|
Size: 4446
Comment:
|
Size: 6290
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| <<TableOfContents(2)>> |
|
| Line 17: | Line 19: |
| 1. Right-click on Subject01, select ''Import MEG/EEG...'' <<BR>><<BR>>In the file format selection box, pick "''MEG/EEG: CTF .ds (*.meg4;*.res4)''". Go to TutorialCtf/data/ directory, select the the ''somMDYO-18av.ds'' directory and click on Open.<<BR>><<BR>> {{attachment:panelImportCtf.gif||height="258px",width="318px"}} | 1. Right-click on Subject01, select ''Import MEG/EEG...'' <<BR>><<BR>>In the file format selection box, pick "''MEG/EEG: CTF .ds (*.meg4;*.res4)''". Go to TutorialCtf/data/ directory, select the the ''somMDYO-18av.ds'' directory and click on Open.<<BR>><<BR>> {{attachment:panelImportCtf.gif||height="258",width="318"}} |
| Line 31: | Line 33: |
| === Display menu === The two menus in the ''Display ''menu display the same thing, but in a different way. You can add the scalp surface easily with the toolbar in the ''Surfaces ''tab, in the main window (''Add a surface'' button). <<BR>><<BR>> {{attachment:channelCtf.gif||height="169px",width="272px"}} {{attachment:channelMeg.gif||height="168px",width="271px"}} |
=== Menu: Display === The two menus in the ''Display ''menu display the same thing, but in a different way. You can add the scalp surface easily with the toolbar in the ''Surfaces ''tab, in the main window (''Add a surface'' button). |
| Line 34: | Line 36: |
| * '''CTF axial gradiometers''': Displays the CTF MEG sensors as they will be used for the forward model computation. The small squares do not have any reality, as CTF coils are circular, but they are modelized like that. Only the coil closest to the head is represented. The red arrows represent the orientation of the coils. This menu is only available for CTF recordings. Read more about [[MegSensorTypes|MEG sensor types]]. * '''MEG''': MEG coils are now represented as small white dots (center of the coils), and tesselated. |
{{attachment:channelCtf.gif||height="169",width="272"}} {{attachment:channelMeg.gif||height="168",width="271"}} * '''CTF axial gradiometers''' (only available for CTF recordings): * All the CTF MEG sensors axial gradiometers, with one coil close to the head, and one parallel and much farther. * This menu displays them as they will be used for the forward model computation. The small squares do not represent exactly the reality, as CTF coils are circular, but they are modelized like that. * Only the coil closest to the head is represented. The red arrows represent the orientation of the coils. Read more about [[MegSensorTypes|MEG sensor types]]. * '''MEG''': MEG sensors are now represented as small white dots (center of the coils close to the head), and tesselated. Red arrows still represent the orientation of the coils. === Menu: Edit channel file === Displays a table with almost all the information about the individual channels. You can use this window to view and edit the channels's properties. {{attachment:channelEdit.gif}} The channel file describe each channel separately, with the following information: * '''Name ''': Name that was given to the channel by the acquisition device. * '''Type ''': Channel type, eg. MEG, EEG, EOG, ECG, EMG, Stim, Other, etc. * Sometimes you will have to change the Type for some sensors; for instance if the EOG channel was saved as a regular EEG channel, you will have to change its type to prevent it from being used in the source reconstruction based on EEG. * '''Comment ''': Description of the channel. * '''Loc ''': Indicates the position in space of the sensor (x,y,z coordinates). One column per coil or per integration point; consult the pages about forward modeling theory if you want to know more about this. You should not modify these values from this interface. * '''Orient ''': Indicates the orientation in space of the sensor (x,y,z coordinates). One column per coil or per integration point. * '''Weight ''': When there are more than one coil or integration point, the Weight field indicates the multiplication factor to apply to each of these points. === Menu: File === Some other fields are present in the channel file that cannot be accessed with the ''Channel editor'' window. You can explore those other fields with the ''File ''menu, selecting ''View .mat files'' or ''Export to Matlab''. As we saw in previous tutorial. {{attachment:channelViewMat.gif}} |
Tutorial: Import and review recordings
Contents
Dataset description
You should already have downloaded this package if you have done correctly the previous tutorial. If it is not the case, go back to the ?previous tutorial and follow all the instructions.
File: |
bst_tutorial_somatotopy_ctf.zip |
Acquisition system: |
CTF MEG, 151 axial gradiometers, La Salpetriere Hospital, Paris |
Protocol: |
Shuffled electrical stimulations of the thumb fingers from both hands. The idea is to get a map of the primary sensory response on the cortex |
Author: |
Data provided courtesy of Sabine Meunier |
Anatomy directory: |
- T1-MRI of the subject in CTF format (.mri) |
Datasets directory: |
- somMGYO-18av.ds: average response for the stimulation of the right thumb (one subject, 400 trials) |
Observations: |
Stimulus occurs at time 0. There's a first tiny wave occuring at about 20ms or so but it's not too clear on all fingers. So if you are to compute cortical maps, start by the 40ms peak which is also of interest and which has much better SNR. |
Import recordings
Go to the Functional data (sorted by subject) view of the Brainstorm database.
Right-click on Subject01, select Import MEG/EEG...
In the file format selection box, pick "MEG/EEG: CTF .ds (*.meg4;*.res4)". Go to TutorialCtf/data/ directory, select the the somMDYO-18av.ds directory and click on Open.
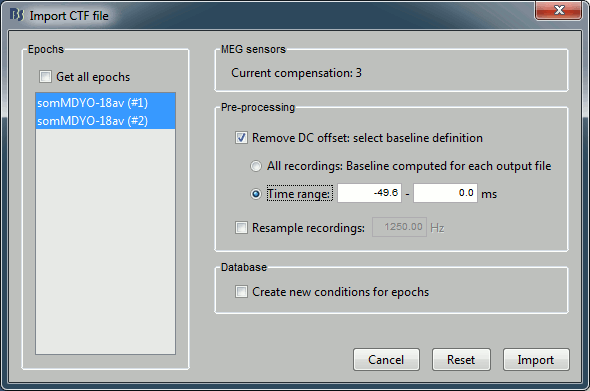
Two blocks of data are available in this file. They are referred in CTF files as Trials, even if they are the results of the averaging of trials from two different conditions.
You can select individual trials if you uncheck the "Use all trials" option.
- You can see that each block of recordings is defined for the following time range: [-49.60ms, 250.40ms]
Check the "Remove DC offset" option, together with "Base on pretrigger". For each sensor, this will compute the average value across the time (on the for the pre-trigger period, ie. where times < 0), and substract this mean to all the time values.
=> In MEG, this operation is always needed, unless it was already applied during one of the pre-processing steps.- Click on Import.
A condition was created, called after the filename. It contains three items:
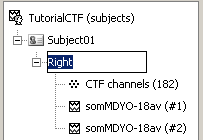
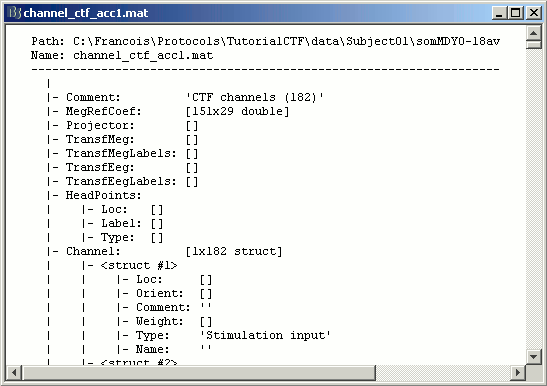
The first is the channel file: description of the positions, names, types, and various properties of the sensors that recorded the data. The value (182) means that there are 182 channels of data in the data files. They are not necessarily MEG channels, it may also includes EEG, EOG, stimulation lines, references, etc.
- The last two represent recording files
Rename the condition into StimRightThumb (F2, or successive left clicks, or right-click>Rename)
Display channel file
Let's explore what you can do with the first file. Right-click on the CTF and try all the menus.
Menu: Display
The two menus in the Display menu display the same thing, but in a different way. You can add the scalp surface easily with the toolbar in the Surfaces tab, in the main window (Add a surface button).
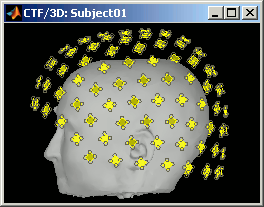
CTF axial gradiometers (only available for CTF recordings):
- All the CTF MEG sensors axial gradiometers, with one coil close to the head, and one parallel and much farther.
- This menu displays them as they will be used for the forward model computation. The small squares do not represent exactly the reality, as CTF coils are circular, but they are modelized like that.
Only the coil closest to the head is represented. The red arrows represent the orientation of the coils. Read more about ?MEG sensor types.
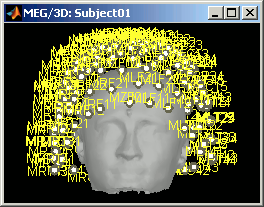
MEG: MEG sensors are now represented as small white dots (center of the coils close to the head), and tesselated. Red arrows still represent the orientation of the coils.
Menu: Edit channel file
Displays a table with almost all the information about the individual channels. You can use this window to view and edit the channels's properties.
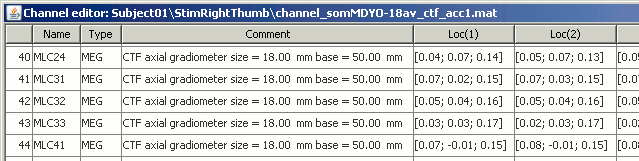
The channel file describe each channel separately, with the following information:
Name : Name that was given to the channel by the acquisition device.
Type : Channel type, eg. MEG, EEG, EOG, ECG, EMG, Stim, Other, etc.
- Sometimes you will have to change the Type for some sensors; for instance if the EOG channel was saved as a regular EEG channel, you will have to change its type to prevent it from being used in the source reconstruction based on EEG.
Comment : Description of the channel.
Loc : Indicates the position in space of the sensor (x,y,z coordinates). One column per coil or per integration point; consult the pages about forward modeling theory if you want to know more about this. You should not modify these values from this interface.
Orient : Indicates the orientation in space of the sensor (x,y,z coordinates). One column per coil or per integration point.
Weight : When there are more than one coil or integration point, the Weight field indicates the multiplication factor to apply to each of these points.
Menu: File
Some other fields are present in the channel file that cannot be accessed with the Channel editor window. You can explore those other fields with the File menu, selecting View .mat files or Export to Matlab. As we saw in previous tutorial.