the second filters the signal in different frequency bands and extract the envelope of the filtered signals using the Hilbert transform. | In Brainstorm we offer two approaches for computing time-frquency decompositions (TF): the first one is based on the convolution of the signal with series of complex Morlet wavelets, the second filters the signal in different frequency bands and extract the envelope of the filtered signals using the Hilbert transform. |
| Line 35: | Line 35: |
| {{attachment:edgeEffect5Hz.gif}} {{attachment:edgeEffect10Hz.gif}} == Time-frequency process == We are going first to compute the time-frequency decomposition for the two averaged recordings we have in the protocol. Drag and drop the two ERF files from ''Right ''and ''Left ''in ''Process1''. Select "Recordings". {{attachment:fileSelection.gif}} Click on Run. Select the process: '''Frequency > Time-frequency (Morlet wavelets)'''. * The only generic option you have access to in this window is the selection of the sensors on which you want to apply the process. You can use this box to select a mixed list of channel types (ex: "MEG") or channel names (ex: "MRT16"). * For now, you can just leave the default "MEG, EEG". There is no EEG channel but it doesn't matter, the extra information is just ignored. {{attachment:processSelection.gif}} Click on the "Edit" button to get access to the options specific to the time-frequency decomposition process. With this time-frequency option window, you can: 1. define bands in time and frequency or analyze the data in all points, 1. configure the time and frequency resolution of the wavelets, 1. define the output type (complex coefficients or squared coefficients / power). {{attachment:timeFreqOptionsBasic.gif}} == Description of the options == |
{{attachment:edgeEffect5Hz.gif||height="244",width="299"}} {{attachment:edgeEffect10Hz.gif||height="245",width="302"}} == Simple example == Let's start with a simple example to explore the options of the time-frequency decomposition process. * In Process1, select the average recordings for the '''deviant '''condition for Run#01. * Select process: '''Frequency > Time-frequency (Morlet wavelets)'''.<<BR>><<BR>> {{attachment:example_process.gif||height="288",width="507"}} * Enter "'''MEG'''" for the sensor types, do '''not '''select the "Spectral flattening" option.<<BR>>Click on the button ['''Edit'''] to see all the process options.<<BR>><<BR>>Time definition: '''Same as input''', Frequency definition: '''Log 1:40:150''', Compute measure: '''Power'''. <<BR>><<BR>> {{attachment:example_options.gif||height="528",width="392"}} == Process options == |
| Line 60: | Line 47: |
| '''Time definition''': * This panel describes the time points for which you will get a value at the end of the computation. * '''Same as input file''': If you select this option, you would get the time-frequency decomposition (TF) for each of the time samples in the input file (here: 375 samples between -50ms to 250ms) * '''Group in time bands''': This option adds another step of computation: * It first computes the TF for each of the time points in the initial file, exactly like with the "same as input file" option * Then for each time band: it groups the corresponding time points by averaging the power of their TF values. * Definition of the time bands: two methods |
'''Time definition''' * '''Same as input file''': The output file has the same time definition as the input file. <<BR>>In this example, it means: 361 samples between -100ms and 500ms. * '''Group in time bands''': This option adds a step of computation. First it computes the TF decomposition for all the input file, then averages the power by time band. To define a time band: |
| Line 72: | Line 55: |
| '''Frequency definition''': * This panel describes the frequencies for which the power will be estimated at each time instant. * '''Linear''': You can specify the frequencies with any Matlab syntax start:step:stop. The default is "1:1:60", which produces 60 values [1, 2, 3, 4, ..., 59, 60] * '''Group in frequency bands''': As for the time definition, this option leads to a two-step process: * It computes the TF for several values in the frequency band (see below how to define a frequency band) * Then for each frequency band: it averages the power of all the frequencies in the band, so that you get only one value for each frequency band (and for each time point and each sensor). * Definition of the frequency bands: |
'''Frequency definition''': Frequencies for which the power will be estimated at each time instant * '''Linear''': You can specify the frequencies with the Matlab syntax start:step:stop. <<BR>>The default is "1:1:60", which produces 60 values [1, 2, 3, 4, ..., 59, 60]. * '''Group in frequency bands''': As for the time definition, this option leads to a two-step process. First it computes the TF decomposition for several values in the frequency band, then it averages the power of TF coefficients per frequency band. To define a frequency band: |
| Line 81: | Line 60: |
| * The name can be any string you want * The frequency definition is a Matlab expression evaluated with an eval() call. If the frequency definition contains only two values, Brainstorm adds two extra values in the middle so that the final averaged value is a bit more robust. * Examples of valid Matlab expressions: * "2,4": evaluates to [2,4], and as there are only two values, it uses the frequency vector [2, 2.66, 3.33, 4] instead * "2, 2.5, 3, 3.5, 4": evaluates to [2 2.5 3 3.5 4] * "2:4": evalues to [2 3 4] * "2:0.1:4": evaluates to a vector of 21 values [2 2.1 2.2 2.3 .... 3.8 3.9 4] |
* The frequency definition is a Matlab expression evaluated with an eval() call. If the frequency definition contains only two values, Brainstorm adds two extra values in the middle so that the final averaged value is a bit more robust. Example of valid expressions:<<BR>><<BR>>"'''2,4'''": Evaluates to [2,4], and then expands to the frequency vector [2, 2.66, 3.33, 4]<<BR>><<BR>>"'''2:0.5:4'''": evalues to [2 2.5 3 3.5 4]<<BR>>"'''2, 2.5, 3, 3.5, 4'''": Evaluates to [2 2.5 3 3.5 4] |
| Line 89: | Line 62: |
| * Click on the "Reset" button to go back to the default settings. '''Morlet wavelet options''': * Central frequency: frequency where the mother wavelet is designed. All other wavelets will be shifted and scaled versions of the mother wavelet * Time resolution (FWHM): desirable temporal resolution of the wavelet at the central frequency (in units of Full Width Half Maximum). Click 'display' to see the resolution of the wavelet in other frequencies * See the first section of this tutorial for a description of these parameters. '''Processing options''': * '''Compute the following measure''': * The TF returns the wavelet coefficients for each frequency/time/sensor, which are complex values. Typically, what you want to display is the power of the coefficients, which is the square of the amplitude: abs(TF).^2. You can choose if you want to apply this transformation or not. * '''Power''': Applies the "power" transformation right after the computation. This causes a loss of the phase information. * '''None''': Save the TF coefficients as they are computed. This can be useful if you plan to use these decompositions for other purposes that require the phase, but the files are going to be twice as big. * If you select "None" here, the power will be computed on the fly for the various displays. * Some combinations of options may disable this choice. Example: if you ask for time bands or frequency bands, the program will have to compute the power before averaging the values; so in these cases the "None" option is disabled. * '''Output''': * '''Save individual TF maps''': This option stops the computation here, and saves in the database one time-frequency file for each input file. * Be very careful using this option because if you have hundreds of trials, it will fill your hard drive very quickly. * One full TF file in this case contains [nSensors x nTime x nFreq] = 3.397.500 values = 25 Mb. * You can however limit the amount of data produced by using time and/or frequency bands. For 15 time bands and 6 frequency bands, the size of each file drops to: 110 Kb. * '''Save average TF maps''': This option is available only when there are more than one input files (number of files you dropped in the Processes tab). * Instead of saving the TF for each file separately, it automatically computes the average of the power of all the TF. * This is a good choice if you do not plan to use independently all the TF files, because it saves a lot of time and disk space. === Computation === Back to our evoked fields. Let's select the following options: * Time: Same as input * Frequency: Linear 1:1:60 * Morlet wavelet: 1Hz / 3s * Output: Power / Individual TF maps (we want one map for the left, and one for the right) Start the computation: * Click on "Ok" * Back to the Pipeline editor, click on "Run". * After a while, two new files will appear in the tree, representing the TF for the two averaged recordings of the two conditions. |
'''Morlet wavelet options''' * '''Central frequency''': Frequency where the mother wavelet is designed. All other wavelets will be shifted and scaled versions of the mother wavelet * '''Time resolution (FWHM)''': Temporal resolution of the wavelet at the central frequency (in units of Full Width Half Maximum). Click [Display] to see the resolution of the wavelet for other frequencies. '''Compute the following measure''': * The convolution of the signal with complex Morlet wavelets returns the complex coefficients for each frequency/time/sensor. Typically, what we display is the power of the coefficients (square of the amplitude: abs(TF).^2). You can choose if you want to apply this transformation or not. * '''Power''': Computes the "power" transformation immediately after the TF decomposition. This discards the phase information, but produces files that are twice smaller and a lot easier to process. * '''None''': Save the TF coefficients as they are computed. This can be useful if you plan to use these decompositions for other purposes that require the phase. * Some combinations of options may disable this choice. If you select frequency bands, the program will have to compute the power before averaging the values, therefore "none" is not an option. |
| Line 129: | Line 76: |
| * Double-click on the new file to display the time-frequency decomposition of the first sensor. <<BR>><<BR>>{{attachment:example_display.gif}} | |
| Line 289: | Line 237: |
| == Scouts time series == * The previous section introduces an interesting tool to display the TF for a single source, but it is hard to extract meaningful information that can be analyzed statistically or compared between subjects this way. A better option is to define cortical regions of interest (scouts, see [[Tutorials/TutScouts|tutorial #8]]), and compute the TF for the average time series of these scouts. * Define the scouts:<<BR>> * Double click for the source file in the Left condition. * Make sure that your LeftSS and RightSS scouts are still available in your cortex surface.If not, go back to tutorial 8, and recreate them <<BR>><<BR>> {{attachment:createScouts.gif}} * Compute the TF decomposition for these scouts: * Drag and drop the Left source file (''MN: MEG(Constr)'') into ''Process1''. * Select the "Process sources" button. * Click on Run. Select "Frequency > Time-frequency (Morlet wavelets)". * As some scouts are available for the input, you will see this new box:<<BR>><<BR>> {{attachment:selectScouts.gif}} * Select "Use scouts time series", and select both scouts in list. * Click on Edit. This time, most of the options are available again. We are going to compute the full TF matrix at once for the two scouts, so we can compute the power right away, together with time and/or frequency bands. * Leave all the options to the default (linear time, linear frequencies, scout function: After), then click on Ok, then Run. * A new file appears in the tree, as a child of the sources file: "Scouts,Power,1:1:60Hz". * To compute the TF of each scout, the operations performed by the program are the following: * Get time series for each source in the scout * If the scout function is applied __before__ the TF: * Apply the "scout function", that combines the different sources to get only one time series for each scout. * Compute the power of the TF for each scout. * If the scout function is applied __after__ the TF: * Compute the power of the TF for each source * Apply the "scout function", that combines the different sources TF to get one TF only for each scout. * The selection of the "scout function" is done in the "Scout" tab. * '''Output''': * '''Save individual TF maps''': This option stops the computation here, and saves in the database one time-frequency file for each input file. * Be very careful using this option because if you have hundreds of trials, it will fill your hard drive very quickly. * One full TF file in this case contains [nSensors x nTime x nFreq] = 3.397.500 values = 25 Mb. * You can however limit the amount of data produced by using time and/or frequency bands. For 15 time bands and 6 frequency bands, the size of each file drops to: 110 Kb. * '''Save average TF maps''': This option is available only when there are more than one input files (number of files you dropped in the Processes tab). * Instead of saving the TF for each file separately, it automatically computes the average of the power of all the TF. * This is a good choice if you do not plan to use independently all the TF files, because it saves a lot of time and disk space. * Right-click on this file and try all the visualization menus available (including "Time series" and "Power spectrum").<<BR>><<BR>> {{attachment:scoutsTfFile.gif}} |
|
| Line 301: | Line 285: |
| Line 305: | Line 290: |
== Scouts time series == * The previous section introduces an interesting tool to display the TF for a single source, but it is hard to extract meaningful information that can be analyzed statistically or compared between subjects this way. A better option is to define cortical regions of interest (scouts, see [[Tutorials/TutScouts|tutorial #8]]), and compute the TF for the average time series of these scouts. * Define the scouts:<<BR>> * Double click for the source file in the Left condition. * Make sure that your LeftSS and RightSS scouts are still available in your cortex surface.If not, go back to tutorial 8, and recreate them <<BR>><<BR>> {{attachment:createScouts.gif}} * Compute the TF decomposition for these scouts: * Drag and drop the Left source file (''MN: MEG(Constr)'') into ''Process1''. * Select the "Process sources" button. * Click on Run. Select "Frequency > Time-frequency (Morlet wavelets)". * As some scouts are available for the input, you will see this new box:<<BR>><<BR>> {{attachment:selectScouts.gif}} * Select "Use scouts time series", and select both scouts in list. * Click on Edit. This time, most of the options are available again. We are going to compute the full TF matrix at once for the two scouts, so we can compute the power right away, together with time and/or frequency bands. * Leave all the options to the default (linear time, linear frequencies, scout function: After), then click on Ok, then Run. * A new file appears in the tree, as a child of the sources file: "Scouts,Power,1:1:60Hz". * To compute the TF of each scout, the operations performed by the program are the following: * Get time series for each source in the scout * If the scout function is applied __before__ the TF: * Apply the "scout function", that combines the different sources to get only one time series for each scout. * Compute the power of the TF for each scout. * If the scout function is applied __after__ the TF: * Compute the power of the TF for each source * Apply the "scout function", that combines the different sources TF to get one TF only for each scout. * The selection of the "scout function" is done in the "Scout" tab. * Right-click on this file and try all the visualization menus available (including "Time series" and "Power spectrum").<<BR>><<BR>> {{attachment:scoutsTfFile.gif}} |
Tutorial 24: Time-frequency
Authors: Francois Tadel, Dimitrios Pantazis, Sylvain Baillet
This tutorial introduces how to compute time-frequency decompositions of MEG/EEG recordings and cortical currents using complex Morlet wavelets and Hilbert transforms.
Contents
- Introduction
- Morlet wavelets
- Edge effects
- Simple example
- Process options
- Normalized time-frequency maps
- Display time-frequency maps
- Display multiple channels
- Power spectrum and time series
- Time and frequency bands
- Scouts time series
- Cortical sources
- Hilbert transform
- On the hard drive
- Additional discussions on the forum
Introduction
Some of the MEG/EEG signal properties are difficult to access in time domain (graphs time/amplitude). A lot of the information of interest is carried by oscillations at certain frequencies, but the amplitude of these oscillations is sometimes a lot lower than the amplitude of the slower components of the signal, making them difficult to observe.
The averaging in time domain may also lead to a cancellation of these oscillations when they are not strictly locked in phase across trials. Averaging trials in time-frequency domain allows to extract the power of the oscillation regardless of the phase shifts. For a better understanding of this topic, we recommend the lecture of the following article: Bertrand O, Tallon-Baudry C (2000)
Oscillatory gamma activity in humans: a possible role for object representation
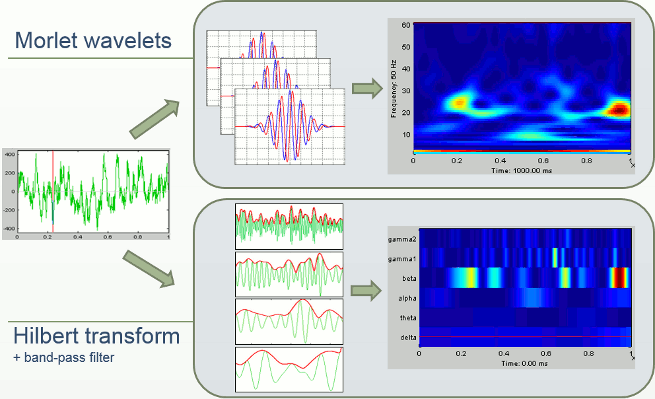
In Brainstorm we offer two approaches for computing time-frquency decompositions (TF): the first one is based on the convolution of the signal with series of complex Morlet wavelets, the second filters the signal in different frequency bands and extract the envelope of the filtered signals using the Hilbert transform.
Morlet wavelets
Complex Morlet wavelets are very popular in EEG/MEG data analysis for time-frequency decomposition. They have the shape of a sinusoid, weighted by a Gaussian kernel, and they can therefore capture local oscillatory components in the time series. An example of this wavelet is shown below, where the blue and red curves represent the real and imaginary part, respectively.
Contrary to the standard short-time Fourier transform, wavelets have variable resolution in time and frequency. When designing the wavelet, we basically decide a trade off between temporal and spectral resolution.
To design the wavelet, we first need to choose a central frequency, ie the frequency where we will define the mother wavelet. All other wavelets will be scaled and shifted versions of the mother wavelet. Unless interested in designing the wavelet at a particular frequency band, the default 1Hz should be fine.
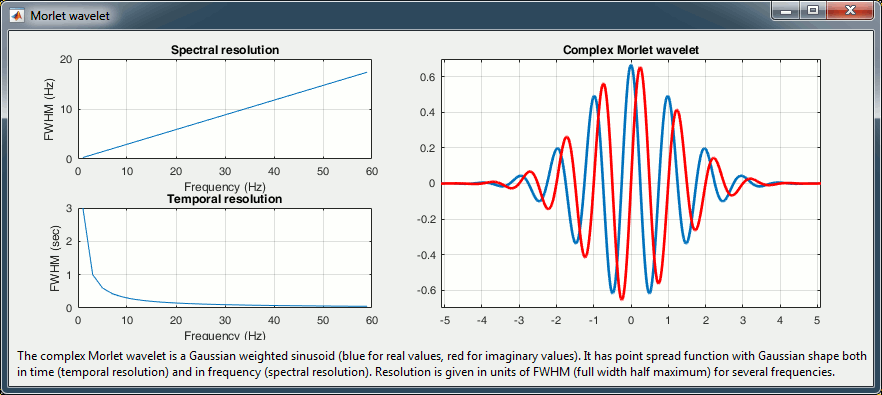
Then, the desirable time resolution for the central frequency should be defined. For example, we may wish to have a temporal resolution of 3 seconds at frequency 1 Hz (default parameters). These two parameters, uniquely define the temporal and spectral resolution of the wavelet for all other frequencies, as shown in the plots below.
Resolution is given in units of Full Width Half Maximum of the Gaussian kernel, both in time and frequency. The relevant plots are given below.
Edge effects
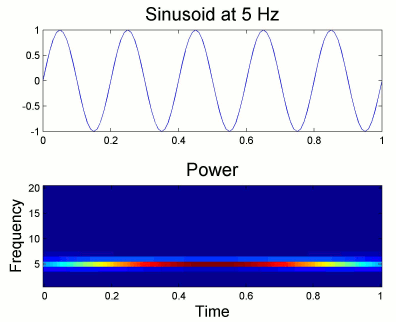
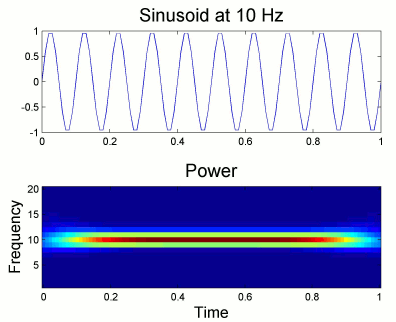
Users should pay attention to edge effects when applying wavelet analysis. Wavelet coefficients are computed by convolving the wavelet kernel with the time series. Similarly to any convolution of signals, there is zero padding at the edges of the time series and therefore the wavelet coefficients are weaker at the beginning and end of the time series.
From the figure above, which designs the Morlet wavelet, we can see that the default wavelet (central frequency 1Hz, FWHM=3sec) has temporal resolution 0.25sec at 5Hz and 0.1sec at 10Hz. In such case, the edge effects are roughly half these times: 125ms in 5Hz and 50ms in 10Hz. Examples of such edge effects are given in the figures below.
Simple example
Let's start with a simple example to explore the options of the time-frequency decomposition process.
In Process1, select the average recordings for the deviant condition for Run#01.
Select process: Frequency > Time-frequency (Morlet wavelets).
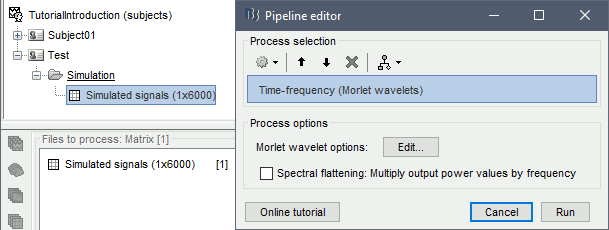
Enter "MEG" for the sensor types, do not select the "Spectral flattening" option.
Click on the button [Edit] to see all the process options.
Time definition: Same as input, Frequency definition: Log 1:40:150, Compute measure: Power.
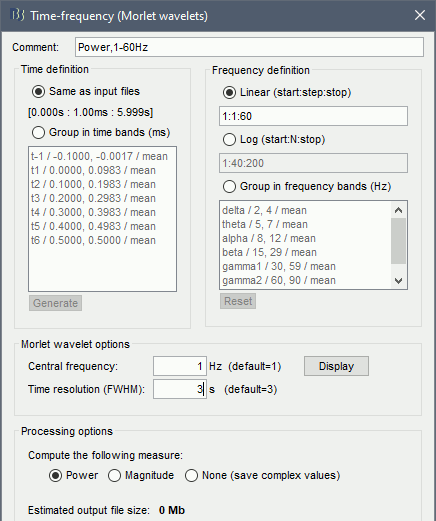
Process options
Comment: String that will be displayed in the database explorer to represent the output file.
Time definition
Same as input file: The output file has the same time definition as the input file.
In this example, it means: 361 samples between -100ms and 500ms.Group in time bands: This option adds a step of computation. First it computes the TF decomposition for all the input file, then averages the power by time band. To define a time band:
Enter your own time bands in the text area, one line per time band, with following format: "name / time definition / function"
- Click on the "Generate" button to automatically create a list of time bands with the same length. You will be asked the maximal length of each time band.
The function is the measure we take to combine the values for all the individual frequencies into one for the frequency band. Possible values are: mean, max, std, median.
Frequency definition: Frequencies for which the power will be estimated at each time instant
Linear: You can specify the frequencies with the Matlab syntax start:step:stop.
The default is "1:1:60", which produces 60 values [1, 2, 3, 4, ..., 59, 60].Group in frequency bands: As for the time definition, this option leads to a two-step process. First it computes the TF decomposition for several values in the frequency band, then it averages the power of TF coefficients per frequency band. To define a frequency band:
One line per frequency band, with the format "name / frequency definition / function"
The frequency definition is a Matlab expression evaluated with an eval() call. If the frequency definition contains only two values, Brainstorm adds two extra values in the middle so that the final averaged value is a bit more robust. Example of valid expressions:
"2,4": Evaluates to [2,4], and then expands to the frequency vector [2, 2.66, 3.33, 4]
"2:0.5:4": evalues to [2 2.5 3 3.5 4]
"2, 2.5, 3, 3.5, 4": Evaluates to [2 2.5 3 3.5 4]The function is the measure we take to combine the values for all the individual frequencies into one for the frequency band. Possible values are: mean, max, std, median.
Morlet wavelet options
Central frequency: Frequency where the mother wavelet is designed. All other wavelets will be shifted and scaled versions of the mother wavelet
Time resolution (FWHM): Temporal resolution of the wavelet at the central frequency (in units of Full Width Half Maximum). Click [Display] to see the resolution of the wavelet for other frequencies.
Compute the following measure:
- The convolution of the signal with complex Morlet wavelets returns the complex coefficients for each frequency/time/sensor. Typically, what we display is the power of the coefficients (square of the amplitude: abs(TF).^2). You can choose if you want to apply this transformation or not.
Power: Computes the "power" transformation immediately after the TF decomposition. This discards the phase information, but produces files that are twice smaller and a lot easier to process.
None: Save the TF coefficients as they are computed. This can be useful if you plan to use these decompositions for other purposes that require the phase.
- Some combinations of options may disable this choice. If you select frequency bands, the program will have to compute the power before averaging the values, therefore "none" is not an option.
Normalized time-frequency maps
Double-click on the new file to display the time-frequency decomposition of the first sensor.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Time-frequency files, once computed, can be processed with Process1 and Process2, exactly as recordings or sources file, as introduced in ?Tutorial #9.
- Only the TF of sensors, clusters and scouts can be processed this way.
- The TF of full source files, as explained above, are saved in the database in an incomplete form, that requires a few more on-the-fly computation steps. The processing engine requires an access to the full TF matrix, which is usually too big to be handled in Matlab for these TF/sources files.
- If you need to average or do some group-analysis on time-frequency data at the source level, the problem has to be simplified a few steps before: for instance by doing your analysis only on a few scouts instead of the full cortex.
Drag and drop one of the TF files in Process1, select the "Time-frequency" button, click on Run.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
- One of the process that is commonly applied on the time-frequency maps is the Z-score, to normalize the values in each frequency band with respect to a baseline. This way, all the frequency bands can be compared directly in terms of amplitude. If we look directly at the power of the Morlet coefficients, the values we observe are always lower in the high frequencies than in the low frequencies. The Z-score fixes this and helps getting figures that are easier to read.
Select the process "Standardize > Compute Z-score (static)". Leave the baseline to the default value (the pre-stimulus interval). Click on run.
- Red values are positive, representing an increase in power with respect with the baseline, blue values are negative, representing a decrease.
The time-frequency map may appear all white or all blue depending on the way the colormap is configured. To edit the colormap, right-click on the figure > Colormap.
Display time-frequency maps
Right-click on the TF file of the Left condition to see what are the possible display options
One channel
- This is the default view for the TF decompositions of full recordings, ie. what would be displayed if you double-click on this file. It shows for one sensor (or one scout/source) the power of each frequency at each time.
- When you load time-frequency related files, the "Display" tab automatically appears in the main Brainstorm window to offer some options for the displays, together with the "current frequency" slider.
Current frequency: Slider that shows the current frequency selected in ALL the figures.
Exactly as the time is Brainstorm, the frequency selection/display is centralized and managed by one control only for all the figures. As a consequence, it is impossible to display TF files with different frequency definitions at the same time. This can be perceived as an annoying limitation, but it allows all the simultaneous displays to be consistent at anytime, and make the interface much more intuitive to manipulate, with lower risks of mistakes in the interpretation of the different figures.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Display tab
List of sensors: This drop-down list shows the current entry displayed in the selected figure. In this case, it is by default the MEG sensor "MLC11". Try to change it, it will update the display of the figure, showing the frequency power for another sensor.
This selection is relative only to the selected figure, and gets updated when you selected another TF figure.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png) -
- ![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Hide edge effects: When this option is selected, the time-frequency coefficients that could not be properly estimated because of a lack of time samples are hidden. It allows you to see only the information that is really reliable in the selected figure. In this example, you can see that it masks a large portion of the window: 375 time samples are not enough to perform a correct time-frequency decomposition. Try to work on larger time windows when using these tools.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Measure: The selection reflects the type of measure that is currently represented in the selected figure. The entries that are enabled depend on the type of data that is saved in the file. In this case, we saved directly the power of the wavelet coefficients in the file, we discarded the angle/phase information, hence the "phase" option is disabled. If we name "power" the values that are saved in the file in the database, the other options would perform the following operations:
Power: Shows the power values unchanged - (left figure)
Magnitude: Shows sqrt(power) - (right figure)
Log: Shows 10 * log10(power)
Mouse and keyboard
Left-click: Selection of current time and frequency.
- If you click anywhere on a TF figure, it will move the current time/frequency cursor to the clicked point, update the time and the frequency sliders in the Brainstorm window, and also update all other visualization figures.
- If you change the current time or the current frequency with the sliders or with another figure, you would observe that the time/frequency cursor position is updated.
Left-click + move: Select a time/frequency range.
Zoom:
Mouse wheel: Zoom/unzoom in time, centered on the current time cursor.
Control + mouse wheel: Zoom/unzoom in frequencies, centered on the current frequency cursor.
Right-click + move, or Control + left-click + move: Move in the zoomed image.
Double-click: Restore initial view.
Note for MacBook users: all the actions involving the mouse wheel can be performed on MacBook laptops by moving up and down two fingers on the pad.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Colormap:
- As explained in the previous tutorials, you can change the colormap by left-clicking on the colorbar on the right of the figure, holding the button, and then by moving the mouse.
- Up/down: Changes the brightness of the colormap.
- Left/Right: Changes the contrast.
This has the same effect than changing the brightness/contrast using the popup menu.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png) -
- ![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Popup menu: Right-click on the figure.
All the menus have been described in the ?tutorial #4 "Exploring the recordings", except for the "Time-frequency selection menu":
(You can see this full menu only if there is a selection defined in the figure)Set selection manually: Does the same thing as drawing a time/freq selection square on a figure, but by typing the values for time and frequency manually.
Export to database: Save the selection for the displayed sensor in a new TF file in the database. However, it is necessary to keep the same time definition for this new file because as it has to stay compatible with the other files it is linked with (here: the recordings file). So in this case, only the selected frequencies are saved, but the time selection is ignored, all the time points are saved.
Remember that only one sensor is exported using this menu, all the others are ignored.Export to file: Same as "Export to database", but the saved file is not registered in the database. The user has to select where to save the file. Unlike the previous menu, only the selected times AND selected frequencies are exported.
Export to Matlab: Same as "Export to file", but the output structure is sent to a variable in the Matlab base workspace instead of being saved to a file.
- Keyboard shortcuts:
Left and right arrows: Change the current time
Page-up and page-down: Also change the current time, but faster (10 time samples instead of one)
Up and down arrows: Change the the sensor displayed in this figure (same as using the drop-list in the Display tab)
Control + up and down arrows: Change the current frequency
Enter: View the original time series for this sensor.
Control + R: View the original MEG recordings (this shortcut and the following are visible in the popup menu)
Control + T: View the time-frequency 2D topography (explained in the next sections)
Control + I: Save as image
Control + D: Dock figure in the Matlab environment.
Display multiple channels
Three menus display the same information (the TF maps of all the channels) with different spatial organizations.
Time-freq: All channels: All the maps are displayed one after the other, in the order they are saved in the file.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Time-freq: 2D Layout: Similarly to the "2D Layout" for the recordings, it shows each the position of each TF map where the sensor is located on a flattened 2D map.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Time-freq: 2D Layout (no overlap): Similar to the "2D Layout", but reorganizing the positions of the small images so that they do not overlap. Overlapping in the "2D Layout" views is not a problem with this old CTF system (151 sensors); it gets much worse with the newer CTF systems (275 sensors), or the Elekta-Neuromag systems (102 chips with 3 sensors at each position). This display is a good intermediate between seeing all the information, and having it spatially organized.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
What to do with these figures:
- Maximize the window to be able to see something
Click: clicking on any small TF image opens a new figure to display only the selected sensor
Shift + click: Same, but opens the original recordings time series of the sensor instead of the TF
Mouse wheel: Zoom / unzoom
Left+right mouse click + move: Move in a zoomed figure.
Power spectrum and time series
These two menus generate similar figures: they represent as a line the evolution of each sensor for one parameter (time or frequency) when the other parameter is fixed.
- Power spectrum: For the given time, show the evolution of all the sensors across the frequencies
- Time series: For the given frequency, show the evolution of all the sensors in time
Open at the same time three figures for the Left/ERF file:
- One channel
- Power spectrum
- Time series.
Then navigate in time and frequencies, and observe how each figure gets updated at each change.
2D topography
Right-click on the TF file in condition Left and select successively the first three topography menus. All these three windows represent the same information, in a slightly different way: a spatial map of the power of the current frequency, for all the sensors at the current time.
- 3D Sensor cap
- 2D Sensor cap
- 2D Disc
Try to move the time and frequency sliders and see what happens. You can open at the same time a "One channel" view, to keep track easily of the current time and frequency.
Keyboard shortcuts:
Left and right arrows: Change the current time
Page-up and page-down: Also change the current time, but faster (10 time samples instead of one)
Up and down arrows: Change the current frequency
Control + E: Display the sensors markers
Control + E again: Display the sensors names
Shift + Click on a sensor: Displays the time-frequency decomposition for that specific sensors.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
2D Layout
The last display mode available for these TF decomposition of recordings is this "2D Layout" topography menu. Right-click on the TF file for the Left condition and select "2D Layout".
This represents spatially the power of the current frequency for all the sensors and all the time points. Try to move the current frequency slider to see how the display changes when increasing the frequency. It is a good example to show that the time resolution increase with the frequency. Below: the "2D Layout" for f=8Hz and f=60Hz.
Useful operations for this window:
Mouse wheel: Zoom / unzoom
Left+right click + move: (or middle click+move) Move into the zoomed window
Shift+mouse wheel: Increase/decrease the sharpness of the time series
Control + mouse wheel: Increase/decrease the length of the time window displayed around the current time.
Click on a line: Select a sensor
Right click + move: Select a group of sensors
Shift + click on a line: Display the time-frequency decomposition for the selected sensor
Right-click: Display popup menu. More options in the "2D Layout options" sub-menu.
Control + E: Display the channels names
All the other shortcuts and the popup menu are already described in ?tutorial #4 "Exploring the recordings".
Time and frequency bands
Frequency bands
Drag and drop the averaged file Left / ERF in the Process tab, click on Run, select the process "Frequency > Time-frequency (Morlet wavelets)". Click on "Edit" to get the options.
- Just change the frequency definition: select "Group in frequency bands", leave the default frequency bands, and click on Ok. Click on Run.
Right-click on the file you've just computed "Power,FreqBands" > "One channel".
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
- You can notice that now the frequency selection is discreet, both in the figure and in the frequency slider, it is not possible to select a frequency in between. At each time and for each sensor, there are now just 6 values, one per frequency band.
- Now if you try to open at the same time the first TF that you have computed "Power,1:1:60Hz", you would get an error message: This frequency definition is not compatible for display with the linear [1...60] scale.
- Display the other possible views (All sensors, 2D Sensor Cap, 2D Layout), and change the current time and the current frequency in different ways (sliders, keyboard, clicks on figures). Play with the open figures until you are completely comfortable with these representations.
Time bands
Compute TF again for file Left / ERF, but this time select the option "Group in time bands" (with "linear" frequency definition). To generate a list of regular time intervals, click on "Generate" and enter "30" ms as the duration of each time band.
- You get 11 time bands: t-2 and t-1 before 0ms, and t1 to t9 after 0ms. Click on Ok to start the computation.
Right-click on this new file "Power,1:1:60Hz" > Time-frequency maps.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
- Observe that the behavior is not the same as with the frequency bands: it is still possible to select a specific time in a time band (using the slider or clicking on the figure). This is because the time definition in the interface is still based on the ERF averaged file time definition, which is "continuous". Only 11 values in time are computed, and when requesting values a specific time point, the interface gets the values associated with the time band that time point belongs to.
- As a consequence, it is possible to open at the same time the first TF file that you have computed: "Power,1:1:60Hz". You would not get an error message like with the frequencies mismatching.
- Display also other types of figures (All sensors, 2D topographies), change current time and frequency several times in several ways (sliders, keyboard, clicks on figures).
Time bands and frequency bands
Scouts time series
The previous section introduces an interesting tool to display the TF for a single source, but it is hard to extract meaningful information that can be analyzed statistically or compared between subjects this way. A better option is to define cortical regions of interest (scouts, see ?tutorial #8), and compute the TF for the average time series of these scouts.
Define the scouts:
- Compute the TF decomposition for these scouts:
Drag and drop the Left source file (MN: MEG(Constr)) into Process1.
- Select the "Process sources" button.
Click on Run. Select "Frequency > Time-frequency (Morlet wavelets)".
As some scouts are available for the input, you will see this new box:
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
- Select "Use scouts time series", and select both scouts in list.
- Click on Edit. This time, most of the options are available again. We are going to compute the full TF matrix at once for the two scouts, so we can compute the power right away, together with time and/or frequency bands.
- Leave all the options to the default (linear time, linear frequencies, scout function: After), then click on Ok, then Run.
- A new file appears in the tree, as a child of the sources file: "Scouts,Power,1:1:60Hz".
- To compute the TF of each scout, the operations performed by the program are the following:
- Get time series for each source in the scout
If the scout function is applied before the TF:
- Apply the "scout function", that combines the different sources to get only one time series for each scout.
- Compute the power of the TF for each scout.
If the scout function is applied after the TF:
- Compute the power of the TF for each source
- Apply the "scout function", that combines the different sources TF to get one TF only for each scout.
- The selection of the "scout function" is done in the "Scout" tab.
Output:
Save individual TF maps: This option stops the computation here, and saves in the database one time-frequency file for each input file.
- Be very careful using this option because if you have hundreds of trials, it will fill your hard drive very quickly.
- One full TF file in this case contains [nSensors x nTime x nFreq] = 3.397.500 values = 25 Mb.
- You can however limit the amount of data produced by using time and/or frequency bands. For 15 time bands and 6 frequency bands, the size of each file drops to: 110 Kb.
Save average TF maps: This option is available only when there are more than one input files (number of files you dropped in the Processes tab).
- Instead of saving the TF for each file separately, it automatically computes the average of the power of all the TF.
- This is a good choice if you do not plan to use independently all the TF files, because it saves a lot of time and disk space.
Right-click on this file and try all the visualization menus available (including "Time series" and "Power spectrum").
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Cortical sources
- How to display the TF decompositions for the source time series?
- It is possible to estimate the TF for each source of the brain, but it would be unrealistic to save this information to a file. The size of the TF matrix would be [nVertices x nTimes x nFrequencies] = [15010 x 375 x 60] complex-double = 5.2 Gb!
- We need then to simplify this problem. Two methods: compute the TF only for a few scouts (next section), or use the linear property of the TF decomposition.
As both the source reconstruction process (ImagingKernel * recordings) and the TF are linear operators, it is possible to exchange them:
TF(Inverse(Recordings)) = Inverse(TF(Recordings))
=> Power(TF(Inverse(Recordings))) = Power(Inverse(TF(Recordings)))So the solution is to estimate the TF of the recordings, and then multiply it on the fly by the ImagingKernel only for the required sources and/or frequency and/or time instant. This is done in a completely transparent way from the user point a view.
- Start the computation of the TF for a source file:
Drag and drop the source file computed for the Left / ERF file in Process1.
- Select the "Process sources" button.
- Click on Run. Select "Time-frequency (Morlet wavelets)". Do NOT check the box "Use scouts time series".
- Click on Edit. There is a new option available, that offers between saving only the kernel and the TF of sensors, or the full TF matrix. Select the optimized solution.
Note that this solution is not compatible with any of the operations that requires to compute explicitly the power of all the TF decomposition: time bands and frequency bands.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Click on Ok, then Run. A new file appears in the database explorer, as a child of the source file. Right-click on it, try all the visualization menus.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
If you understood well how to use the visualization of both the sources and the time-frequency maps, the manipulation of these figures should be rather intuitive. Change several times the current time and the current frequency in different ways. Change current time and frequency so that you can observe the main response in the somatosensory cortex (eg. t = 46.40ms, f = 45Hz).
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Shift + click on the cortex surface: This is a useful shortcut that you should remember. It displays the TF decomposition of the selected source.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Right-click on the brain: This selects the closest vertex and displays the popup menu at the same time. The first three menus are relative to the source that was just clicked.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Hilbert transform
On the hard drive
Right click one of the TF files, and select the menu File > View file contents, to have a look to what is the actual contents of these structures.
TF: [nSensors x nTime x nFrequency] matrix containing all the values of the time-frequency decomposition (complex wavelet coefficients, or power of amplitudes)
- nSensors: Number of sensors for which the TF has been estimated
- nTime:
If there are time bands defined, nTime = size(TimeBands,1) = number of time bands
- If there are no time bands (linear time sampling): nTime = length(Time)
- nFrequency:
- If there are frequency bands defined, nFrequency = size(Freqs,1) = number of frequency bands
- If there are no frequency bands (linear frequency sampling): nFrequency = length(Freqs)
Comment: String displayed in the database explorer to represent the file.
DataType: Explains from what kind of data this file was computed. Possible values are: 'data', 'results', 'scout, 'matrix'...
Time: Time vector used to estimate this file.
TimeBands: Description of the time bands. Cell array {nTimeBands x 3}, where each line represents a time band {'band_name', 'time definition', 'function'}
Freqs:
- If frequency bands: Cell array {nFreqBands x 3}, where each line represents a frequency band {'band_name', 'frequency definition', 'function'}
- In linear frequency sampling: vector containing all the frequencies
RowNames: Cell array of strings that describes each row of the TF matrix. In this specific case, it would be the list of all the MEG sensor names. But it could also be a list of names of scouts or clusters.
Measure: Contains the name of the function that was applied right after the computation of the wavelet coefficients. So it represents the type of data contained in the TF matrix. Possible values:
- 'none': No measure applied, TF contains the complex wavelet coefficients.
- 'power': TF contains the power for each frequency, ie. the square of the amplitude: abs(coefficients).^2
- 'magnitude': abs(coefficients)
- 'log': 10 * log10( abs(coefficients).^2)
- 'phase': angle(coefficients)
Method: String that identifies the process that generated the file; possible values: {'morlet', 'fft', 'psd', 'hilbert', 'corr', 'cohere', 'granger', 'plv'}
DataFile: Initial file from which this file was computed. In the database explorer, the TF file will be shown as a child of this DataFile file.
nAvg: Number of trials that were averaged to obtain this file (copied from the recordings file at computation)
Options: Options that were selected in the time-frequency options window.
History: List of operations performed on this file (better visualized with popup menu File > View file history)
Document file tags
Additional discussions on the forum
Time and frequency resolution: http://neuroimage.usc.edu/forums/showthread.php?1848