|
Size: 4497
Comment:
|
Size: 4495
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 41: | Line 41: |
Using the anatomy templates
Author: Francois Tadel
Contents
Changing the default anatomy
When you create a new protocol, the program makes a copy of the ICBM152 or Colin27 anatomy and sets it as the default for the protocol. It means that you will be able to use this template brain as a substitute for the subjects without an individual MRI, or as the common brain for group analysis.
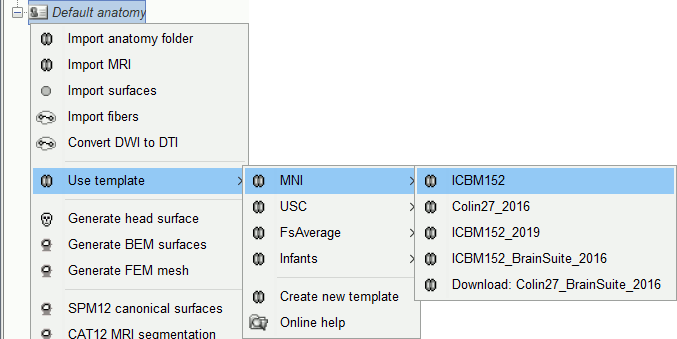
Other sets of MRI+surfaces are available to replace the Colin27 anatomy. Right-click on (Default anatomy) > Use template. If a package is not currently available on your system, it will be downloaded from the Brainstorm website and saved in $HOME/.brainstorm/templates.
If you click on any of the download options, it downloads it into your $HOME/.brainstorm/templates folder, then the list of files in the (default anatomy) folder is replaced with the new template.
Group analysis
When performing a group analysis with multiple subjects for which you have the individual MRI scans, you need to project the sources estimated on each subject on a common template, as explained in this tutorial: Group analysis.
FreeSurfer templates
The available options are:
Colin27: Average of 27 scans of the same head, processed with FreeSurfer 5.3: more information
Colin27_2012: Previous version of the default anatomy distributed with Brainstorm
ICBM152: Non-linear average of 152 subjects, processed with FreeSurfer 5.3: more information
FSAverage: Average of 40 subjects using a spherical averaging described in (Fischl et al. 1999).
It is the default FreeSurfer brain: please register here if you are using it.
More images: http://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#FSAverage_templateInfant7w: 7-week infant brain with the antomical atlas presented in (Kabdebon et al. 2014).
Oreilly_1y: 1 year old infant brain with Tzourio-Mazoyer surface atlas (Li et al. 2015, Shi et al. 2011): http://neuroimage.usc.edu/forums/showthread.php?2123-Atlas-for-1-year-old-babies
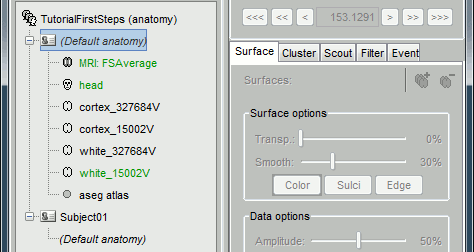
They all include the following information:
- T1 MRI volume
- Cortex surface: high-resolution (~300.000 vertices) and low-resolution (15.000 vertices)
- Head surface: based on the head used for FSAverage in the MNE software
FreeSurfer spherical registration of each hemisphere, with which we can co-register the individual brains processed with FreeSurfer with the selected default anatomy
FreeSurfer surface-based atlases: Desikan-Killiany, Destrieux, Brodman, Mindboggle
(plus Yeo2011 and PALS for FSAverage only)
The atlases will be discussed in the following tutorials. For more information on the interactions between FreeSurfer and Brainstorm: read this tutorial.
BrainSuite templates
Modify the default MRI fiducials
The fiducial points (Nasion, LPA, RPA) used in your recordings might not be the same as the ones used in the anatomy templates in Brainstorm (Colin27, ICBM152, FSAverage). By default, the LPA/RPA points are defined at the junction between the tragus and the helix, as represented with the red dot in the Coordinates systems page.
If you want to use an anatomy template but you are using a different convention when digitizing the position of these points, you have to modify the default positions of the template with the MRI Viewer.
- Go to the anatomy view
In (default anatomy), right-click on the MRI > Edit MRI
- Modify the position of the fiducial points to match your own convention
- Click on [Save], it will update the surfaces to match the new coordinate system