|
Size: 5462
Comment:
|
Size: 5442
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| In EEG, a lot of the analysis can be done directly on the electrodes values. This tutorial explains how to group sensors in a cluster and overlay channels from different conditions in the same graph. | In EEG, most of the analysis can be done directly on the electrode values. This tutorial explains how to group sensors in a cluster and overlay channels from different conditions in the same graph. |
| Line 31: | Line 31: |
| * '''Saving''': The clusters are not saved neither in the database files, nor in the Brainstorm user preferences, therefore if you close Brainstorm you would your clusters. To keep them between sessions, you can use the ['''Save'''] and ['''Open'''] buttons on the right side of the tab. | * '''Saving''': The clusters are not saved in the database files, nor in the Brainstorm user preferences, therefore if you close Brainstorm you would lose your clusters. To keep them between sessions, you can use the ['''Save'''] and ['''Open'''] buttons on the right side of the tab. |
| Line 34: | Line 34: |
| We have now two clusters available in the list. The second one contains only one channel, displaying it corresponds to displaying the channel time series. The first one contains multiple channels, we need to apply a function to group them into one signal only. | We now have two clusters available in the list. The second one contains only one channel, displaying it corresponds to displaying the channel time series. The first one contains multiple channels, we need to apply a function to group them into one signal only. |
| Line 37: | Line 37: |
| * Select one cluster and click on the '''[Display clusters]''' button (third from the left in the toolbar). <<BR>><<BR>> {{attachment:cluster_view1.gif||height="149",width="434"}} * Select the two clusters at once and click on '''[''''''Display clusters]''' again. <<BR>><<BR>> {{attachment:cluster_view2.gif||height="140",width="344"}} * To see the two signals in the same graph, select the option '''"Overlay: Cluster"'''. Click on '''[Display]'''. <<BR>><<BR>> {{attachment:cluster_view3.gif||height="124",width="438"}} |
* Select one cluster and click on the '''[Display cluster]''' button (third from the left in the toolbar). <<BR>><<BR>> {{attachment:cluster_view1.gif||height="149",width="434"}} * Select the two clusters at once and click on '''[''''''Display cluster]''' again. <<BR>><<BR>> {{attachment:cluster_view2.gif||height="140",width="344"}} * To have the two signals in the same graph, select the option '''"Overlay: Cluster"'''. Click on '''[Display]'''. <<BR>><<BR>> {{attachment:cluster_view3.gif||height="141",width="500"}} |
| Line 41: | Line 41: |
| * In the database explorer: Double-click on the '''deviant average '''of the same run.<<BR>>Click on '''[Display clusters]''' again, now it will display the cluster values for the deviant condition.<<BR>><<BR>> {{attachment:cluster_view4.gif||height="130",width="439"}} * Unselect '''"Overlay: Cluster"''' and select '''"Overlay: Conditions"''', it will group the signals in a different way: one graph per cluster, each one showing the two condition averages. <<BR>><<BR>> {{attachment:cluster_view5.gif||height="129",width="438"}} |
* In the database explorer: Double-click on the '''deviant average '''of the same run.<<BR>>Click on '''[Display cluster]''' again, now it displays the clusters values for both conditions.<<BR>><<BR>> {{attachment:cluster_view4.gif||height="148",width="500"}} * Unselect '''"Overlay: Cluster"''' and select '''"Overlay: Conditions"''', it will group the signals in a different way: one graph per cluster, each one showing the two condition averages. <<BR>><<BR>> {{attachment:cluster_view5.gif||height="147",width="500"}} |
| Line 45: | Line 45: |
| Once the clusters are defined, you can use apply them to any number of files in the database in one click. | Once the clusters are defined, you can apply them to any number of files in the database in one click. |
Tutorial 19: Clusters of sensors
Authors: Francois Tadel, Elizabeth Bock, Sylvain Baillet
In EEG, most of the analysis can be done directly on the electrode values. This tutorial explains how to group sensors in a cluster and overlay channels from different conditions in the same graph.
In MEG, we tend to avoid working at the sensor level because the signals are ambiguous, we work mostly in the source space, which is the topic of the next tutorials. If you are planning to work only with MEG recordings, you may skip this tutorial.
Cluster tab
The Cluster tab is not shown by default in the Brainstorm interface.
Click on the [+] button next to the Scout tab > Add: Cluster.
If the list of tabs is too busy, you can close some of them in the same way.
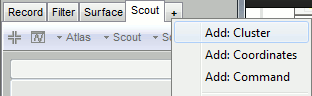
The Cluster tab is similar to the Scout tab, but much simpler. It contains a toolbar to create and edit clusters, a list of existing clusters and a few display options at the bottom.
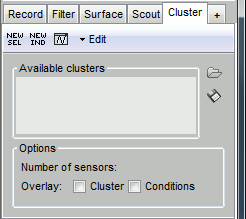
[NEW SEL]: Create a cluster containing the channels that are currently selected in the interface.
[NEW IND]: Create a cluster with channels selected by index, name or type.
[Display clusters time series]: Shows the time series of the selected clusters for all the files that are currently open in the interface.
Edit: Menus to modify the selected clusters.
Creating clusters
A cluster is a group of one or multiple channels. There are two ways for creating a new one.
Display the MEG recordings for the standard average of Run#01 (double-click on the file).
Add a 2D topography (Ctrl+T), display the sensors and select a group of them (right-click+move).
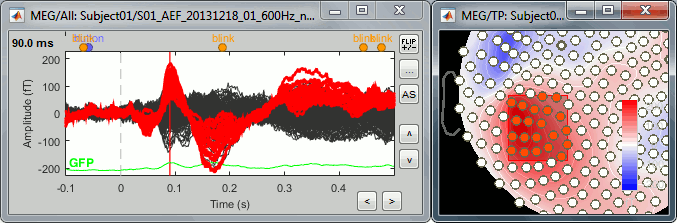
In the Cluster tab, click on [NEW SEL]. It creates a new cluster "c1".
Double-click on it to rename it (or menu Edit > Rename).In the Cluster tab, click on [NEW IND]. Enter the name of the sensor "MLT12". Rename it.
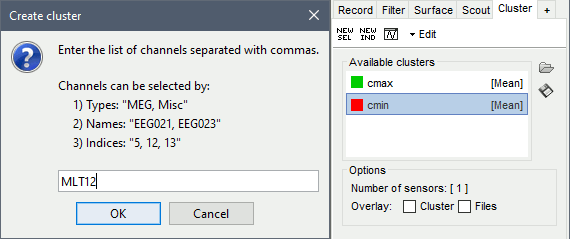
Selection: When you change the selected cluster in the list, it selects the corresponding sensors in all the figures. The number of channels involved in the selection is displayed at the bottom of the tab. You can select multiple clusters at once, holding the Ctrl/Shift/Command button while clicking.
Saving: The clusters are not saved in the database files, nor in the Brainstorm user preferences, therefore if you close Brainstorm you would lose your clusters. To keep them between sessions, you can use the [Save] and [Open] buttons on the right side of the tab.
Displaying clusters
We now have two clusters available in the list. The second one contains only one channel, displaying it corresponds to displaying the channel time series. The first one contains multiple channels, we need to apply a function to group them into one signal only.
The cluster function used to group multiple signals into one is shown on the right of the cluster list. By default it uses a simple average ("Mean"). You can edit this cluster function with the menu Edit > Set cluster function.
Select one cluster and click on the [Display cluster] button (third from the left in the toolbar).
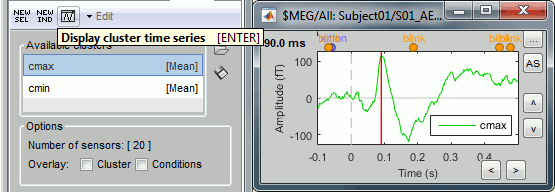
Select the two clusters at once and click on [Display cluster] again.
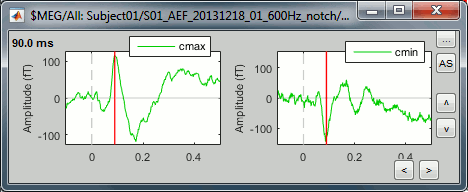
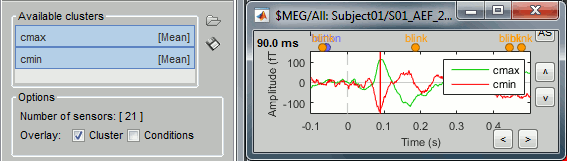
To have the two signals in the same graph, select the option "Overlay: Cluster". Click on [Display].
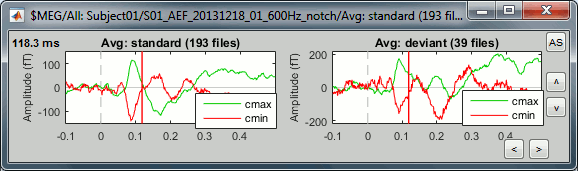
In the database explorer: Double-click on the deviant average of the same run.
Click on [Display cluster] again, now it displays the clusters values for both conditions.
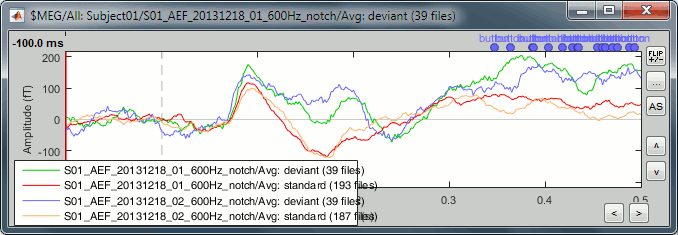
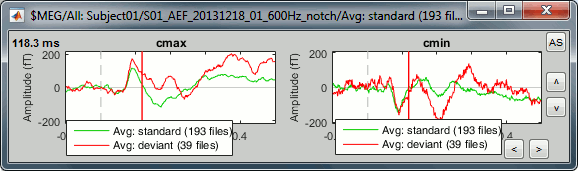
Unselect "Overlay: Cluster" and select "Overlay: Conditions", it will group the signals in a different way: one graph per cluster, each one showing the two condition averages.
From the database explorer
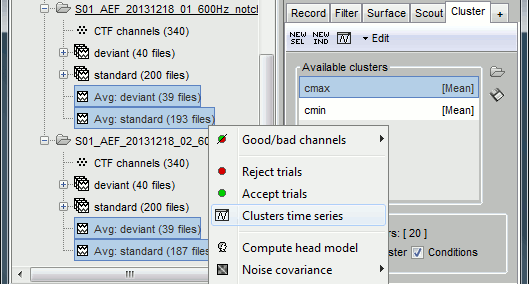
Once the clusters are defined, you can apply them to any number of files in the database in one click.
- Close all the figures with the [X] button in the top-right corner of the Brainstorm figure.
In the Cluster tab, select only one of the two clusters, select Overlay:Condition.
In the database explorer, select all the averages for both runs. Right-click on one of the selected files > Clusters time series. On this graph, you can verify that the selected cluster behaves in the same way in the two acquisition runs.