|
Size: 85
Comment:
|
Size: 26887
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Writing your own tutorials = User-defined processes in ~user/.brainstorm/process |
= How to write your own process = Brainstorm offers a flexible plug-in structure. All the operations available when using the Process1 and Process2 tabs, which means most of the Brainstorm features, are in fact written as plug-ins. If you are interested in running your own code from the Brainstorm interface and benefit from the powerful database and visualization systems, the best option is probably for you to create your own process functions. It can take some time to get used to this logic but it is time well invested: you will be able to exchange code easily with your collaborators and the methods you develop could immediately reach thousands of users. Once your functions are stable, we can integrate them in the main Brainstorm distribution and maintain the code for you to ensure it stays compatible with the future releases of the software. This tutorial looks long and complicated, but don't let it scare you. Putting your code in a process is not so difficult. Most of it is a reference manual that details all the possible options, you don't need to understand it completely. The last part explains how to copy an existing process and modify it to do what you want. {{attachment:introPipeline.gif||height="283",width="489"}} == Alternative == Before you start reading this long technical document, you should be aware of a simpler solution to execute your own code from the Brainstorm interface. The process "Pre-process > '''Run Matlab command'''" is simple but very powerful. It loads the files in input and run them through a piece of Matlab code that you can edit freely. It can extend a lot the flexibility of the Brainstorm pipeline manager, providing an easy access to any Matlab function or script. {{attachment:runMatlab.gif}} == Process folders == A Brainstorm plug-in, or "process", is a single Matlab .m script that is automatically identified and added to the menus in the pipeline editor. Two folders are parsed for plug-ins: * '''brainstorm3/toolbox/process/functions''':<<BR>>Brainstorm "official" processes, included in the main distribution of the software * '''$HOME/.brainstorm/process''':<<BR>>User folder, to develop new processes or overwrite Brainstorm functions If you write a valid process function, name the script "process_...m" and place it in one of those folders, it will become automatically available in the pipeline editor menus, when you use the Process1 or Process2 tabs. Send it to another Brainstorm user and your code will be automatically available into the other person's Brainstorm interface. It is a very efficient solution for exchanging code without the nightmare of understanding what are the inputs of the functions (units of the values, dimensions of the matrices, etc.) == Structure of the process scripts == === Sub-functions === A process function must be named "process_...m" and located in one of the two process folders in order to be recognized by the software. Let's call our example function "process_test.m". It contains at least 4 functions: * '''process_test'''(): The first line of the script must contain a function with the same name as the .m script. It contains only a call to the Brainstorm script macro_methodcall. This allows to call sub-functions of the process_test.m script from other scripts, using the syntax: process_test('!FunctionName', arguments) * '''!GetDescription'''(): Returns a structure that describes the process: name, category, accepted inputs, options, etc. This function is called when Brainstorm parses the process folders to find all the valid processes. It informs the pipeline editor on how the process has to be integrated in the interface. * '''!FormatComment'''(): Returns a string that identifies the process in the interface. In the pipeline editor window, when the process is selected or when its options are modified, this function is called to update the process description line. Most processes would return simply the field sProcess.Comment, some other would add some options in the description (example: Pre-process > Band-pass filter, or Average > Average files). * '''Run'''(): Function called when the process is executed, either from the interface (after clicking on the Run button of the pipeline editor) or from a Matlab script (call to bst_process('!CallProcess', 'process_test', ...)). While the three first functions are descriptive, this one really does something. It receives the files placed in the Process1 or Process2 boxes, does its job and returns the output of the computation to Brainstorm. You are free to add as many sub-functions as needed to the process file. If your process needs some sub-functions to run, it is preferable to copy the full code directly into the "process_test.m" code, rather than leaving it in separate functions. This way it prevents from spreading sub-functions everywhere, which are later lost or forgotten in the distribution when the process is deleted. It might be uncomfortable at the beginning if you are not used to work with scripts with over 100 lines, but you'll get used to it, the Matlab code editor offers many solutions to make long scripts easy to edit (cells, code folding...). It makes your process easier to maintain and to exchange with other users, which is important in the long run. === Optional function: Compute() === A process can be designed to be called at the same time: * from the Brainstorm context, to work as a plug-in, and * directly from the Matlab command line or from a script, independently from the Brainstorm database and plug-in system. In this case, we can leave what is specific to the Brainstorm structure in the Run() function, and move the real computation to additional sub-functions. In this case, we recommend that you respect the following convention: name the main external sub-function '''Compute'''(). The following example will help clarifying this concept. === Example: Z-score === Let's take the example of the process "Standardize > Z-score (static)", which is described in the function '''process_zscore.m'''. The function '''Run()''' reads and tests the options defined by the user and then calls Compute(), which is responsible for calculating the Z-score normalization. {{{ function sInput = Run(sProcess, sInput) % Get inputs iBaseline = panel_time('GetTimeIndices', [...]); [...] % Compute zscore sInput.A = Compute(sInput.A, iBaseline); [...] end }}} The function '''Compute'''() calls another function '''!ComputeStat'''(): {{{ function A = Compute(A, iBaseline) % Calculate mean and standard deviation [meanBaseline, stdBaseline] = ComputeStat(A(:, iBaseline,:)); % Compute zscore A = bst_bsxfun(@minus, A, meanBaseline); A = bst_bsxfun(@rdivide, A, stdBaseline); end }}} {{{ function [meanBaseline, stdBaseline] = ComputeStat(A) % Compute baseline statistics stdBaseline = std(A, 0, 2); meanBaseline = mean(A, 2); % Remove null variance values stdBaseline(stdBaseline == 0) = 1e-12; end }}} This mechanism allows us to access this Z-score function at different levels. We can call it as a Brainstorm process that takes Brainstorm structures in input (this is usually not done manually): {{{ sInput = process_zscore('Run', sProcess, sInput); }}} Or as regular functions that takes standard Matlab matrices in input: {{{ % Generate some random signal F = rand(1,500); ind = 1:100; % Normalize the signal F = process_zscore('Compute', F, ind); % Or just calculate its average and standard deviation [Favg, Fstd] = process_zscore('ComputeStat', F); }}} == Process description == The function !GetDescription() creates a structure sProcess that documents the process: its name, the way it is supposed to be used in the interface and all the options it needs. It contains the following fields: * '''Comment''': String, represents the process in the "Add process" menus of the pipeline editor window. * '''!FileTag''': String that is added to the description of the output files, in the case of "Filter" processes. In the example of the Z-score process, !FileTag='| zscore'. If you apply the Z-score process on a file named "MN: MEG", the file created by the process is named "MN: MEG | zscore". This file tag is also added to the file name. * '''Category''': String that defines how the process is supposed to behave.<<BR>>The possible values are defined in the next section: 'Filter', 'File', 'Custom'. * '''!SubGroup''': Sub-menu in which you want the process to appear in the menus of the pipeline editor. It can be an existing category (eg. 'Pre-processing', 'Standardize', etc) or a new category. * '''Index''': Integer value that indicates a relative position in the "Add process" menus. For example, if your process sets Index=411, it would be displayed after the Z-score process (Index=410). Two processes can have the same index, in this case the one displayed first is the one that is read first in the process folder. If you set the Index to zero, it would be ignored and not displayed in the menus of the pipeline editor. * '''isSeparator''': Display a separator bar after the process in the pipeline editor menus. * '''!InputTypes''': Cell array of strings that represents the possible input types ('raw', 'data', 'results', 'timefreq', 'matrix'). This information is used to determine if a process is available in a specific interface configuration. For example: In the Process1 tab, if the "Process sources" button is selected, only the processes that have 'results' in their list !InputTypes, will be marked as available. All the others will be greyed out. * '''!OutputTypes''': Cell array of strings with the same dimension as !InputTypes. It defines, for each input type, what is the type of the files in output. For example: a process that has !InputTypes={'data','results'} and !OutputTypes={'timefreq','timefreq'} transforms recordings in time-frequency objects, and sources in time-frequency objects. Now if !OutputTypes={'data','results'}, the type of the new files is the same as the one from the input file. * '''nInputs''': Integer, defines the number of inputs of the process. If nInputs=1, the process appears in the Process1 tab. If nInputs=2, the process appears in the Process2 tab. * '''nMinFiles''': Integer, minimum number of files required by the process to run (0, 1, 2 or more). * '''processDim''': For the "Filter" processes, defines along which dimensions the process is allowed to split the input data matrix while processing it, if it's too big to be processed at once. Possible values: * 1: Split in blocks of signals (example: Band-pass filter or Sinusoid removal) * 2: Split in time blocks (example: EEG average reference or Apply SSP) * Empty: Does not allow the process to split the input data matrix (default) * '''isSourceAbsolute''': For the processes that accept source maps in input (type 'results'), this option defines if we want to process the real source values or their absolute values. Possible values: * -1: Never process the absolute values of the sources * 0: Offer it as an option, but disabled by default * 1: Offer it as an option, and enables it by default * 2: Always process the absolute values of the sources * '''isPaired''': Applies only to the processes with two inputs (Process2), defines if the process needs pairs of files in input. If isPaired=1, the first file in FilesA and the first file in FilesB are processed together, the second files are processed together and so on. Example: paired t-test. * '''options''': List of options that are offered to the user in the pipeline editor window. This variable is a structure, where each field represents an option. Not all the fields have to be defined in the function !GetDescription(). The missing ones will be set to their default values, as defined in db_template('!ProcessDesc'). == Definition of the options == === Options structure === The field sProcess.options describes the list of options that are displayed in the pipeline editor window when the process is selected. It is a structure with one field per option. If we have an option named "overwrite", it is described in the structure sProcess.options.overwrite. Every option is a structure with the following fields: * '''Type''': String, defines the type of option (checkbox, text field, etc.) * '''Comment''': String, describes the option in the pipeline editor window * '''Value''': Default value for this option. The type of this variable depends of the "Type" field * '''!InputTypes''': [optional] Cell array of the types of input files for which the option is shown * '''Hidden''': [optional] If set to 1, the option is not displayed in the pipeline editor, but passed to the Run function in the sProcess structure. It can be a way to pass additional parameters to the Run function without overloading the user interface. === Examples === Example of two options defined in process_zscore.m: {{{ % === Baseline time window sProcess.options.baseline.Comment = 'Baseline:'; sProcess.options.baseline.Type = 'baseline'; sProcess.options.baseline.Value = []; % === Sensor types sProcess.options.sensortypes.Comment = 'Sensor types or names (empty=all): '; sProcess.options.sensortypes.Type = 'text'; sProcess.options.sensortypes.Value = 'MEG, EEG'; sProcess.options.sensortypes.InputTypes = {'data'}; }}} === User preferences === Note that the default values defined in sProcess.options are usually displayed only once. When the user modifies the option, the new value is saved in the user preferences and offered as the default the next time the process is selected in the pipeline editor. If you modify the Value field in your process function, the default offered when you select the process in the pipeline editor may not change accordingly. This means that another default has been saved in the user preferences. To reset all the options to their real default values (as defined in the process functions), you can use the menu '''Reset options''' in the Pipeline menu of pipeline editor window. === Option types === * ''''checkbox'''': Simple check box, to enable or disable something * Comment: String, displayed next to the checkbox * Value: 0 (not checked) or 1 (checked) * Example: process_average * ''''radio'''': List of radio buttons, to select between multiple choices * Comment: Cell array of strings, each string is a possible choice * Value: Integer, index of the selected entry * Example: process_average * ''''combobox'''': Drop-down list, to select between multiple choice * Comment: String displayed before the drop-down list * Value: {iSelected, {'entry1', 'entry2', ...}} (iSelected is the index of the selected entry) * Example: process_headmodel * ''''text'''': Simple text field * Comment: String displayed before the text field * Value: String * Example: process_add_tag * ''''textarea'''': Multi-line text editor * Comment: String displayed above the text area * Value: String * Example: process_matlab_eval * ''''value'''': Text field to edit numerical values with fixed precision * Comment: String displayed before the text field * Value: {value, units, precision} * value: Numerical value entered by the user * units: String that represents the units, displayed after the field * precision: Number of decimals after the point (0=integer) * Example: process_bandpass * ''''range'''': Two text fields to enter an interval, [start, stop] * Comment: String displayed before the text fields * Value: {[start,stop], units, precision} * [start,stop]: Two numerical values entered by the user * units: String that represents the units, displayed after the two fields * precision: Number of decimals after the point (0=integer) * Example: process_evt_detect * ''''timewindow'''': Similar to 'range', but offers by default the time range of the first file to process * Example: process_average_time * ''''baseline'''': Same as 'timewindow', but offers by default the time segment before 0 * If there are no negative times, uses the full time range * ''''poststim'''': Same as 'timewindow', but offers by default the time segment after 0 * If there are no negative times, uses the full time range * ''''label'''': Simple text label, no user input * Comment: String, accepts HTML input * Value: Ignored * Example: process_average * ''''filename'''': Select a file or a folder (text box + button "...") * Comment: String displayed before the text box * Value: !SelectOptions cell array, see details in the code of the example process * Example: process_evt_import * ''''datafile'''': Same as 'filename', but specific to MEG/EEG files that have to be imported or linked in the database * ''''channelname'''': Drop-down list to select a channel, can be edited directly by typing the channel name * Comment: String displayed before the drop-down list * Value: String, name of the selected channel * Example: process_evt_detect * ''''subjectname'''': Drop-down list to select a subject, can be edited directly by typing the subject name * Comment: String displayed before the drop-down list * Value: String, name of the selected subject * Example: process_import_data_raw * ''''groupbands'''': Edit a list of time or frequency bands with a text editor * Comment: String displayed above the text area * Value: Cell array that describes the frequency band, one row per band:<<BR>>{'band_name', 'band_range', 'band_function'}<<BR>>The default frequency bands can be obtained with: bst_get('DefaultFreqBands') * Example: process_tf_bands * ''''cluster'''': Select a set of clusters or scouts in a list * Comment: Ignored * Value: Array of structures representing the scouts selected by the user * Example: process_extract_cluster * ''''cluster_confirm'''': Same as 'cluster', but with an additional checkbox on the top. If the checkbox is not selected by the user, the cluster/scout lists is greyed out and the returned Value will be []. * Example: process_fft * ''''atlas'''': Select from a drop-down list an atlas available in the the first input source file. The atlas selected by default in the list is the last one that one selected in the Scout tab with displaying the source file. * Comment: String displayed before the drop-down list * Value: String, name of the atlas selected by the user * Example: process_source_atlas * ''''editpref'''': Show a button 'Edit' that opens a user-defined option panel * Comment: {'panel_function_name', 'Comment'} * Value: Structure returned by the function panel_function_name>!GetPanelContents() * Example: process_hilbert / panel_timefreq_options == Categories of process == There are three different types of processes: '''Filter''', '''File''', '''Custom'''. The category of the process is defined by the field sProcess.Category. For the processes with two sets of inputs files (Process2), the logic is the same but the category are called: '''Filter2''', '''File2''', '''Custom'''. === Category: 'Filter' and 'Filter2' === Brainstorm processes independently each file in the input list (the files that have been dropped in the Process1 or Process2 files lists) and is responsible for the following operations: * reading the input files, * possibly splitting them if they are too big, * writing the output files on the hard drive, * referencing them in the database. In the process, the function Run(): * receives the data matrix to process, one file at a time, * applies some operation on it, * returns the processed values. Advantages: All the complicated things are taken care of automatically, the functions can be very short. Limitations: There is no control over the file names and locations, one file in input = one file in output, and the file type cannot be changed (!InputTypes=!OutputTypes). For example, let's consider one of the simplest processes: process_absolute.m. It just calculates the absolute value of the input data matrix. The Run() function is only one line long: {{{ function sInput = Run(sProcess, sInput) sInput.A = abs(sInput.A); end }}} The sInput structure gives lots of information about the input file coming from the database, and one additional fields "A" that contains the block of data to process. This process just applies the function abs() to the data sInput.A and returns modified values. A new file is created by Brainstorm in the database to store this result. === Category: 'File' and 'File2' === Brainstorm processes independently each file in the input list. It creates a structure sInput that documents the input file but does not load the data in the "A" field, as in the Filter case. In the process, the function Run() is called once for each input file and is responsible for: * reading the input file, * processing it, * saving the results in a new file on the hard drive, [optional] * referencing the new file in the database. [optional] * return the path of the new file The resulting functions are much longer, but this time the process is free do anything, there are no restrictions. The outline of the typical Run() function can be described as following: {{{ function OutputFile = Run(sProcess, sInput) % Load input file DataMat = in_bst_data(sInput.FileName); % Apply some function to the data in DataMat OutputMat = some_function(DataMat); % Generate a new file name in the same folder OutputFile = bst_process('GetNewFilename', bst_fileparts(sStudy.FileName), fileType); % Save the new file save(OutputFile, '-struct', 'OutputMat'); % Reference OutputFile in the database: db_add_data(sInput.iStudy, OutputFile, OutputMat); end }}} === Category: 'Custom' === Similar to the previous case "File", but this time all the input files are passed at once to the process. The function Run() is called only once. It receives all the input file names in an array of structures "sInputs". From that, it can create zero, one or many files. The list of output files is returned in a cell array of strings "!OutputFiles". {{{ function OutputFiles = Run(sProcess, sInputs) % Load input files % Do something interesting % Save new files % Reference the new files in the database % Return all the new file names in the cell-array OutputFiles end }}} == Input description == The structure sInput contains the following fields: * '''iStudy''': Index of the study, get the structure with sStudy=bst_get('Study', sInput.iStudy) * '''iItem''': Index of file in its category (example for recordings: sStudy.Data(sInput.iItem) * '''!FileName''': Input file name (file path relative to the protocol data folder) * '''!FileType''': Input file type ('raw', 'data', 'results', 'timefreq', 'matrix') * '''Comment''': Comment field of the input file (what is displayed in the database explorer) * '''Condition''': Name of the condition/folder in which the file is located * '''!SubjectFile''': Relative path to the subject file (brainstormsubject.mat) * '''!SubjectName''': Name of the subject * '''!DataFile''': For types 'results' or 'timefreq', path of the parent file in the database explorer * '''!ChannelFile''': Relative path to the channel file * '''!ChannelTypes''': Cell array of channel types available for the input file * '''A''': Loaded data matrix, only in the case of a Filter process == Running a process == Three ways to run a process: * From the '''interface''': select the process in the pipeline editor window, then click on the Run button. * From a script generated by Brainstorm, using '''bst_process''': select the process in the pipeline editor window, select the menu "Generate .m script", then copy-paste the call to bst_process('!CallProcess', 'process_name', options). It can be executed from a script or directly from the Matlab command window * Directly calling a function within the process: with the syntax: "'''process_name'''('!FunctionName', arguments)" == Create your own process == The easiest way for you to write your own plug-in function is to start working from an existing example. There are over a hundred processes currently available in the main Brainstorm distribution. Take some time to find one that is close to what you are planning to do. By order of importance: same category (Filter/File/Custom), same types of input and output files, similar options, same logic. Then follow those guidelines: * Copy the original process_...m script to your $HOME/.brainstorm/process folder * Rename it so that it doesn't interfere with the existing functions: pick a name that represents what your function does and avoid using capital letters in the file name * Edit the function name in the first line of the process, so that it matches the script file name * Edit the comments in the header of the function * Edit the some fields in the function !GetDescription functions: * sProcess.Comment: String that represents the process in the pipeline editor menu * sProcess.!SubGroup and sProcess.Index, so that it appears where you want in the list * sProcess.options: Read the above documentation * Edit the !FormatComment() function if needed. If it contains some complicated code, just replace it with "Comment = sProcess.Comment;" * Edit the Run() function * Delete all the other sub-functions * Post on the Brainstorm forums all the questions you need * Your process should be automatically available in the menus in the pipeline editor window. It is not, it might be because there are some errors in it: check your Matlab command window. If you see a message "Invalid plug-in function", Matlab could not execute the code correctly, check the syntax. * For development/debugging purpose, remember that you don't have to use the Brainstorm interface to execute your code. It can be annoying to select hundreds of times per day your process from the pipeline editor. == Examples == Here are some additional sample processes that can help you for specific tasks: * Generate a head model / leadfield matrix: [[attachment:process_headmodel_test.m|process_headmodel_test]] * Generate an inverse model / source matrix: [[attachment:process_beamformer_test.m|process_beamformer_test]] * Load all the trials in input at once and process them: [[attachment:process_example_customavg.m|process_example_customavg]] == Feedback == <<EmbedContent(http://neuroimage.usc.edu/brainstorm3_register/get_feedback.php?Tutorials/TutUserProcess)>> |
How to write your own process
Brainstorm offers a flexible plug-in structure. All the operations available when using the Process1 and Process2 tabs, which means most of the Brainstorm features, are in fact written as plug-ins.
If you are interested in running your own code from the Brainstorm interface and benefit from the powerful database and visualization systems, the best option is probably for you to create your own process functions. It can take some time to get used to this logic but it is time well invested: you will be able to exchange code easily with your collaborators and the methods you develop could immediately reach thousands of users. Once your functions are stable, we can integrate them in the main Brainstorm distribution and maintain the code for you to ensure it stays compatible with the future releases of the software.
This tutorial looks long and complicated, but don't let it scare you. Putting your code in a process is not so difficult. Most of it is a reference manual that details all the possible options, you don't need to understand it completely. The last part explains how to copy an existing process and modify it to do what you want.
Alternative
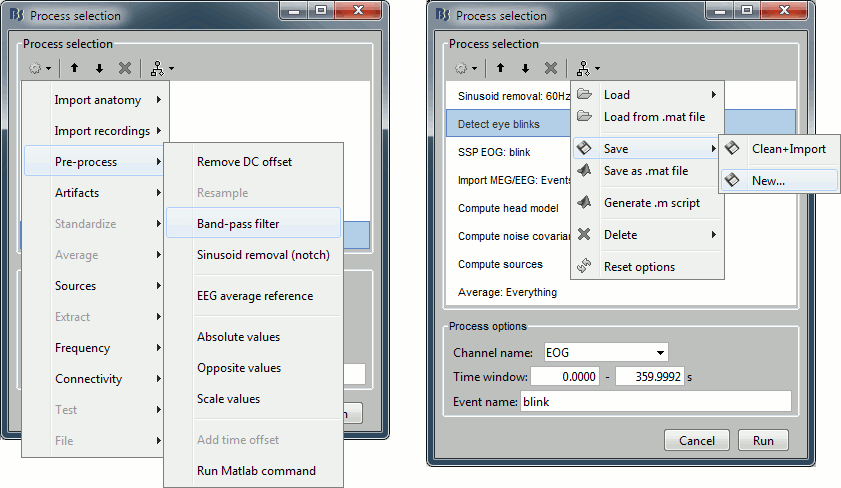
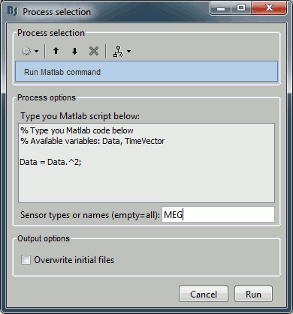
Before you start reading this long technical document, you should be aware of a simpler solution to execute your own code from the Brainstorm interface.
The process "Pre-process > Run Matlab command" is simple but very powerful. It loads the files in input and run them through a piece of Matlab code that you can edit freely. It can extend a lot the flexibility of the Brainstorm pipeline manager, providing an easy access to any Matlab function or script.
Process folders
A Brainstorm plug-in, or "process", is a single Matlab .m script that is automatically identified and added to the menus in the pipeline editor. Two folders are parsed for plug-ins:
brainstorm3/toolbox/process/functions:
Brainstorm "official" processes, included in the main distribution of the software$HOME/.brainstorm/process:
User folder, to develop new processes or overwrite Brainstorm functions
If you write a valid process function, name the script "process_...m" and place it in one of those folders, it will become automatically available in the pipeline editor menus, when you use the Process1 or Process2 tabs.
Send it to another Brainstorm user and your code will be automatically available into the other person's Brainstorm interface. It is a very efficient solution for exchanging code without the nightmare of understanding what are the inputs of the functions (units of the values, dimensions of the matrices, etc.)
Structure of the process scripts
Sub-functions
A process function must be named "process_...m" and located in one of the two process folders in order to be recognized by the software. Let's call our example function "process_test.m". It contains at least 4 functions:
process_test(): The first line of the script must contain a function with the same name as the .m script. It contains only a call to the Brainstorm script macro_methodcall. This allows to call sub-functions of the process_test.m script from other scripts, using the syntax: process_test('FunctionName', arguments)
GetDescription(): Returns a structure that describes the process: name, category, accepted inputs, options, etc. This function is called when Brainstorm parses the process folders to find all the valid processes. It informs the pipeline editor on how the process has to be integrated in the interface.
FormatComment(): Returns a string that identifies the process in the interface. In the pipeline editor window, when the process is selected or when its options are modified, this function is called to update the process description line. Most processes would return simply the field sProcess.Comment, some other would add some options in the description (example: Pre-process > Band-pass filter, or Average > Average files).
Run(): Function called when the process is executed, either from the interface (after clicking on the Run button of the pipeline editor) or from a Matlab script (call to bst_process('CallProcess', 'process_test', ...)). While the three first functions are descriptive, this one really does something. It receives the files placed in the Process1 or Process2 boxes, does its job and returns the output of the computation to Brainstorm.
You are free to add as many sub-functions as needed to the process file. If your process needs some sub-functions to run, it is preferable to copy the full code directly into the "process_test.m" code, rather than leaving it in separate functions. This way it prevents from spreading sub-functions everywhere, which are later lost or forgotten in the distribution when the process is deleted. It might be uncomfortable at the beginning if you are not used to work with scripts with over 100 lines, but you'll get used to it, the Matlab code editor offers many solutions to make long scripts easy to edit (cells, code folding...). It makes your process easier to maintain and to exchange with other users, which is important in the long run.
Optional function: Compute()
A process can be designed to be called at the same time:
- from the Brainstorm context, to work as a plug-in, and
- directly from the Matlab command line or from a script, independently from the Brainstorm database and plug-in system.
In this case, we can leave what is specific to the Brainstorm structure in the Run() function, and move the real computation to additional sub-functions. In this case, we recommend that you respect the following convention: name the main external sub-function Compute(). The following example will help clarifying this concept.
Example: Z-score
Let's take the example of the process "Standardize > Z-score (static)", which is described in the function process_zscore.m. The function Run() reads and tests the options defined by the user and then calls Compute(), which is responsible for calculating the Z-score normalization.
function sInput = Run(sProcess, sInput)
% Get inputs
iBaseline = panel_time('GetTimeIndices', [...]);
[...]
% Compute zscore
sInput.A = Compute(sInput.A, iBaseline);
[...]
endThe function Compute() calls another function ComputeStat():
function A = Compute(A, iBaseline)
% Calculate mean and standard deviation
[meanBaseline, stdBaseline] = ComputeStat(A(:, iBaseline,:));
% Compute zscore
A = bst_bsxfun(@minus, A, meanBaseline);
A = bst_bsxfun(@rdivide, A, stdBaseline);
endfunction [meanBaseline, stdBaseline] = ComputeStat(A)
% Compute baseline statistics
stdBaseline = std(A, 0, 2);
meanBaseline = mean(A, 2);
% Remove null variance values
stdBaseline(stdBaseline == 0) = 1e-12;
endThis mechanism allows us to access this Z-score function at different levels. We can call it as a Brainstorm process that takes Brainstorm structures in input (this is usually not done manually):
sInput = process_zscore('Run', sProcess, sInput);Or as regular functions that takes standard Matlab matrices in input:
% Generate some random signal
F = rand(1,500); ind = 1:100;
% Normalize the signal
F = process_zscore('Compute', F, ind);
% Or just calculate its average and standard deviation
[Favg, Fstd] = process_zscore('ComputeStat', F);
Process description
The function GetDescription() creates a structure sProcess that documents the process: its name, the way it is supposed to be used in the interface and all the options it needs. It contains the following fields:
Comment: String, represents the process in the "Add process" menus of the pipeline editor window.
FileTag: String that is added to the description of the output files, in the case of "Filter" processes. In the example of the Z-score process, FileTag='| zscore'. If you apply the Z-score process on a file named "MN: MEG", the file created by the process is named "MN: MEG | zscore". This file tag is also added to the file name.
Category: String that defines how the process is supposed to behave.
The possible values are defined in the next section: 'Filter', 'File', 'Custom'.SubGroup: Sub-menu in which you want the process to appear in the menus of the pipeline editor. It can be an existing category (eg. 'Pre-processing', 'Standardize', etc) or a new category.
Index: Integer value that indicates a relative position in the "Add process" menus. For example, if your process sets Index=411, it would be displayed after the Z-score process (Index=410). Two processes can have the same index, in this case the one displayed first is the one that is read first in the process folder. If you set the Index to zero, it would be ignored and not displayed in the menus of the pipeline editor.
isSeparator: Display a separator bar after the process in the pipeline editor menus.
InputTypes: Cell array of strings that represents the possible input types ('raw', 'data', 'results', 'timefreq', 'matrix'). This information is used to determine if a process is available in a specific interface configuration. For example: In the Process1 tab, if the "Process sources" button is selected, only the processes that have 'results' in their list InputTypes, will be marked as available. All the others will be greyed out.
OutputTypes: Cell array of strings with the same dimension as InputTypes. It defines, for each input type, what is the type of the files in output. For example: a process that has InputTypes={'data','results'} and OutputTypes={'timefreq','timefreq'} transforms recordings in time-frequency objects, and sources in time-frequency objects. Now if OutputTypes={'data','results'}, the type of the new files is the same as the one from the input file.
nInputs: Integer, defines the number of inputs of the process. If nInputs=1, the process appears in the Process1 tab. If nInputs=2, the process appears in the Process2 tab.
nMinFiles: Integer, minimum number of files required by the process to run (0, 1, 2 or more).
processDim: For the "Filter" processes, defines along which dimensions the process is allowed to split the input data matrix while processing it, if it's too big to be processed at once. Possible values:
- 1: Split in blocks of signals (example: Band-pass filter or Sinusoid removal)
- 2: Split in time blocks (example: EEG average reference or Apply SSP)
- Empty: Does not allow the process to split the input data matrix (default)
isSourceAbsolute: For the processes that accept source maps in input (type 'results'), this option defines if we want to process the real source values or their absolute values. Possible values:
- -1: Never process the absolute values of the sources
- 0: Offer it as an option, but disabled by default
- 1: Offer it as an option, and enables it by default
- 2: Always process the absolute values of the sources
isPaired: Applies only to the processes with two inputs (Process2), defines if the process needs pairs of files in input. If isPaired=1, the first file in FilesA and the first file in FilesB are processed together, the second files are processed together and so on. Example: paired t-test.
options: List of options that are offered to the user in the pipeline editor window. This variable is a structure, where each field represents an option.
Not all the fields have to be defined in the function GetDescription(). The missing ones will be set to their default values, as defined in db_template('ProcessDesc').
Definition of the options
Options structure
The field sProcess.options describes the list of options that are displayed in the pipeline editor window when the process is selected. It is a structure with one field per option. If we have an option named "overwrite", it is described in the structure sProcess.options.overwrite. Every option is a structure with the following fields:
Type: String, defines the type of option (checkbox, text field, etc.)
Comment: String, describes the option in the pipeline editor window
Value: Default value for this option. The type of this variable depends of the "Type" field
InputTypes: [optional] Cell array of the types of input files for which the option is shown
Hidden: [optional] If set to 1, the option is not displayed in the pipeline editor, but passed to the Run function in the sProcess structure. It can be a way to pass additional parameters to the Run function without overloading the user interface.
Examples
Example of two options defined in process_zscore.m:
% === Baseline time window
sProcess.options.baseline.Comment = 'Baseline:';
sProcess.options.baseline.Type = 'baseline';
sProcess.options.baseline.Value = [];
% === Sensor types
sProcess.options.sensortypes.Comment = 'Sensor types or names (empty=all): ';
sProcess.options.sensortypes.Type = 'text';
sProcess.options.sensortypes.Value = 'MEG, EEG';
sProcess.options.sensortypes.InputTypes = {'data'};
User preferences
Note that the default values defined in sProcess.options are usually displayed only once. When the user modifies the option, the new value is saved in the user preferences and offered as the default the next time the process is selected in the pipeline editor.
If you modify the Value field in your process function, the default offered when you select the process in the pipeline editor may not change accordingly. This means that another default has been saved in the user preferences. To reset all the options to their real default values (as defined in the process functions), you can use the menu Reset options in the Pipeline menu of pipeline editor window.
Option types
'checkbox': Simple check box, to enable or disable something
- Comment: String, displayed next to the checkbox
- Value: 0 (not checked) or 1 (checked)
- Example: process_average
'radio': List of radio buttons, to select between multiple choices
- Comment: Cell array of strings, each string is a possible choice
- Value: Integer, index of the selected entry
- Example: process_average
'combobox': Drop-down list, to select between multiple choice
- Comment: String displayed before the drop-down list
- Value: {iSelected, {'entry1', 'entry2', ...}} (iSelected is the index of the selected entry)
- Example: process_headmodel
'text': Simple text field
- Comment: String displayed before the text field
- Value: String
- Example: process_add_tag
'textarea': Multi-line text editor
- Comment: String displayed above the text area
- Value: String
- Example: process_matlab_eval
'value': Text field to edit numerical values with fixed precision
- Comment: String displayed before the text field
- Value: {value, units, precision}
- value: Numerical value entered by the user
- units: String that represents the units, displayed after the field
- precision: Number of decimals after the point (0=integer)
- Example: process_bandpass
'range': Two text fields to enter an interval, [start, stop]
- Comment: String displayed before the text fields
- Value: {[start,stop], units, precision}
- [start,stop]: Two numerical values entered by the user
- units: String that represents the units, displayed after the two fields
- precision: Number of decimals after the point (0=integer)
- Example: process_evt_detect
'timewindow': Similar to 'range', but offers by default the time range of the first file to process
- Example: process_average_time
'baseline': Same as 'timewindow', but offers by default the time segment before 0
- If there are no negative times, uses the full time range
'poststim': Same as 'timewindow', but offers by default the time segment after 0
- If there are no negative times, uses the full time range
'label': Simple text label, no user input
- Comment: String, accepts HTML input
- Value: Ignored
- Example: process_average
'filename': Select a file or a folder (text box + button "...")
- Comment: String displayed before the text box
Value: SelectOptions cell array, see details in the code of the example process
- Example: process_evt_import
'datafile': Same as 'filename', but specific to MEG/EEG files that have to be imported or linked in the database
'channelname': Drop-down list to select a channel, can be edited directly by typing the channel name
- Comment: String displayed before the drop-down list
- Value: String, name of the selected channel
- Example: process_evt_detect
'subjectname': Drop-down list to select a subject, can be edited directly by typing the subject name
- Comment: String displayed before the drop-down list
- Value: String, name of the selected subject
- Example: process_import_data_raw
'groupbands': Edit a list of time or frequency bands with a text editor
- Comment: String displayed above the text area
Value: Cell array that describes the frequency band, one row per band:
{'band_name', 'band_range', 'band_function'}
The default frequency bands can be obtained with: bst_get('DefaultFreqBands')- Example: process_tf_bands
'cluster': Select a set of clusters or scouts in a list
- Comment: Ignored
- Value: Array of structures representing the scouts selected by the user
- Example: process_extract_cluster
'cluster_confirm': Same as 'cluster', but with an additional checkbox on the top. If the checkbox is not selected by the user, the cluster/scout lists is greyed out and the returned Value will be [].
- Example: process_fft
'atlas': Select from a drop-down list an atlas available in the the first input source file. The atlas selected by default in the list is the last one that one selected in the Scout tab with displaying the source file.
- Comment: String displayed before the drop-down list
- Value: String, name of the atlas selected by the user
- Example: process_source_atlas
'editpref': Show a button 'Edit' that opens a user-defined option panel
- Comment: {'panel_function_name', 'Comment'}
Value: Structure returned by the function panel_function_name>GetPanelContents()
- Example: process_hilbert / panel_timefreq_options
Categories of process
There are three different types of processes: Filter, File, Custom. The category of the process is defined by the field sProcess.Category. For the processes with two sets of inputs files (Process2), the logic is the same but the category are called: Filter2, File2, Custom.
Category: 'Filter' and 'Filter2'
Brainstorm processes independently each file in the input list (the files that have been dropped in the Process1 or Process2 files lists) and is responsible for the following operations:
- reading the input files,
- possibly splitting them if they are too big,
- writing the output files on the hard drive,
- referencing them in the database.
In the process, the function Run():
- receives the data matrix to process, one file at a time,
- applies some operation on it,
- returns the processed values.
Advantages: All the complicated things are taken care of automatically, the functions can be very short.
Limitations: There is no control over the file names and locations, one file in input = one file in output, and the file type cannot be changed (InputTypes=OutputTypes).
For example, let's consider one of the simplest processes: process_absolute.m. It just calculates the absolute value of the input data matrix. The Run() function is only one line long:
function sInput = Run(sProcess, sInput)
sInput.A = abs(sInput.A);
endThe sInput structure gives lots of information about the input file coming from the database, and one additional fields "A" that contains the block of data to process. This process just applies the function abs() to the data sInput.A and returns modified values. A new file is created by Brainstorm in the database to store this result.
Category: 'File' and 'File2'
Brainstorm processes independently each file in the input list. It creates a structure sInput that documents the input file but does not load the data in the "A" field, as in the Filter case.
In the process, the function Run() is called once for each input file and is responsible for:
- reading the input file,
- processing it,
- saving the results in a new file on the hard drive, [optional]
- referencing the new file in the database. [optional]
- return the path of the new file
The resulting functions are much longer, but this time the process is free do anything, there are no restrictions. The outline of the typical Run() function can be described as following:
function OutputFile = Run(sProcess, sInput)
% Load input file
DataMat = in_bst_data(sInput.FileName);
% Apply some function to the data in DataMat
OutputMat = some_function(DataMat);
% Generate a new file name in the same folder
OutputFile = bst_process('GetNewFilename', bst_fileparts(sStudy.FileName), fileType);
% Save the new file
save(OutputFile, '-struct', 'OutputMat');
% Reference OutputFile in the database:
db_add_data(sInput.iStudy, OutputFile, OutputMat);
end
Category: 'Custom'
Similar to the previous case "File", but this time all the input files are passed at once to the process.
The function Run() is called only once. It receives all the input file names in an array of structures "sInputs". From that, it can create zero, one or many files. The list of output files is returned in a cell array of strings "OutputFiles".
function OutputFiles = Run(sProcess, sInputs)
% Load input files
% Do something interesting
% Save new files
% Reference the new files in the database
% Return all the new file names in the cell-array OutputFiles
end
Input description
The structure sInput contains the following fields:
iStudy: Index of the study, get the structure with sStudy=bst_get('Study', sInput.iStudy)
iItem: Index of file in its category (example for recordings: sStudy.Data(sInput.iItem)
FileName: Input file name (file path relative to the protocol data folder)
FileType: Input file type ('raw', 'data', 'results', 'timefreq', 'matrix')
Comment: Comment field of the input file (what is displayed in the database explorer)
Condition: Name of the condition/folder in which the file is located
SubjectFile: Relative path to the subject file (brainstormsubject.mat)
SubjectName: Name of the subject
DataFile: For types 'results' or 'timefreq', path of the parent file in the database explorer
ChannelFile: Relative path to the channel file
ChannelTypes: Cell array of channel types available for the input file
A: Loaded data matrix, only in the case of a Filter process
Running a process
Three ways to run a process:
From the interface: select the process in the pipeline editor window, then click on the Run button.
From a script generated by Brainstorm, using bst_process: select the process in the pipeline editor window, select the menu "Generate .m script", then copy-paste the call to bst_process('CallProcess', 'process_name', options). It can be executed from a script or directly from the Matlab command window
Directly calling a function within the process: with the syntax: "process_name('FunctionName', arguments)"
Create your own process
The easiest way for you to write your own plug-in function is to start working from an existing example. There are over a hundred processes currently available in the main Brainstorm distribution. Take some time to find one that is close to what you are planning to do. By order of importance: same category (Filter/File/Custom), same types of input and output files, similar options, same logic. Then follow those guidelines:
- Copy the original process_...m script to your $HOME/.brainstorm/process folder
- Rename it so that it doesn't interfere with the existing functions: pick a name that represents what your function does and avoid using capital letters in the file name
- Edit the function name in the first line of the process, so that it matches the script file name
- Edit the comments in the header of the function
Edit the some fields in the function GetDescription functions:
- sProcess.Comment: String that represents the process in the pipeline editor menu
sProcess.SubGroup and sProcess.Index, so that it appears where you want in the list
- sProcess.options: Read the above documentation
Edit the FormatComment() function if needed. If it contains some complicated code, just replace it with "Comment = sProcess.Comment;"
- Edit the Run() function
- Delete all the other sub-functions
- Post on the Brainstorm forums all the questions you need
- Your process should be automatically available in the menus in the pipeline editor window. It is not, it might be because there are some errors in it: check your Matlab command window. If you see a message "Invalid plug-in function", Matlab could not execute the code correctly, check the syntax.
- For development/debugging purpose, remember that you don't have to use the Brainstorm interface to execute your code. It can be annoying to select hundreds of times per day your process from the pipeline editor.
Examples
Here are some additional sample processes that can help you for specific tasks:
Generate a head model / leadfield matrix: process_headmodel_test
Generate an inverse model / source matrix: process_beamformer_test
Load all the trials in input at once and process them: process_example_customavg
Feedback
