|
Size: 74
Comment:
|
Size: 4508
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| ## page was renamed from TutImportAnatomy | |
| Line 3: | Line 2: |
| Objectives: Import anatomy for CTF MEG tutorial. <<TableOfContents>> == Create protocol == 1. Download '''bst_tutorial_somatotopy_ctf.zip''' file ([[http://neuroimage.usc.edu/brainstorm3_register/download.php|Download]] section), and unzip it somewhere. Data description in the next tutorial. * Warning: Do '''NOT '''unzip it in the Brainstorm '''database or program directories'''. This package contains files in original file formats, and need to be imported through the interface to be processed with Brainstorm. 1. Create a new protocol called ''TutorialCTF ''with the following options: * "No, use individual anatomy", because the individual subject anatomy is available * "No, use one channel file per condition (MEG)", because we are going to process regular MEG data in the next tutorial 1. Create a new subject (let's keep the default name: ''Subject01'') == Import MRI == 1. Go to the ''Anatomy ''view of the protocol files (very first button in Brainstorm window toolbar). 1. Right click on subject node, and select menu "''Import MRI...''"<<BR>><<BR>> {{attachment:menuImportMri.gif}} 1. Select file "''TutorialCtf/Anatomy/01.mri''". 1. If you didn't modified any Brainstorm option yet, you should get an error message, saying "Unrecognized format, try switching byte order". This is because not all the computers process the binary data the same way. To learn more about endianness, you can consult [[http://en.wikipedia.org/wiki/Endianness|this Wikipedia page]]. Depending on the type of computer which wrote the binary files, you may need to read them as ''little-endian'' or ''big-endian''. To change this setting, go to the ''Options > Byte order'' menu in main window. You may need to modify this setting again to import other files.<<BR>><<BR>> {{attachment:menuByteOrder.gif}} 1. Try to import the MRI again. It should be working now. The MRI viewer is displayed, together with a message box that tells you what to do. Follow the instructions. The MRI is already well oriented so you can directly process with the fiducials selection. You will see that three fiducials were already identified in the MRI file (NAS, LPA, RPA); you just need to add the three other ones (AC, PC, IH). If you are not sure how to do this, consult this page: CoordinateSystems. == Import surfaces == The MRI is now available in the anatomy folder for ''Subject01''. Let's now proceed with the surfaces. The envelopes we are going to import in this tutorial were extracted using [[Links|BrainVISA]] software, and their names and types may differ from other software solutions. The next tutorials will show you how to import surfaces from other programs ([[Links|BrainSuite]], [[Links|FreeSurfer]]). 1. Right click on the subject node again, and click on "''Import surfaces...''". Select '''at the same time''' (holding ''Shift ''or ''Ctrl ''key) all the files in the directory "''TutorialCtf/Anatomy/BrainVISA''" (head, Lhemi, Rhemi). Click on "Open" and answer ''Yes ''to the question: "Align surfaces with MRI now?". * Note: The coordinate system used to localize in 3D the vertices of each surface depends on the software that created them. When importing the surfaces, BrainStorm registers them with the MRI using its fiducials points (NAS, LPA, RPA). This simple algorithm needs the head (ie. scalp) surface, so if you try to import only the cortical surface, it will not be well registered with the MRI, and all the following computations will fail. 1. You should see the three surfaces in the database tree. <<BR>><<BR>> {{attachment:treeSurfaces.gif}} * One has been identified automatically as a ''scalp'' (=''head'') surface, because its original filename contained the "''scalp''" keyword. Usually, Brainstorm identifies file types with this kind of tags in the file names. * The two other files where not identified automatically. They contain the envelopes of left and right hemispheres of the brain, but ''Lhemi ''and ''Rhemi ''are not keywords that are identified automatically by Brainstorm. * Try to display the surfaces by double-clicking on them, or with the popup menu ''Display''. 1. In the ''Surfaces ''tab, the numbers of faces and vertices are displayed == What happend on the harddrive == If you are curious to know how all this anatomy information was stored in the database directory on the hard drive, you can have a look in your ''brainstorm_database/TutorialCTF'' directory. |
Import individual anatomy
Objectives: Import anatomy for CTF MEG tutorial.
Contents
Create protocol
Download bst_tutorial_somatotopy_ctf.zip file (Download section), and unzip it somewhere. Data description in the next tutorial.
Warning: Do NOT unzip it in the Brainstorm database or program directories. This package contains files in original file formats, and need to be imported through the interface to be processed with Brainstorm.
Create a new protocol called TutorialCTF with the following options:
- "No, use individual anatomy", because the individual subject anatomy is available
- "No, use one channel file per condition (MEG)", because we are going to process regular MEG data in the next tutorial
Create a new subject (let's keep the default name: Subject01)
Import MRI
Go to the Anatomy view of the protocol files (very first button in Brainstorm window toolbar).
Right click on subject node, and select menu "Import MRI..."
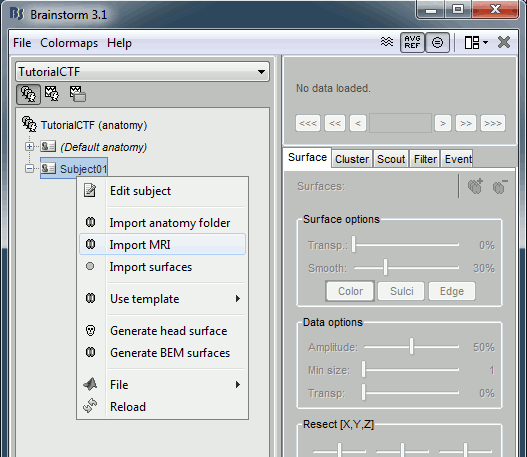
Select file "TutorialCtf/Anatomy/01.mri".
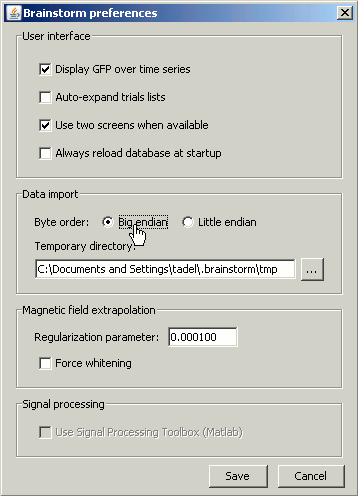
If you didn't modified any Brainstorm option yet, you should get an error message, saying "Unrecognized format, try switching byte order". This is because not all the computers process the binary data the same way. To learn more about endianness, you can consult this Wikipedia page. Depending on the type of computer which wrote the binary files, you may need to read them as little-endian or big-endian. To change this setting, go to the Options > Byte order menu in main window. You may need to modify this setting again to import other files.
Try to import the MRI again. It should be working now. The MRI viewer is displayed, together with a message box that tells you what to do. Follow the instructions. The MRI is already well oriented so you can directly process with the fiducials selection. You will see that three fiducials were already identified in the MRI file (NAS, LPA, RPA); you just need to add the three other ones (AC, PC, IH). If you are not sure how to do this, consult this page: CoordinateSystems.
Import surfaces
The MRI is now available in the anatomy folder for Subject01. Let's now proceed with the surfaces. The envelopes we are going to import in this tutorial were extracted using ?BrainVISA software, and their names and types may differ from other software solutions. The next tutorials will show you how to import surfaces from other programs (?BrainSuite, ?FreeSurfer).
Right click on the subject node again, and click on "Import surfaces...". Select at the same time (holding Shift or Ctrl key) all the files in the directory "TutorialCtf/Anatomy/BrainVISA" (head, Lhemi, Rhemi). Click on "Open" and answer Yes to the question: "Align surfaces with MRI now?".
Note: The coordinate system used to localize in 3D the vertices of each surface depends on the software that created them. When importing the surfaces, BrainStorm registers them with the MRI using its fiducials points (NAS, LPA, RPA). This simple algorithm needs the head (ie. scalp) surface, so if you try to import only the cortical surface, it will not be well registered with the MRI, and all the following computations will fail.
You should see the three surfaces in the database tree.
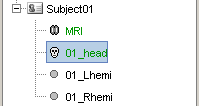
One has been identified automatically as a scalp (=head) surface, because its original filename contained the "scalp" keyword. Usually, Brainstorm identifies file types with this kind of tags in the file names.
The two other files where not identified automatically. They contain the envelopes of left and right hemispheres of the brain, but Lhemi and Rhemi are not keywords that are identified automatically by Brainstorm.
Try to display the surfaces by double-clicking on them, or with the popup menu Display.
In the Surfaces tab, the numbers of faces and vertices are displayed
What happend on the harddrive
If you are curious to know how all this anatomy information was stored in the database directory on the hard drive, you can have a look in your brainstorm_database/TutorialCTF directory.
