|
Size: 5389
Comment:
|
Size: 6820
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| The default approach for the source estimation in Brainstorm is to limit the source space to the cortex surface. This choice is motivated by the assumption that most of the activity we record in MEG and EEG comes from the cerebral cortex. Constraining the source reconstruction to a surface works well when this assumption is verified, and the results we obtain are much easier to review that a full volume. | The default approach for the source estimation in Brainstorm is to limit the source space to the cortex surface. This choice is motivated by the assumption that most of the activity we record in MEG and EEG comes from the cerebral cortex. Constraining the source reconstruction to a surface works well when this assumption is verified, and the results we obtain are much easier to review than a full volume. |
| Line 13: | Line 13: |
| * Select the protocol TutorialIntroduction . Go to the view "Functional data (sorted by subjects)". * In the imported Run01, right-click on the channel file > '''Compute head model '''> '''MRI volume'''. <<BR>>The various options available a described below.<<BR>><<BR>> {{attachment:gridOptions.gif||height="420",width="673"}} |
* Select the protocol TutorialIntroduction. Go to the view "Functional data (sorted by subjects)". * In the imported Run01, right-click on the channel file > '''Compute head model '''> '''MRI volume'''. <<BR>><<BR>> {{attachment:gridOptions.gif||height="420",width="673"}} |
| Line 16: | Line 16: |
| * '''Generate from cortex surface''': Creates a grid that samples the head model in an adaptive way, denser at the surface of the brain (it is closer to the sensors therefore the expected spatial resolution of the MEG/EEG source estimation is higher), and sparser at the center of the brain. This grid is created with following algorithm: | * '''Generate from cortex surface''': Creates a grid that samples the brain volume in an adaptive way, denser at the surface (we expect a higher spatial resolution close to the sensors), and sparser at the center of the brain. This grid is created with following algorithm: |
| Line 21: | Line 21: |
| * '''Regular grid''': Creates a regular grid with one grid point every few millimeters (option Grid resolution). This grid is isotropic grid, ie. all the points are equally spaced. | * '''Regular grid''': Creates a regular grid with one grid point every few millimeters (option Grid resolution) in the three directions (x,y,z). |
| Line 23: | Line 23: |
| * '''Load from a file/variable''': If you already have a grid of points you would like to use, you can import it directly from a file or from a Matlab variable (only one field, [N,,points,,x3] values). | * '''Load from a file/variable''': If you already have a grid of points you would like to use, you can import it from a file or a Matlab variable (only one field, [N,,points,,x3] values). |
| Line 25: | Line 25: |
| * '''Use template grid''': This option is only available when a template source grid has been defined on the default anatomy. For more information, see tutorial [[http://neuroimage.usc.edu/brainstorm/Tutorials/CoregisterSubjects#Volume_source_models|Group analysis: Subject coregistration]]. | * '''Use template grid''': This option is only available when a template source grid has been defined on the default anatomy. For more information, see the tutorial [[http://neuroimage.usc.edu/brainstorm/Tutorials/CoregisterSubjects#Volume_source_models|Group analysis: Subject coregistration]]. |
| Line 27: | Line 27: |
| * '''Preview''': For each combination of parameters, you can see the total number of grid points at the bottom of the option window. Click on [Preview] to see the grid of dipoles. Screen captures below show an adaptive grid (left) and a regular grid (right). <<BR>><<BR>> {{attachment:grid.gif||height="259",width="652"}} * Leave all the intial options, and click on Ok. A new head model is computed. <<BR>><<BR>> {{attachment:treeHeadmodel.gif}} |
* '''Preview''': For each combination of parameters, you can see the total number of grid points at the bottom of the option window. Click on [Preview] to see the grid. The screen captures below show an adaptive grid (left) and a regular grid (right). <<BR>><<BR>> {{attachment:grid.gif||height="259",width="652"}} * Leave all the intial options and click [OK]. A new head model is computed. <<BR>><<BR>> {{attachment:treeHeadmodel.gif}} |
| Line 32: | Line 32: |
| * Right-click on the volume head model > '''Compute sources [2016]''' > Unconstrained '''dSPM'''. <<BR>><<BR>> {{attachment:dspm.gif}} * Note that only the "Unconstrained" source orientation is available. In the previous tutorials, the source space was the cortical surface, and we were able to use the normals to this this surface to constrain the model. For a random grid of points, we cannot privilege one orientation more than another. Three orthogonal dipoles will be defined at each point of the grid. * Right-click on the volume dSPM sources for the Deviant average > '''Cortical activations'''. <<BR>><<BR>> {{attachment:treeSource.gif}} |
* Right-click on this head model > '''Compute sources [2016]''' > Unconstrained '''dSPM'''. <<BR>><<BR>> {{attachment:dspm.gif}} * Note that only the '''Unconstrained''' dipole orientation is available. In the previous tutorials, the source space was the cortical surface, and we were able to use the normals to this surface to constrain the model. For a random grid of points, we cannot privilege one orientation more than another. Three orthogonal dipoles will be defined at each point of the grid. * Right-click on the dSPM sources for the Deviant average > '''Cortical activations'''. <<BR>><<BR>> {{attachment:treeSource.gif}} |
| Line 38: | Line 38: |
| The regions of interest on the surface were introduced in the tutorial [[Tutorials/Scouts|Scouts]]. In a similar way, we can also create regions of interests from the volume sources. In this context, a scout is a subset of the the grid points used to estimate the sources. We cannot create them in the default atlas "User scouts", which is reserved to scouts created directly on the cortex surface. | The regions of interest on the surface were introduced in the tutorial [[Tutorials/Scouts|Scouts]]. In a similar way, we can also create regions of interests from the volume sources. In this context, a scout is a subset of the the grid of points used to estimate the sources. We cannot create them in the default atlas "User scouts", which is reserved to scouts created directly on the cortex surface. |
| Line 40: | Line 40: |
| * Right-click on the volume dSPM sources for the Deviant average > '''Display on MRI (3D)'''. | * Right-click on the dSPM sources for the Deviant average > '''Display on MRI (3D)'''. |
| Line 42: | Line 42: |
| * From the scout tab, create a new volume atlas: menu Atlas > New atlas > '''Volume scouts'''. {{attachment:atlas_volume.gif}} * Create a new volume atlas: menu Atlas > New atlas > Volume scouts. The create scouts by clicking on the MRI slices in a 3D view. To move the slices: right-click an move the mouse. |
* From the scout tab: menu Atlas > New atlas > '''Volume scouts'''. {{attachment:atlas_volume.gif||height="245",width="670"}} * Note that this volume atlas will be valid only for this grid of sources. If you compute another volume head model with a different grid, this atlas won't be available. To help you keep track of this, the name of the new atlas includes the number of grid points (eg. "Volume scouts 15004"). * You can start drawing scouts with the 3D figure: Click on the button '''[Create scout]''', then click on one of the slices to define the center of your scout. The interface detects the coordinates of the point you clicked in the MRI and finds the closest point in the source grid. * To be able to select the region you want, you need to '''move the slices''' before creating the scout: right-click and move the mouse in the direction of the slice, or use the sliders in the section Resect in the Surface tab. * '''Grow''' the ROI with the buttons in the Scout tab ([<<], [<], [>], [>>]) and rename it (double-click). The dots represent the points of the grid that are part of the scout. The surface around them is their convex envelope, to give an idea of the spatial extent of the region. * Create two scouts for the left and right primary auditory cortices: '''A1L''', '''A1R'''. <<BR>><<BR>> {{attachment:scouts_3d.gif||height="282",width="578"}} * Display the scouts time series (first button in the Scout toolbar). With the option "'''Values:Absolute'''" selected, the values are averaged separately for the three orientations (x,y,z), then the norm of these three orientations is displayed: sqrt(x^2^+y^2^+z^2^).<<BR>><<BR>> {{attachment:scouts_ts.gif||height="221",width="482"}} * If you select the option "'''Values:Relative'''", you can observe the amplitude along the three orientations separately instead of the norm. <<BR>><<BR>> {{attachment:scouts_relative.gif||height="222",width="523"}} |
Volume source estimation
Authors: Francois Tadel, John C Mosher
The default approach for the source estimation in Brainstorm is to limit the source space to the cortex surface. This choice is motivated by the assumption that most of the activity we record in MEG and EEG comes from the cerebral cortex. Constraining the source reconstruction to a surface works well when this assumption is verified, and the results we obtain are much easier to review than a full volume.
However, when studying the activity from deeper regions of the brain or when processing recordings from patients with serious anatomical abnormalities, this cortical constraint is not always adapted. This tutorial explains how to construct a grid of dipoles that samples the full brain volume.
Compute a volume head model
The example below uses the protocol TutorialIntroduction created in the introduction tutorials.
Select the protocol TutorialIntroduction. Go to the view "Functional data (sorted by subjects)".
In the imported Run01, right-click on the channel file > Compute head model > MRI volume.
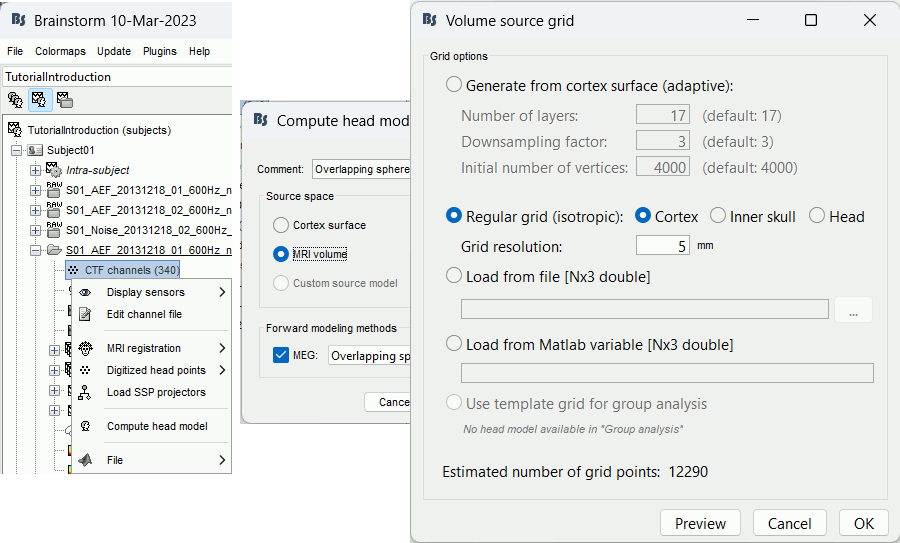
Generate from cortex surface: Creates a grid that samples the brain volume in an adaptive way, denser at the surface (we expect a higher spatial resolution close to the sensors), and sparser at the center of the brain. This grid is created with following algorithm:
Start with a brain envelope with a given number of vertices (Initial number of vertices).
Shrink the previous layer, and downsample it by a given factor (option Downsampling factor).
Repeat this operation [Number of layers] times, or until there are no more vertices.
Regular grid: Creates a regular grid with one grid point every few millimeters (option Grid resolution) in the three directions (x,y,z).
Load from a file/variable: If you already have a grid of points you would like to use, you can import it from a file or a Matlab variable (only one field, [Npointsx3] values).
Use template grid: This option is only available when a template source grid has been defined on the default anatomy. For more information, see the tutorial Group analysis: Subject coregistration.
Preview: For each combination of parameters, you can see the total number of grid points at the bottom of the option window. Click on [Preview] to see the grid. The screen captures below show an adaptive grid (left) and a regular grid (right).
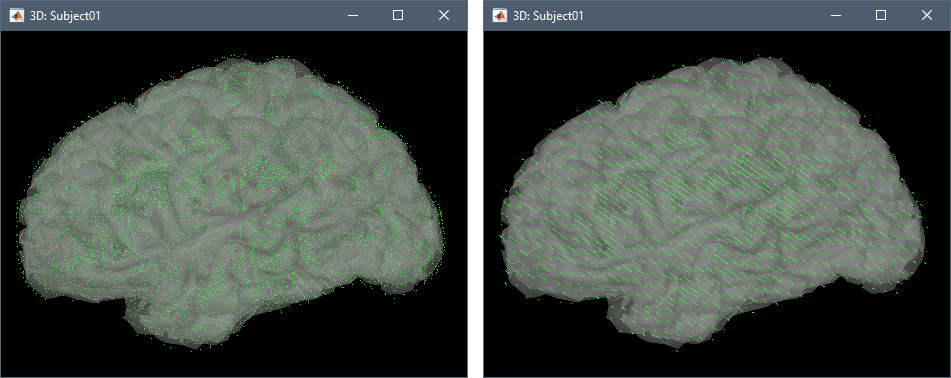
Leave all the intial options and click [OK]. A new head model is computed.

Compute sources
- Make sure your new volume head model is selected as the default one (displayed in green).
Right-click on this head model > Compute sources [2016] > Unconstrained dSPM.
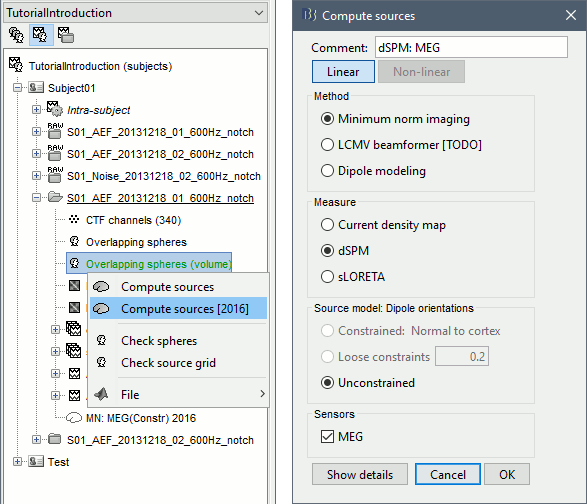
Note that only the Unconstrained dipole orientation is available. In the previous tutorials, the source space was the cortical surface, and we were able to use the normals to this surface to constrain the model. For a random grid of points, we cannot privilege one orientation more than another. Three orthogonal dipoles will be defined at each point of the grid.
Right-click on the dSPM sources for the Deviant average > Cortical activations.
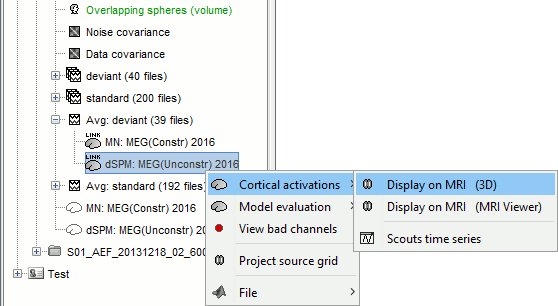
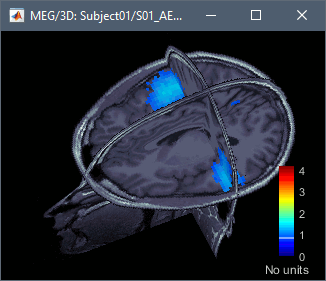
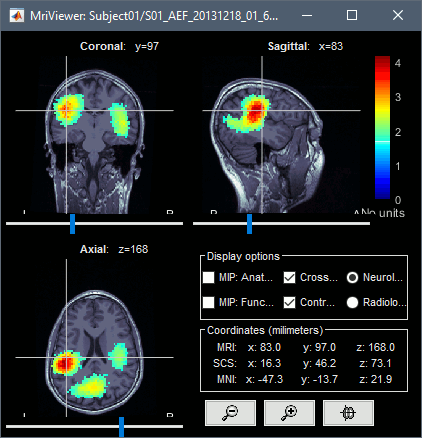
The two menus Display on MRI (3D) and Display on MRI (MRI Viewer) were already introduced in the tutorial Source estimation. You can use the sliders in the Surface tab to adjust the display. The bilateral response in the auditory cortices is very clear starting from 60ms.
Volume scouts
The regions of interest on the surface were introduced in the tutorial Scouts. In a similar way, we can also create regions of interests from the volume sources. In this context, a scout is a subset of the the grid of points used to estimate the sources. We cannot create them in the default atlas "User scouts", which is reserved to scouts created directly on the cortex surface.
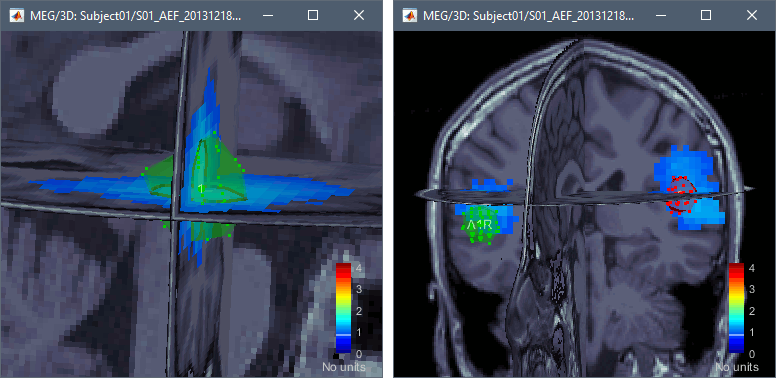
Right-click on the dSPM sources for the Deviant average > Display on MRI (3D).
Go to 60ms and in the Surface tab, set the amplitude threshold to 20%.
From the scout tab: menu Atlas > New atlas > Volume scouts.
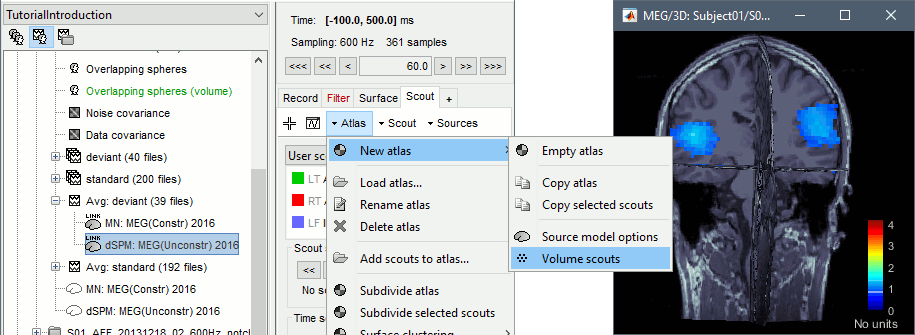
- Note that this volume atlas will be valid only for this grid of sources. If you compute another volume head model with a different grid, this atlas won't be available. To help you keep track of this, the name of the new atlas includes the number of grid points (eg. "Volume scouts 15004").
You can start drawing scouts with the 3D figure: Click on the button [Create scout], then click on one of the slices to define the center of your scout. The interface detects the coordinates of the point you clicked in the MRI and finds the closest point in the source grid.
To be able to select the region you want, you need to move the slices before creating the scout: right-click and move the mouse in the direction of the slice, or use the sliders in the section Resect in the Surface tab.
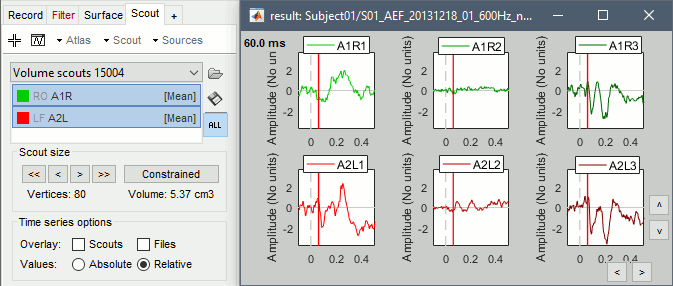
Grow the ROI with the buttons in the Scout tab ([<<], [<], [>], [>>]) and rename it (double-click). The dots represent the points of the grid that are part of the scout. The surface around them is their convex envelope, to give an idea of the spatial extent of the region.
Create two scouts for the left and right primary auditory cortices: A1L, A1R.
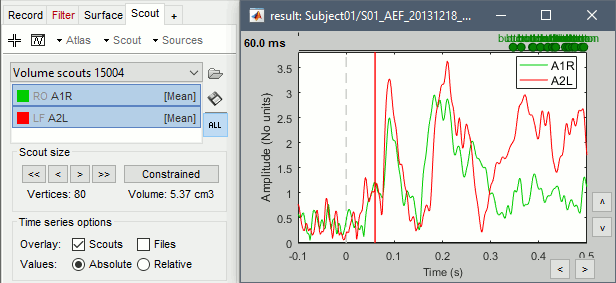
Display the scouts time series (first button in the Scout toolbar). With the option "Values:Absolute" selected, the values are averaged separately for the three orientations (x,y,z), then the norm of these three orientations is displayed: sqrt(x2+y2+z2).
If you select the option "Values:Relative", you can observe the amplitude along the three orientations separately instead of the norm.