|
Size: 8151
Comment:
|
Size: 8140
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 14: | Line 14: |
| We need three realistic layers extracted from the subject's MRI (scalp, inner skull, outer skull), plus the source space (cortical surface). Those layers are sometimes complicated to generate. In the case you do not have any way to calculate the inner skull and outer skull surfaces, Brainstorm can generate rough approximations based on the subject's cortex and head surfaces and Colin27's inner and outer skull surfaces.The surfaces created with Brainstorm are by construction non-intersecting. | We need three realistic layers extracted from the subject's MRI (scalp, inner skull, outer skull), plus the source space (cortical surface). These layers are sometimes complicated to generate. In the case you do not have any way to calculate the inner skull and outer skull surfaces, Brainstorm can generate rough approximations based on the subject's cortex and head surfaces and ICBM152's inner and outer skull surfaces.The surfaces created with Brainstorm are by construction non-intersecting. |
| Line 24: | Line 24: |
| * The default values are high (1922 vertices per layer), to ensure that by default the results are good. If you can afford it computationally, you should use those '''1922 vertices''' per layer. If you have to decrease those values, you can try for instance with '''1082 vertices '''(scalp) and '''642 vertices '''(outer skull and innser skull). * In output, you will probably not get the number of vertices you are asked for. The algorithm generating those surfaces uses a fixed set of apolar spheres with the following number of vertices: 273, 362, 482, 642, 812, 1082, 1442, 1922, 2432, 2562, 3242, 4322... * If you are using the default Colin27 anatomy, you do not need to run this, you can directly use the head/outerskull/innerskull surfaces in the the default anatomy folder. The number of vertices is much lower, but we know this anatomy to be stable, you should not need to recalculate new surfaces with more vertices. |
* The default values are high (1922 vertices per layer), to ensure that by default the results are good. If you can afford it computationally, you should use these '''1922 vertices''' per layer. If you have to decrease these values, you can try for instance with '''1082 vertices '''(scalp) and '''642 vertices '''(outer skull and innser skull). * In output, you will probably not get the number of vertices you are asked for. The algorithm generating these surfaces uses a fixed set of apolar spheres with the following number of vertices: 273, 362, 482, 642, 812, 1082, 1442, 1922, 2432, 2562, 3242, 4322... * If you are using the default ICBM152 anatomy, you do not need to run this, you can directly use the head/outerskull/innerskull surfaces in the the default anatomy folder. The number of vertices is much lower, but we know this anatomy to be stable, you should not need to recalculate new surfaces with more vertices. |
| Line 39: | Line 39: |
| 1. Once downloaded, the calculation of the head model will start automatically. You may wait for a very long time. Be patient, it's worth it...<<BR>><<BR>> {{attachment:openmeegWait.gif}} | 1. Once downloaded, the calculation of the head model will automatically start. You may wait for a very long time. Be patient, it's worth it...<<BR>><<BR>> {{attachment:openmeegWait.gif}} |
| Line 76: | Line 76: |
| * Gramfort, T. Papadopoulo, E. Olivi, M. Clerc. OpenMEEG: opensource software for quasistatic bioelectromagnetics, [[http://www.biomedical-engineering-online.com/content/9/1/45|BioMedical Engineering OnLine 45:9, 2010]] * Kybic J, Clerc M, Abboud T, Faugeras O, Keriven R, Papadopoulo T. A common formalism for the integral formulations of the forward EEG problem. [[http://ieeexplore.ieee.org/xpls/abs_all.jsp?isnumber=30034&arnumber=1375158&count=10&index=1|IEEE Transactions on Medical Imaging, 24:12-28, 2005.]] [[ftp://ftp-sop.inria.fr/odyssee/Publications/2005/kybic-clerc-etal:05.pdf|[pdf]]] |
* Gramfort A, Papadopoulo T, Olivi E, Clerc M<<BR>>[[http://www.biomedical-engineering-online.com/content/9/1/45|OpenMEEG: opensource software for quasistatic bioelectromagnetics]]<<BR>>BioMedical Engineering OnLine 45:9, 2010 * Kybic J, Clerc M, Abboud T, Faugeras O, Keriven R, Papadopoulo T<<BR>>[[http://ieeexplore.ieee.org/xpls/abs_all.jsp?isnumber=30034&arnumber=1375158&count=10&index=1|A common formalism for the integral formulations of the forward EEG problem]]<<BR>>IEEE Transactions on Medical Imaging, 24:12-28, 2005 | [[ftp://ftp-sop.inria.fr/odyssee/Publications/2005/kybic-clerc-etal:05.pdf|pdf]] |
| Line 79: | Line 79: |
| <<EmbedContent(http://neuroimage.usc.edu/brainstorm3_register/get_feedback.php?Tutorials/TutBem)>> | <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/TutBem)>> |
OpenMEEG BEM head model
Authors: Francois Tadel, Alexandre Gramfort
This forward model uses a symmetric boundary element method (symmetric BEM) and was developed by the French public research institute INRIA (website). It uses three realistic layers (scalp, inner skull, outer skull). The goal of this forward solution is mostly for EEG users, to provide a better model than the existing "3-shell sphere". It is not necessary for MEG users, as the "overlapping spheres" method gives relatively similar results but faster. Nevertheless, for demonstration purpose or in case you want to use the same head model for both EEG and MEG data, we are going to compute this solution on MEG recordings, using the existing TutorialCTF protocol.

Requirements
On Windows 64bit systems, you need to install an extra Microsoft software package to run OpenMEEG: Microsoft Visual C++ 2010 Redistributable Package (x64).
http://www.microsoft.com/en-us/download/details.aspx?id=14632
BEM surfaces
We need three realistic layers extracted from the subject's MRI (scalp, inner skull, outer skull), plus the source space (cortical surface). These layers are sometimes complicated to generate. In the case you do not have any way to calculate the inner skull and outer skull surfaces, Brainstorm can generate rough approximations based on the subject's cortex and head surfaces and ICBM152's inner and outer skull surfaces.The surfaces created with Brainstorm are by construction non-intersecting.
- Select the TutorialCTF protocol
- Go to the anatomy view (first button on top of the database explorer)
Right-click on Subject01 > Generate BEM surfaces...You can set the approximate number of vertices you want for each layer. For the example, leave the default parameters and click on Ok.
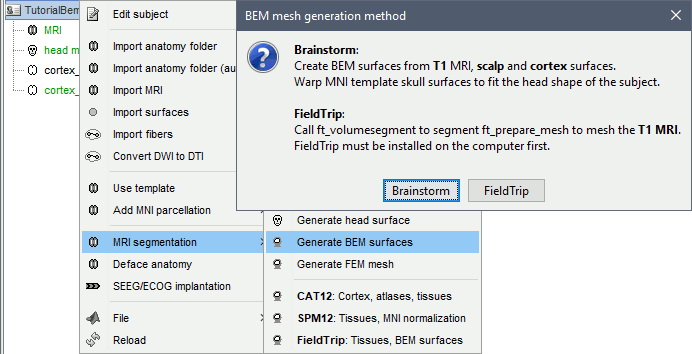
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
At the end of the process, three new surfaces are added to the database:
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
How to set the number of vertices per layer
- The more vertices you have for each layer, the more accurate your head model will be. If there are not enough vertices, OpenMEEG may consider that the surfaces are intersecting and crash before the computation, or it may finish but at the end the model may be instable with a few vertices with values much higher than all the others.
- On the other hand, using more vertices on each layer increases a lot the amount of memory necessary and the computation time. If your system hangs, if it takes 24hrs to run or if you get "Out of memory" errors, you have to decrease the number of vertices for each layer.
The default values are high (1922 vertices per layer), to ensure that by default the results are good. If you can afford it computationally, you should use these 1922 vertices per layer. If you have to decrease these values, you can try for instance with 1082 vertices (scalp) and 642 vertices (outer skull and innser skull).
- In output, you will probably not get the number of vertices you are asked for. The algorithm generating these surfaces uses a fixed set of apolar spheres with the following number of vertices: 273, 362, 482, 642, 812, 1082, 1442, 1922, 2432, 2562, 3242, 4322...
- If you are using the default ICBM152 anatomy, you do not need to run this, you can directly use the head/outerskull/innerskull surfaces in the the default anatomy folder. The number of vertices is much lower, but we know this anatomy to be stable, you should not need to recalculate new surfaces with more vertices.
Forward model
- Switch to the view "Functional data (sorted by subjects)"
Right-click on Subject01 / Right > Compute head model. Select OpenMEEG BEM in the list. Click on Run.
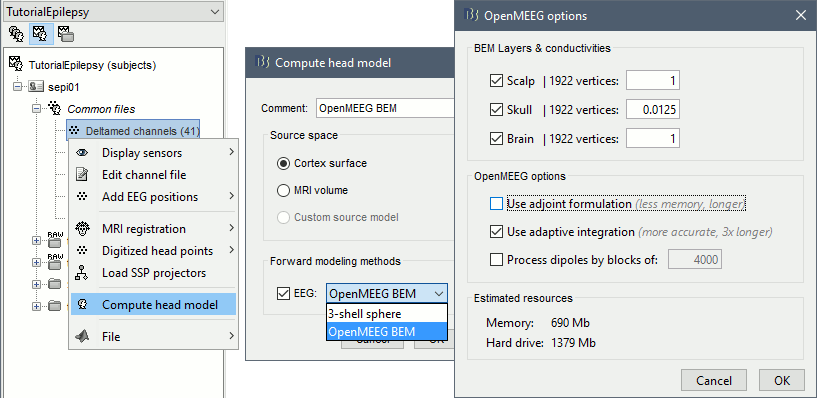
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
- Explanations of the options:
BEM Layers & conductivities: In this panel you can select how many layers you want to use for the BEM model. The possible options in our case are Scalp (=head), Skull (=outer skull), and Brain (=inner skull). The values on the right are the relative conductivities of each layer.
- For EEG: Use all the layers
- For MEG: Use only the inner skull layer
Use adjoint formulation: Decrease significantly the amount of memory that is necessary in memory and on the hard drive, but much longer.
Use adaptive integration: This produces a much more accurate solution, mostly in the cases of dipoles (vertices of the cortex surface) that are very close to the inner skull layer. You should keep this option selected.
Process dipoles by blocks: If you consistently run into crashes, it's probably because you don't have enough memory available. You can try to process the dipoles by smaller blocks. It would be longer but would probably work. Do not select unless you cannot compute the solution with having memory errors.
Download OpenMEEG software? Yes.
This will download the latest version of OpenMEEG binaries for your operating system from INRIA website, and install it in Brainstorm user folder (~username/.brainstorm/openmeeg/)
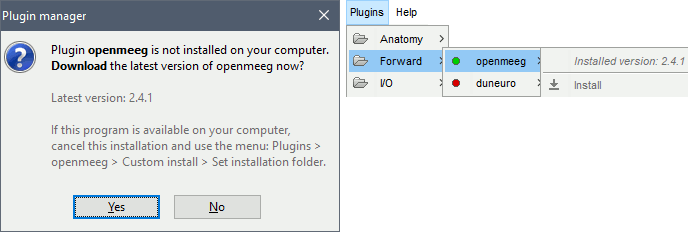
Once downloaded, the calculation of the head model will automatically start. You may wait for a very long time. Be patient, it's worth it...

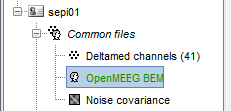
You have now a new head model in your condition, that you can use exactly the same way you would use your previous "Overlapping sphere" model.
Additional discussions on the forum
Download problems:
Binary issues / Error #-1073741515:
Library issues / Error #1:
Memory allocation failed:
Intersecting meshes:
Format of the Gain matrix in the headmodel file:
License
OpenMEEG is distributed under the French opensource license CeCILL-B. It is intended to give users the freedom to modify and redistribute the software. It is therefore compatible with popular opensource licenses such as the GPL and BSD licenses. The CeCILL-B license imposes to anybody distributing a software incorporating OpenMEEG the obligation to give credits (by citing the appropriate publications), in order for all contributions to be properly identified and acknowledged.
The references to be acknowledged are:
Gramfort A, Papadopoulo T, Olivi E, Clerc M
OpenMEEG: opensource software for quasistatic bioelectromagnetics
BioMedical Engineering OnLine 45:9, 2010Kybic J, Clerc M, Abboud T, Faugeras O, Keriven R, Papadopoulo T
A common formalism for the integral formulations of the forward EEG problem
IEEE Transactions on Medical Imaging, 24:12-28, 2005 | pdf
