|
Size: 690
Comment:
|
Size: 7395
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| ## page was renamed from Tutorials/TutFirstStart | |
| Line 2: | Line 3: |
| Objectives: Create a protocol with one subject, that uses the default anatomy Colin27. |
|
| Line 6: | Line 9: |
| 1. Domewhere else, create an empty folder to store your Brainstorm database (for instance "''brainstorm_database''", or "''brainstorm_protocols''"). 1. Do '''not '''create this folder in the '''brainstorm3 program folder''' 1. This directory will be managed completely automatically by Brainstorm 1. '''Never put your original data files''' in this directory: it could prevent Brainstorm to manage it efficiently 1. All the data in Brainstorm database need to be imported via the GUI |
1. With some Matlab installations, you may observe an error message in the command window, about the compilation of the MEX files. Some files from the permutation tests functions need to be compiled on your own operating system, and you may not be able to use some statistical tests if you have such an error. You may ignore this problem, it will be fixed later. 1. The ''Licence Agreement'' will be display. Read the licence file, and click on "Agree" if you agree with it. 1. If you have two screens on your computer, you may have a message asking if you want to use your second screen. You can try to say yes: you will have the Brainstorm window on primary screen and the data figures on the secondary one). If does not work properly, go to the menu Options > GUI, and uncheck the menu "Use two screens when available". 1. Then you will be asked to specify Brainstorm database directory. Read carefully the ''Important notes'' and then create or select an empty directory to store your Brainstorm database (for instance "''brainstorm_database''", or "''brainstorm_protocols''"). 1. After a last message asking you to create or load a protocol, you should see the main Brainstorm window == Main interface window == {{attachment:main_window.gif}} == Create first protocol == 1. Click on the ''Protocol selection'' drop list, and select ''Create new protocol''. 1. Edit the protocol name and enter: "!TutorialFirstSteps". It will automatically update the paths (''Anatomy path'' and ''Datasets path''. 1. Select the "''MNI Colin27''" as the default anatomy. * This anatomy (MRI & surfaces) will be available if needed by all the subjects in the protocol. * This can be changed later, using the "''Use default''" menu in the subjects' popup menu. 1. ''Subjects configuration'' panel: * These are the default settings that will be applied when creating new subjects. It is then possible to override those settings for each subject. * Our objective is to create one subject that uses the default anatomy Colin27, so select the option "''Yes, use protocol's default anatomy''". * The option selected for ''Default channel file'' is not important because we are not going to import recordings (and channel files, ie. files that describe each sensor: position, orientation, name...). This point will be described in another tutorial. 1. Once you get something like the following figure, click on ''Create''.<<BR>><<BR>> {{attachment:createNewProtocol.gif}} 1. Confirm that you want to create the directories 1. The ''MRI Viewer'' window opens automatically and displays the Colin27 brain, and a message box informs you that you should select some points on the MRI. Those fiducial points are used to register the MRI with the MEG or EEG sensors, when you import recordings. We don't need to define them now, because we are not going to import any recordings in this turorial, but let's play with the MRI Viewer Viewer. == MRI Viewer and fiducials selection == {{attachment:mriViewer.gif}} * You can navigate through the volume by clicking anywhere in the slices, or by using the sliders below each image. * There are four small buttons above each image. They may be useful when you import your own MR volumes, if they are not oriented in the way Brainstorm want them (e.g. you see the ''Sagittal ''title on top of the coronal view; or the small white "''P''", which is supposed to indicate the posterior part of the head, points the nose). Leave your mouse a few seconds over button one to read their description tooltip. Click on them and try to understand what they are doing. * Images' contrast can be changed with the mouse: click and hold the right button on one image, and move your mouse up and down (this operation will be later referred as ''right-click+move''). * You can define six points on the MRI: * Three to define the [[CoordinateSystems|Subject Coordinate System (SCS)]]: Nasion (NAS), Left pre-auricular point (LPA), Right pre-auricular point (RPA), * Three to define the[[CoordinateSystems|Talairach coordinate system]] (referred here as ''Normalized CS''): Anterior commissure (AC), Posterior commissure (PC), and any Interhemispheric point (IH).<<BR>> => To know more about [[CoordinateSystems|how to define the fiducial points]]. * To define a point, position the three views a the point your want to define (pointed by the 3 crosses), and click on the ''Set ''button. Later you can click on ''View ''to move the views to this point again. * You can see that all the points are already defined on this MRI, but you should '''always '''check their positions. For many reasons, each research group has a different way to define those points. So you may not position the ears or the nasion exactly the same way as we do. So do not hesitate to move the fiducials to better fit your needings; if not the results might be much less accurate. * The coordinates panel shows the (x,y,z) coordinates in all the coordinate systems managed by Brainstorm: MRI, SCS, and Talairach. * Once you're done, click on ''Cancel ''(unless you really want to save all your experimentations). == Protocol exploration == The protocol is created and you can now see it in the database explorer, in Brainstorm window. It is represented by the top-most node in the tree. . {{attachment:treeNewProtocol.gif}} * You can switch between anatomy and functional data with the first three buttons in the toolbar. They are no subjects in the database yet, so the ''Functional data'' views are completely empty. However, in the ''Anatomy ''view, there is a ''(Default anatomy)'' node; it contains the MRI and surfaces that can be used by default for the subjects without individual anatomy. * Display the ''Anatomy ''view, click on the small "+" to expand the contents of the ''(Default anatomy)'' node. There is one MRI and several surfaces, identified by different icons. * All you can do with each node (anatomy files, subjects, protocol), is accessible by right-clicking on it. Take 10 minutes to explore all the popup menus by yourself. Don't be afraid to click everywhere you want, at this point you cannot damage anything. * If you explored well, you should have found the "''New subject''" menu in the protocol's popup menu. Click on it.<<BR>><<BR>> {{attachment:createNewSubject.gif}} * Usually, the only thing you need to do here is to edit the name of the subject. * But you can also override here the protocol's defaults regarding the sharing of anatomy and channel file between subjects and conditions. For the example, leave selected the "''Yes, use default anatomy''" option. The click on ''Save''. * You should see a the new subject in both ''Anatomy ''and ''Functional data'' views. * In the Anatomy view, the subject node only contains a link called (Default anatomy). It is here just to remind that the subject is using the default ''MNI Colin27'' anatomy.<<BR>> {{attachment:subjectDefaultAnat.gif}} == Surface visualization == |
Tutorials: First steps
Objectives: Create a protocol with one subject, that uses the default anatomy Colin27.
Contents
Running Brainstorm for the first time
- Start Matlab, go to brainstorm3 directory, and type "brainstorm3" in Matlab command window.
- With some Matlab installations, you may observe an error message in the command window, about the compilation of the MEX files. Some files from the permutation tests functions need to be compiled on your own operating system, and you may not be able to use some statistical tests if you have such an error. You may ignore this problem, it will be fixed later.
The Licence Agreement will be display. Read the licence file, and click on "Agree" if you agree with it.
If you have two screens on your computer, you may have a message asking if you want to use your second screen. You can try to say yes: you will have the Brainstorm window on primary screen and the data figures on the secondary one). If does not work properly, go to the menu Options > GUI, and uncheck the menu "Use two screens when available".
Then you will be asked to specify Brainstorm database directory. Read carefully the Important notes and then create or select an empty directory to store your Brainstorm database (for instance "brainstorm_database", or "brainstorm_protocols").
- After a last message asking you to create or load a protocol, you should see the main Brainstorm window
Main interface window
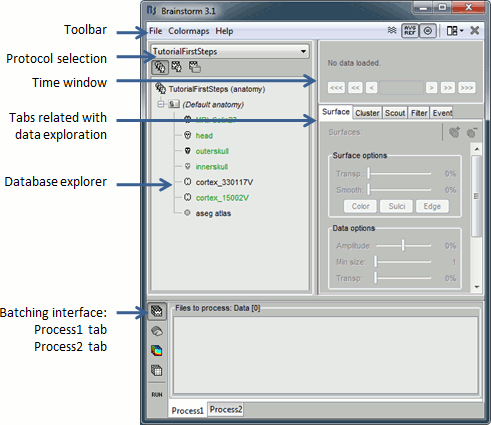
Create first protocol
Click on the Protocol selection drop list, and select Create new protocol.
Edit the protocol name and enter: "TutorialFirstSteps". It will automatically update the paths (Anatomy path and Datasets path.
Select the "MNI Colin27" as the default anatomy.
This anatomy (MRI & surfaces) will be available if needed by all the subjects in the protocol.
This can be changed later, using the "Use default" menu in the subjects' popup menu.
Subjects configuration panel:
- These are the default settings that will be applied when creating new subjects. It is then possible to override those settings for each subject.
Our objective is to create one subject that uses the default anatomy Colin27, so select the option "Yes, use protocol's default anatomy".
The option selected for Default channel file is not important because we are not going to import recordings (and channel files, ie. files that describe each sensor: position, orientation, name...). This point will be described in another tutorial.
Once you get something like the following figure, click on Create.
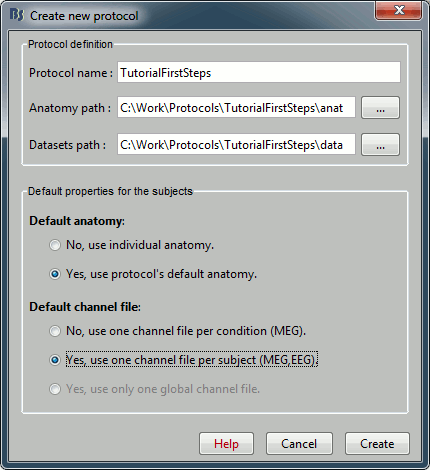
- Confirm that you want to create the directories
The MRI Viewer window opens automatically and displays the Colin27 brain, and a message box informs you that you should select some points on the MRI. Those fiducial points are used to register the MRI with the MEG or EEG sensors, when you import recordings. We don't need to define them now, because we are not going to import any recordings in this turorial, but let's play with the MRI Viewer Viewer.
MRI Viewer and fiducials selection
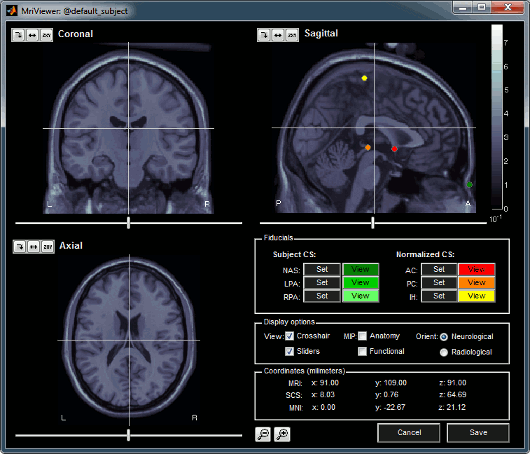
- You can navigate through the volume by clicking anywhere in the slices, or by using the sliders below each image.
There are four small buttons above each image. They may be useful when you import your own MR volumes, if they are not oriented in the way Brainstorm want them (e.g. you see the Sagittal title on top of the coronal view; or the small white "P", which is supposed to indicate the posterior part of the head, points the nose). Leave your mouse a few seconds over button one to read their description tooltip. Click on them and try to understand what they are doing.
Images' contrast can be changed with the mouse: click and hold the right button on one image, and move your mouse up and down (this operation will be later referred as right-click+move).
- You can define six points on the MRI:
Three to define the Subject Coordinate System (SCS): Nasion (NAS), Left pre-auricular point (LPA), Right pre-auricular point (RPA),
Three to define theTalairach coordinate system (referred here as Normalized CS): Anterior commissure (AC), Posterior commissure (PC), and any Interhemispheric point (IH).
=> To know more about how to define the fiducial points.To define a point, position the three views a the point your want to define (pointed by the 3 crosses), and click on the Set button. Later you can click on View to move the views to this point again.
You can see that all the points are already defined on this MRI, but you should always check their positions. For many reasons, each research group has a different way to define those points. So you may not position the ears or the nasion exactly the same way as we do. So do not hesitate to move the fiducials to better fit your needings; if not the results might be much less accurate.
- The coordinates panel shows the (x,y,z) coordinates in all the coordinate systems managed by Brainstorm: MRI, SCS, and Talairach.
Once you're done, click on Cancel (unless you really want to save all your experimentations).
Protocol exploration
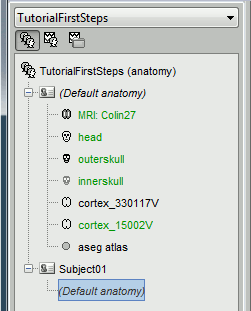
The protocol is created and you can now see it in the database explorer, in Brainstorm window. It is represented by the top-most node in the tree.
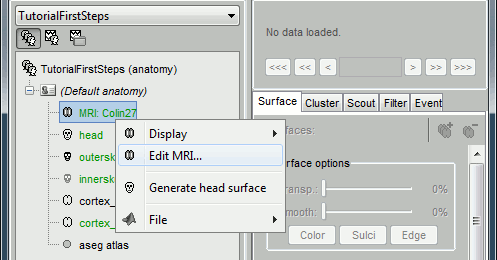
You can switch between anatomy and functional data with the first three buttons in the toolbar. They are no subjects in the database yet, so the Functional data views are completely empty. However, in the Anatomy view, there is a (Default anatomy) node; it contains the MRI and surfaces that can be used by default for the subjects without individual anatomy.
Display the Anatomy view, click on the small "+" to expand the contents of the (Default anatomy) node. There is one MRI and several surfaces, identified by different icons.
- All you can do with each node (anatomy files, subjects, protocol), is accessible by right-clicking on it. Take 10 minutes to explore all the popup menus by yourself. Don't be afraid to click everywhere you want, at this point you cannot damage anything.
If you explored well, you should have found the "New subject" menu in the protocol's popup menu. Click on it.
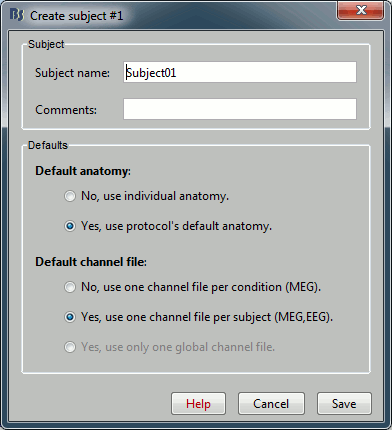
- Usually, the only thing you need to do here is to edit the name of the subject.
But you can also override here the protocol's defaults regarding the sharing of anatomy and channel file between subjects and conditions. For the example, leave selected the "Yes, use default anatomy" option. The click on Save.
You should see a the new subject in both Anatomy and Functional data views.
In the Anatomy view, the subject node only contains a link called (Default anatomy). It is here just to remind that the subject is using the default MNI Colin27 anatomy.