|
Size: 5059
Comment:
|
← Revision 85 as of 2022-11-28 10:28:33 ⇥
Size: 24051
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 2: | Line 2: |
| ''Authors: Francois Tadel, Guiomar Niso, Elizabeth Bock. '' This tutorials introduces how to download a resting state recordings from the OMEGA database, and how to process them into Brainstorm. |
''Authors: Francois Tadel, Guiomar Niso, Elizabeth Bock, [[https://www.neurospeed-bailletlab.org/sylvain-baillet|Sylvain Baillet]]'' This tutorial introduces how to download resting state recordings from the OMEGA database and process them into Brainstorm. The goal is to evaluate where in the brain is distributed the power in specific frequency bands at rest. This example includes only 5 healthy subjects, but the same pipeline can be applied over larger populations. Note that the operations used here are not detailed, the goal of this tutorial is not to introduce Brainstorm to new users. For in-depth explanations of the interface and theoretical foundations, please refer to the [[http://neuroimage.usc.edu/brainstorm/Tutorials#Get_started|introduction tutorials]]. The processing pipeline is published in this article:<<BR>>Niso G, Tadel F, Bock E, Cousineau M, Santos A, Baillet S, [[https://www.frontiersin.org/articles/10.3389/fnins.2019.00284/abstract|Brainstorm Pipeline Analysis of Resting-State Data from the Open MEG Archive]], Frontiers in Neuroscience, Mar 2019 |
| Line 8: | Line 12: |
| == License == The Open MEG Archive (OMEGA) is the fruit of a collaborative effort by the McConnell Brain Imaging Centre and the Université de Montréal to build a centralised repository in which to regroup MEG data for open dissemination. This continuously expanding repository also contains anatomical MRI volumes, demographic and questionnaire information, and eventually will feature other forms of electrophysiological data (e.g. EEG, field, and cell recordings). OMEGA features the technological framework for multi-centric data aggregation, and is amongst the largest freely available resting-state MEG datasets presently available. |
== OMEGA database == The Open MEG Archive (OMEGA) is the fruit of a collaborative effort by the McConnell Brain Imaging Centre and the Université de Montréal to build a centralized repository in which to regroup MEG data for open dissemination. This continuously expanding repository also contains anatomical MRI volumes, demographic and questionnaire information, and eventually will feature other forms of electrophysiological data (e.g. EEG, field, and cell recordings). OMEGA features the technological framework for multi-centric data aggregation, and is amongst the largest freely available resting-state MEG datasets presently available. Website: https://www.mcgill.ca/bic/resources/omega |
| Line 17: | Line 21: |
| * 5 subjects * 5 minute resting sessions, eyes open |
* 5 subjects x 5 minute resting sessions, eyes open |
| Line 23: | Line 26: |
| * Anti-aliasing low-pass filter at 600Hz, files saved with the CTF 3rd order gradient compensation. * Recorded channels (340): * 1 Stim channel indicating the presentation times of the audio stimuli: UPPT001 (#1) * 1 Audio signal sent to the subject: UADC001 (#316) * 1 Response channel recordings the finger taps in response to the deviants: UDIO001 (#2) * 26 MEG reference sensors (#5-#30) * 274 MEG axial gradiometers (#31-#304) * 2 EEG electrodes: Cz, Pz (#305 and #306) * 1 ECG bipolar (#307) * 2 EOG bipolar (vertical #308, horizontal #309) * 12 Head tracking channels: Nasion XYZ, Left XYZ, Right XYZ, Error N/L/R (#317-#328) * 20 Unused channels (#3, #4, #310-#315, #329-340) * The data in this dataset has been formatted along the BIDS specifications (Brain Imaging Data Structure, http://bids.neuroimaging.io) |
* Anti-aliasing low-pass filter at 600Hz * Files may be saved with or without the CTF 3rd order gradient compensation applied. * Recorded channels (at least 297): * 26 MEG reference sensors (#2-#27) * 270 MEG axial gradiometers (#28-#297) * 1 ECG bipolar (EEG057/#298) - Not available in the empty room recordings * 1 vertical EOG bipolar (EEG058/#299) - Not available in the empty room recordings * 1 horizontal EOG bipolar (EEG059/#300) - Not available in the empty room recordings * The data in this dataset has been organized according to the BIDS specifications<<BR>>(Brain Imaging Data Structure, http://bids.neuroimaging.io) |
| Line 38: | Line 37: |
| * 3D digitization using a Polhemus Fastrak device driven by Brainstorm. The .pos files contain: * The position of the center of CTF coils * The position of the anatomical references we use in Brainstorm: Nasion and connections tragus/helix, as illustrated [[http://neuroimage.usc.edu/brainstorm/CoordinateSystems#Pre-auricular_points_.28LPA.2C_RPA.29|here]] * Around 150 head points distributed on the hard parts of the head (no soft tissues) |
3D digitization using a Polhemus Fastrak device driven by Brainstorm. The .pos files contain: * The center of the CTF coils, * The anatomical references we use in Brainstorm: nasion and ears as illustrated [[http://neuroimage.usc.edu/brainstorm/CoordinateSystems#Pre-auricular_points_.28LPA.2C_RPA.29|here]], * Around 100 head points distributed on the hard parts of the head (no soft tissues). |
| Line 44: | Line 44: |
| * Structural T1 image recorded with a 1.5T MRI | * Structural T1 image (defaced for anonymization purposes) |
| Line 48: | Line 48: |
| Note that while the subjects shared here have their anatomy already processed with FreeSurfer to simplify the tutorial, this is not currently the case for data shared in OMEGA. You would therefore need to generate surfaces yourself as explained in the [[https://neuroimage.usc.edu/brainstorm/Tutorials/ImportAnatomy#Import_the_anatomy|anatomy tutorial]]. |
|
| Line 49: | Line 51: |
| * First, make sure you have at least '''20Gb''' of free space on your hard drive (raw files: 12Gb). | * First, make sure you have at least '''40Gb''' of free space on your hard drive (raw files: 12Gb). |
| Line 51: | Line 53: |
| * Go to the [[http://neuroimage.usc.edu/bst/download.php|Download]] page, and download the file: '''sample_omega.zip ''' | * The data is hosted on the OpenNeuro website (click on the icon "Download dataset" at the top):<<BR>>https://openneuro.org/datasets/ds000247 * The safest alternative is to use the Amazon AWS command line tools: * Install AWS CLI: https://aws.amazon.com/cli/ (no need to create an account) * Linux/MacOS: * In a terminal, type: pip install awscli * Windows: * Download and install the program (64bit version, most likely) * Open a command line terminal (Windows menu, type "cmd", enter) * Go to the AWSCLI install folder (type: cd "C:\Program Files\Amazon\AWSCLI") * Start the download (synchronization with an Amazon S3 repository): * Create a new folder "ds000247" somewhere on a large hard drive (NOT in the Brainstorm program or database folders), referred to below as <target_directory>. * In the same terminal, type: * aws s3 sync --no-sign-request s3://openneuro.org/ds000247 <target_directory> |
| Line 61: | Line 78: |
| == BIDS specification == The [[http://bids.neuroimaging.io|Brain Imaging Data Structure (BIDS)]] standard for neuroimaging data organization was first established for MRI and fMRI ([[http://www.nature.com/articles/sdata201644|Gorgolewski, 2016]]). It is based on simple file formats and folder structures that can readily expand to additional data modalities. An extension of the BIDS for MEG datasets has now been published. The data distributed with this tutorial is organized following this prototype (which may change in the future). One objective is that analysis pipelines designed with major analysis tools (such as Brainstorm, FieldTrip, MNE, SPM and others) can be readily applied without requiring software or pipeline redevelopments. Brainstorm includes a set of functions that can load automatically any dataset following the BIDS specification. The files that are read by Brainstorm are the following: '''OMEGA_RestingState_sample'''/ * '''derivatives'''/: Everything that cannot be considered as raw data * '''freesurfer'''/: Result of the FreeSurfer segmentation for all the subjects * '''sub-000X_ses-mri'''/'''anat'''/: Output of FreeSurfer for subject 000X (session 0001) * '''sub-000X'''/: Raw data for subject 000X * '''ses-0001'''/: Only one session per subject (the subject visited the center only once) * '''sub-0002_ses-0001_scans.tsv''': BIDS tab-separated text file listing the MEG recordings and corresponding acquisition dates * '''anat'''/: Anatomical MRI scans for subject 000X (not used if FreeSurfer output available) * '''sub-000X_ses-0001_T1w.nii.gz''': T1 scan * '''meg'''/: Unprocessed MEG recordings * '''sub-000X_ses-0001'''_'''task-rest_run-01_meg.ds''': Resting MEG recordings * '''sub-emptyroom'''/: Empty room measures at different dates (used for noise modelling) * '''ses-XXX'''/: Various sessions of noise recordings (can be matched by date with the subjects' recordings using the *_scans.tsv BIDS files). * '''sub-emptyroom_ses-XXX_scans.tsv''': BIDS tab-separated text file listing the MEG recordings and corresponding acquisition dates * '''meg'''/: Unprocessed MEG empty room recordings * '''sub-000X_ses-XXX_task-noise_run-01_meg.ds''' |
|
| Line 62: | Line 104: |
| * The datasets organized following the [[http://bids.neuroimaging.io|BIDS specifications]] can be imported semi-automatically. * Select the menu File > Load protocol > '''Import BIDS dataset''' > Select the folder '''sample_omega'''. * Keep the default values for all the questions that may be asked during the import process (eg. number of vertices in the cortex surfaces). Once finished, you should be able to see 5 subjects in your database explorer: anatomy, and subject and noise recordings. <<BR>><<BR>> * The files that have been imported with this process are the following: * sample_omega/derivatives/freesurfer/sub-000*_ses-0001: MRIs processed with FreeSurfer. * sample_omega/sub-000*/ses-0001/meg/*_task-rest_run-01_meg.ds: Resting MEG recordings * sample_omega/sub-000*/ses-0001/meg/*_task-noise_run-01_meg: Empty room recordings |
* Select the menu File > Load protocol > '''Import BIDS dataset''' > Select folder '''OMEGA_RestingState_sample'''.<<BR>><<BR>> {{attachment:bids_import.gif}} * '''Folder to import''': The root folder of the BIDS dataset to import. * '''Names of subjects''': If you want to import only a few specific subjects from the dataset, list them here separated with commas. Otherwise, leaving the field empty imports all the subjects available in the dataset. * '''Number of vertices''': If the segmentation of the MRI with [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer|FreeSurfer]], [[https://neuroimage.usc.edu/brainstorm/Tutorials/SegCAT12|CAT12]], [[https://neuroimage.usc.edu/brainstorm/Tutorials/SegBrainSuite|BrainSuite]] or [[https://neuroimage.usc.edu/brainstorm/Tutorials/SegBrainVisa|BrainVISA]] is available in the [[https://neurostars.org/t/bids-derivatives-for-freesurfer/4839|derivatives folder]], this process will import it just like it would do with the menu [[https://neuroimage.usc.edu/brainstorm/Tutorials/ImportAnatomy#Import_the_anatomy|Import anatomy folder]]. Default fiducials positions are computed using the affine MNI normalization (maff8). This process includes downsampling the cortex surface to a specific number of vertices, which you can specify with this option. * '''MNI normalization''': For the first volume imported into the database, when it is not part of a segmentation folder from the derivatives (FreeSurfer or other), computes the [[https://neuroimage.usc.edu/brainstorm/CoordinateSystems#MNI_coordinates|MNI normalization]] using the selected method. * '''Align sensors using headpoints''': When importing EEG or MEG sensor definitions, if there are enough digitized head points, this option enables running automatically the process [[https://neuroimage.usc.edu/brainstorm/Tutorials/ChannelFile#Automatic_registration|Refine registration with head points]]. * '''Import multiple anat sessions''': If this option is ''not'' selected, a subject with multiple anat sessions would lead to the creation of multiple subjects in the Brainstorm database. When selected, all the anatomy volumes from multiple sessions are imported into the anatomy of the same subject. * Keep the default values for all the questions that may be asked during the import process (eg. number of vertices in the cortex surfaces). Once done, you should be able to access the data for the 5 subjects in your database explorer: anatomy, and subject and noise recordings. <<BR>><<BR>> {{attachment:load_bids.gif||width="700"}} * It is important to understand how the spatial coregistration between the anatomy (MRI and FreeSurfer surfaces) and the MEG sensors was done here, as it is not a trivial question. The two procedures available in Brainstorm for this registration are described in the tutorial [[http://neuroimage.usc.edu/brainstorm/Tutorials/ChannelFile#Automatic_registration|Channel file & MRI registration]]: using three anatomical fiducials NAS/LPA/RPA and/or using the digitized head shape. In this dataset we '''do not have access to the positions of the fiducials''' in the MRI, however, the digitized head shapes (same coordinate system as the MEG) are available in the files *_headshape.pos. Therefore, we defined automatically approximate positions for the fiducials NAS/LPA/RPA in the MRI using standard [[http://neuroimage.usc.edu/brainstorm/CoordinateSystems#MNI_coordinates|MNI coordinates]], with the expectation that this approximate registration would later be refined using the head shape. This is done in the next section. * Another solution could have been to first mark manually the positions of the anatomical landmarks using the MRI viewer in Brainstorm. The menu '''File > Batch MRI fiducials''' opens the MRI viewer for each subject, asks for marking the NAS/LPA/RPA landmarks, and creates a file '''fiducials.m''' in the FreeSurfer folder. When importing the FreeSurfer folders, these fiducials are used by default. * '''TODO''': The BIDS-MEG standards are still in development and do not include yet a final solution for saving these anatomical fiducials in the BIDS data structure. When this feature is implemented, we will add the corresponding fields in this example dataset and will update the import function to read the correct fiducials positions automatically. == Refine the MEG-MRI registration == * At this point, the registration MEG/MRI is based only on the three approximate anatomical landmarks NAS/LPA/RPA (standard positions from the MNI template) and must improved. All the MRI scans were anonymized (defaced) so all the head points below the nasion cannot be used. We will try to refine this registration using the additional head points that were digitized above the nasion. * Drag and drop the rest recordings for all the subjects (exluding sub-emptyroom) into the Process1 box, and click on [Run]. * Select process: '''Import > Import anatomy > Remove head points''', Z=0 * Add process: '''Import > Channel file > Refine registration''', then run the execution.<<BR>><<BR>> {{attachment:registration_refine.gif||width="666",height="256"}} * Check the results for one subject: right-click on CTF channels > MRI registration > Check. <<BR>><<BR>> {{attachment:registration_after.gif||width="471",height="229"}} == Pre-processing == * Switch to the functional view of the protocol (second button above the database explorer). * Drag and drop all the subjects (including sub-emptyroom) into the Process1 box, and click on [Run]. * Select process: '''Import > Import recordings >''' '''Convert to continuous (CTF)''': Continuous * Add process: '''Frequency >''' '''Power spectrum density (Welch)''': All file, 4s, 50% overlap, Individual <<BR>><<BR>> {{attachment:psd_process.gif||width="573",height="325"}} * Run, then check the quality of the PSD for all the files, as documented in [[http://neuroimage.usc.edu/brainstorm/Tutorials/ArtifactsFilter#Interpretation_of_the_PSD|this tutorial]]. <<BR>><<BR>> {{attachment:psd_before.gif||width="343",height="152"}} * In Process1, keep the same files selected and click on [Run]. * Select process: '''Pre-process >''' '''Notch filter''': 60 120 180 240 300 Hz, Process the entire file at once * Add process: '''Pre-process >''' '''Band-pass filter''': High-pass filter at 0.3Hz, 60dB, Process entire file. * Add process: '''Frequency >''' '''Power spectrum density (Welch)''': Same options <<BR>><<BR>> {{attachment:filter_process.gif||width="597",height="342"}} * Run, then check the PSD after filtering. <<BR>><<BR>> {{attachment:psd_after.gif||width="389",height="140"}} * Delete all the folders corresponding to the original recordings (_meg) and the notch filtered data (_meg_notch). Only keep the fully processed files (_meg_notch_high). <<BR>><<BR>> {{attachment:delete_original.gif}} <<BR>><<BR>> {{attachment:delete_after.gif}} == Artifact cleaning == We will now run the automatic procedure for cleaning the heartbeats, as described in the introduction tutorials ([[Tutorials/ArtifactsDetect|detection]], [[Tutorials/ArtifactsSsp|SSP]]). The results we want to illustrate here are robust enough, the recordings do not need to be processed any further. If you want to improve the quality of the data with more manual cleaning (blinks, saccades, movements, bad segments), please refer to the introduction tutorials. * In Process1, select all the subject rest recordings (all the subjects, excluding sub-emptyroom). * Select process: '''Events > Detect heartbeats''': ECG, All file, cardiac * Add process: '''Artifacts > SSP: Heartbeats''': cardiac, MEG, Use existing SSP * Add process: '''File > Snapshot''': Sensor/MRI registration * Add process: '''File > Snapshot''': SSP projectors<<BR>><<BR>> {{attachment:ssp_process.gif||width="548",height="304"}} * Run, then review the registration and the SSP topographies for all the subjects. Make sure that the subjects' heads are well aligned in the MEG, and that the spatial components that are removed are really representative of the cardiac artifact. <<BR>><<BR>> {{attachment:ssp_report.gif||width="489",height="378"}} == Source estimation == * In Process1, select all the noise recordings (all the recordings in sub-emptyroom). * Run process: '''Sources > Compute covariance''': All file, Noise covariance, Copy to other folders, Copy to other subjects, '''Match by acquisition date'''. <<BR>><<BR>> {{attachment:noiscov_process.gif}} * The option '''Match noise and subject recordings by acquisition date''' uses the recordings dates read from the *_scans.tsv files in the BIDS dataset to copy in each rest recording session the noise covariance corresponding to its acquisition date. The empty room recordings with the closest acquisition date (in number of days, before or after) is used to estimate the noise covariance for each subject. * In Process1, select all the subject rest recordings (all the subjects, excluding sub-emptyroom). * Run process: '''Sources > Compute head model''': Cortex surface, MEG=Overlapping spheres. <<BR>><<BR>> {{attachment:headmodel_process.gif||width="407",height="348"}} * Run process: '''Sources > Compute sources [2016]''': Kernel only, one per file, dSPM, constrained. <<BR>><<BR>> {{attachment:inverse_process.gif||width="629",height="289"}} == Power maps == We have now access to continuous recordings in source space, we can estimate the average power over this resting period, for various frequency bands. We propose here to compute the PSD from the first 100s instead of the full 600s rest recordings, it is faster and leads to very similar results. For more stable estimates, you can compute the the PSD over all the data available instead. * In Process1, select all the subject rest recordings ('''task-rest'''), click the '''[Process sources]''' button. * Select process: '''Frequency > Power spectrum density (Welch)''': [0,100s], Window=4s, 50% overlap, Group in frequency bands (use the default frequency bands), Save individual PSD values. * Add process: '''Standardize > Spectrum normalization''': Relative power (divide by total power). This normalization is done independently for each source, the output for each frequency band is a percentage of how much this frequency band contributes to the total power of the source signal. * Add process: '''Sources > Project on default anatomy''': Cortex. * Add process: '''Sources > Spatial smoothing''': FWHM=3mm, Overwrite.<<BR>><<BR>> {{attachment:psd_sources_process.gif||width="656",height="359"}} <<BR>><<BR>> {{attachment:psd_sources_normalize.gif||width="634",height="330"}} * Execute this pipeline, then open the results for the different subjects and visualize the different frequency bands. They should look similar across subjects. This measure of relative power is very robust and usually do not vary much between subjects or duration of the input signals. * The values in this figure represent the relative power of each source. At each vertex of the cortex surface, the value between 0 and 1 indicates what is the contribution of the current frequency band to the total power in the signal. To obtain comparable colormaps across multiple figures, you can set a fixed maximum (right-click on a figure > Colormap: Timefreq > Maximum: Custom).<<BR>><<BR>> {{attachment:psd_sources_proj.gif||width="667",height="278"}} * We will now average these files across subjects. In Process1, select all the projected subject averages in the ''Group analysis'' folder, and click on the button '''[Process time-freq]'''. * Run process: '''Average > Average files''': Everything, Arithmetic average, Do not match signals <<BR>><<BR>> {{attachment:psd_sources_avgsubj.gif||width="489",height="459"}} * Double-click on the average to display it. To create the figures below, right-click on the figure > Snapshot > Frequency contact sheet. <<BR>><<BR>> {{attachment:psd_sources_left.gif||width="353",height="152"}} {{attachment:psd_sources_top.gif||width="279",height="179"}} * We can relate these figures with associations commonly reported in the literature: * '''Theta''': Mostly frontal. * '''Alpha''': Parieto-occipital. * '''Beta''': Central regions (motor cortex). * '''Gamma''': Pre-frontal and occipital. However, the occipital sources could be related with muscle activity in the neck. We haven't cleaned the recordings much, we should go back to the recordings and mark as bad all the segments in which we observe a lot of muscle activity. * '''Delta''': Questionable - these source maps showing high values at the poles of the frontal and temporal lobes are typical of eye movements. We should try doing some cleaning of the ocular artifacts (blinks and saccades) and compute these maps again. * You can compute the same results in the head volume. You just have to use a volume head model on a group grid instead of the one we used here. This is explained in the tutorial [[http://neuroimage.usc.edu/brainstorm/Tutorials/CoregisterSubjects#Volume_source_models|Group analysis]]. This seems to confirm that the power in the delta and gamma bands is mostly coming from non-brain regions (eyes and neck muscles). <<BR>><<BR>> {{attachment:psd_volume_delta.gif||width="297",height="293"}} {{attachment:psd_volume_gamma.gif||width="297",height="294"}} * You can produce similar plots at the sensor level: Compute the PSD by frequency band of the rest recordings for each subject, then compute the average across subjects (not as reliable as averaging in source space, as [[http://neuroimage.usc.edu/brainstorm/Tutorials/Workflows#Average-1|explained here]]). <<BR>><<BR>> {{attachment:psd_avgsubj.gif||width="636",height="250"}} == Scripting == The following script from the Brainstorm distribution reproduces the analysis presented in this tutorial page: [[https://github.com/brainstorm-tools/brainstorm3/blob/master/toolbox/script/tutorial_omega.m|brainstorm3/toolbox/script/tutorial_omega.m]] <<HTML(<div style="border:1px solid black; background-color:#EEEEFF; width:720px; height:500px; overflow:scroll; padding:10px; font-family: Consolas,Menlo,Monaco,Lucida Console,Liberation Mono,DejaVu Sans Mono,Bitstream Vera Sans Mono,Courier New,monospace,sans-serif; font-size: 13px; white-space: pre;">)>><<EmbedContent("http://neuroimage.usc.edu/bst/viewcode.php?f=tutorial_omega.m")>><<HTML(</div >)>> == Additional documentation == '''Articles''' * '''BIDS: '''Gorgolewski, K.J., Auer, T., Calhoun, V.D., Craddock, R.C., Das, S., Duff, E.P., Flandin, G., Ghosh, S.S., Glatard, T., Halchenko, Y.O. and Handwerker, D.A. (2016), [[https://www.nature.com/articles/sdata201644|The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments]], Scientific Data, 3, p.160044. * '''MEG-BIDS: '''Niso G, Flandin G, Gramfort, A Henson, RN, Jas M, Litvak V, Moreau JT, Oostenveld R, SchoffelenJ-M, Tadel F, Wexler J, Baillet S (2018), [[https://dx.doi.org/10.1038/sdata.2018.110|MEG-BIDS, the brain imaging data structure extended to magnetoencephalography]], Scientific Data, 5: 180110 EP * '''EEG-BIDS:''' Pernet CR, Appelhoff S, Flandin G, Phillips C, Delorme A, Oostenveld R (2018), [[https://doi.org/10.31234/osf.io/63a4y|BIDS-EEG: an extension to the Brain Imaging Data Structure (BIDS) Specification for electroencephalography]], PsyArXiv doi:10.31234/osf.io/63a4y '''Forum''' * Normalize PSD based on resting-state before statistical tests: https://neuroimage.usc.edu/forums/t/psd-normalization-based-on-the-resting-psd/37854 |
Resting state recordings from the OMEGA database
Authors: Francois Tadel, Guiomar Niso, Elizabeth Bock, Sylvain Baillet
This tutorial introduces how to download resting state recordings from the OMEGA database and process them into Brainstorm. The goal is to evaluate where in the brain is distributed the power in specific frequency bands at rest. This example includes only 5 healthy subjects, but the same pipeline can be applied over larger populations.
Note that the operations used here are not detailed, the goal of this tutorial is not to introduce Brainstorm to new users. For in-depth explanations of the interface and theoretical foundations, please refer to the introduction tutorials.
The processing pipeline is published in this article:
Niso G, Tadel F, Bock E, Cousineau M, Santos A, Baillet S, Brainstorm Pipeline Analysis of Resting-State Data from the Open MEG Archive, Frontiers in Neuroscience, Mar 2019
Contents
OMEGA database
The Open MEG Archive (OMEGA) is the fruit of a collaborative effort by the McConnell Brain Imaging Centre and the Université de Montréal to build a centralized repository in which to regroup MEG data for open dissemination. This continuously expanding repository also contains anatomical MRI volumes, demographic and questionnaire information, and eventually will feature other forms of electrophysiological data (e.g. EEG, field, and cell recordings). OMEGA features the technological framework for multi-centric data aggregation, and is amongst the largest freely available resting-state MEG datasets presently available. Website: https://www.mcgill.ca/bic/resources/omega
You are free to use all data in OMEGA for research purposes; however, we ask that you please cite the following reference in your publications if you have used data from OMEGA:
Niso G, Rogers C, Moreau JT, Chen LY, Madjar C, Das S, Bock E, Tadel F, Evans A, Jolicoeur P, Baillet S
OMEGA: The Open MEG Archive, Neuroimage, 2015.
Presentation of the experiment
Experiment
- 5 subjects x 5 minute resting sessions, eyes open
MEG acquisition
Acquisition at 2400Hz, with a CTF 275 MEG system,
- Recorded at the Montreal Neurological Institute in 2015-2016.
- Anti-aliasing low-pass filter at 600Hz
- Files may be saved with or without the CTF 3rd order gradient compensation applied.
- Recorded channels (at least 297):
- 26 MEG reference sensors (#2-#27)
- 270 MEG axial gradiometers (#28-#297)
- 1 ECG bipolar (EEG057/#298) - Not available in the empty room recordings
- 1 vertical EOG bipolar (EEG058/#299) - Not available in the empty room recordings
- 1 horizontal EOG bipolar (EEG059/#300) - Not available in the empty room recordings
The data in this dataset has been organized according to the BIDS specifications
(Brain Imaging Data Structure, http://bids.neuroimaging.io)
Head shape and fiducial points
3D digitization using a Polhemus Fastrak device driven by Brainstorm. The .pos files contain:
- The center of the CTF coils,
The anatomical references we use in Brainstorm: nasion and ears as illustrated here,
- Around 100 head points distributed on the hard parts of the head (no soft tissues).
Subject anatomy
- Structural T1 image (defaced for anonymization purposes)
Processed with FreeSurfer 5.3
- The anatomical fiducials (NAS, LPA, RPA) have already been marked and saved in the files fiducials.m
Note that while the subjects shared here have their anatomy already processed with FreeSurfer to simplify the tutorial, this is not currently the case for data shared in OMEGA. You would therefore need to generate surfaces yourself as explained in the anatomy tutorial.
Download and installation
First, make sure you have at least 40Gb of free space on your hard drive (raw files: 12Gb).
- The OMEGA database is not in it its final release stage, so we temporarily distribute the 5 subjects used in this tutorial from the Brainstorm website.
The data is hosted on the OpenNeuro website (click on the icon "Download dataset" at the top):
https://openneuro.org/datasets/ds000247- The safest alternative is to use the Amazon AWS command line tools:
Install AWS CLI: https://aws.amazon.com/cli/ (no need to create an account)
- Linux/MacOS:
- In a terminal, type: pip install awscli
- Windows:
- Download and install the program (64bit version, most likely)
- Open a command line terminal (Windows menu, type "cmd", enter)
- Go to the AWSCLI install folder (type: cd "C:\Program Files\Amazon\AWSCLI")
- Linux/MacOS:
- Start the download (synchronization with an Amazon S3 repository):
Create a new folder "ds000247" somewhere on a large hard drive (NOT in the Brainstorm program or database folders), referred to below as <target_directory>.
- In the same terminal, type:
aws s3 sync --no-sign-request s3://openneuro.org/ds000247 <target_directory>
- Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder).
Start Brainstorm (Matlab scripts or stand-alone version). For help, see the Installation page.
Select the menu File > Create new protocol. Name it "TutorialOmega" and select the options:
"No, use individual anatomy",
"No, use one channel file per condition".
BIDS specification
The Brain Imaging Data Structure (BIDS) standard for neuroimaging data organization was first established for MRI and fMRI (Gorgolewski, 2016). It is based on simple file formats and folder structures that can readily expand to additional data modalities.
An extension of the BIDS for MEG datasets has now been published. The data distributed with this tutorial is organized following this prototype (which may change in the future). One objective is that analysis pipelines designed with major analysis tools (such as Brainstorm, FieldTrip, MNE, SPM and others) can be readily applied without requiring software or pipeline redevelopments.
Brainstorm includes a set of functions that can load automatically any dataset following the BIDS specification. The files that are read by Brainstorm are the following:
OMEGA_RestingState_sample/
derivatives/: Everything that cannot be considered as raw data
freesurfer/: Result of the FreeSurfer segmentation for all the subjects
sub-000X_ses-mri/anat/: Output of FreeSurfer for subject 000X (session 0001)
sub-000X/: Raw data for subject 000X
ses-0001/: Only one session per subject (the subject visited the center only once)
sub-0002_ses-0001_scans.tsv: BIDS tab-separated text file listing the MEG recordings and corresponding acquisition dates
anat/: Anatomical MRI scans for subject 000X (not used if FreeSurfer output available)
sub-000X_ses-0001_T1w.nii.gz: T1 scan
meg/: Unprocessed MEG recordings
sub-000X_ses-0001_task-rest_run-01_meg.ds: Resting MEG recordings
sub-emptyroom/: Empty room measures at different dates (used for noise modelling)
ses-XXX/: Various sessions of noise recordings (can be matched by date with the subjects' recordings using the *_scans.tsv BIDS files).
sub-emptyroom_ses-XXX_scans.tsv: BIDS tab-separated text file listing the MEG recordings and corresponding acquisition dates
meg/: Unprocessed MEG empty room recordings
sub-000X_ses-XXX_task-noise_run-01_meg.ds
Import the dataset
Select the menu File > Load protocol > Import BIDS dataset > Select folder OMEGA_RestingState_sample.
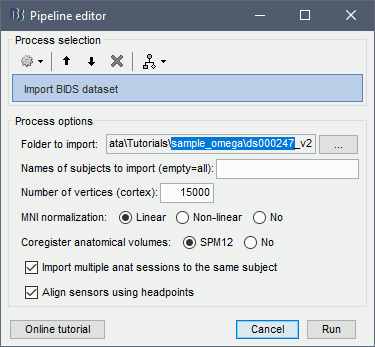
Folder to import: The root folder of the BIDS dataset to import.
Names of subjects: If you want to import only a few specific subjects from the dataset, list them here separated with commas. Otherwise, leaving the field empty imports all the subjects available in the dataset.
Number of vertices: If the segmentation of the MRI with FreeSurfer, CAT12, BrainSuite or BrainVISA is available in the derivatives folder, this process will import it just like it would do with the menu Import anatomy folder. Default fiducials positions are computed using the affine MNI normalization (maff8). This process includes downsampling the cortex surface to a specific number of vertices, which you can specify with this option.
MNI normalization: For the first volume imported into the database, when it is not part of a segmentation folder from the derivatives (FreeSurfer or other), computes the MNI normalization using the selected method.
Align sensors using headpoints: When importing EEG or MEG sensor definitions, if there are enough digitized head points, this option enables running automatically the process Refine registration with head points.
Import multiple anat sessions: If this option is not selected, a subject with multiple anat sessions would lead to the creation of multiple subjects in the Brainstorm database. When selected, all the anatomy volumes from multiple sessions are imported into the anatomy of the same subject.
Keep the default values for all the questions that may be asked during the import process (eg. number of vertices in the cortex surfaces). Once done, you should be able to access the data for the 5 subjects in your database explorer: anatomy, and subject and noise recordings.
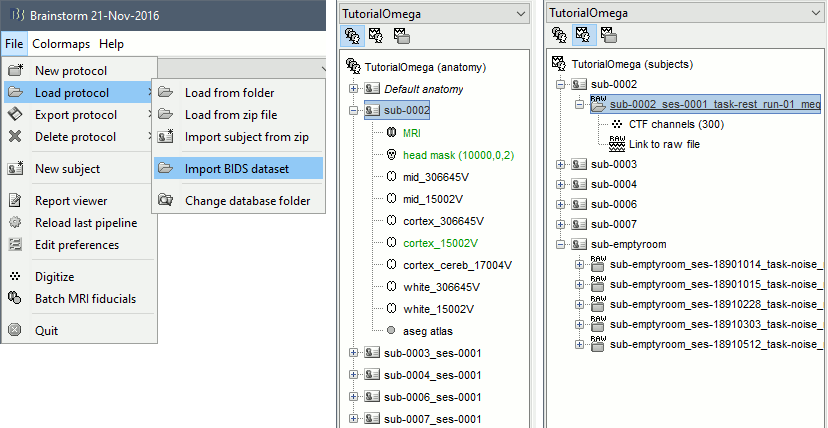
It is important to understand how the spatial coregistration between the anatomy (MRI and FreeSurfer surfaces) and the MEG sensors was done here, as it is not a trivial question. The two procedures available in Brainstorm for this registration are described in the tutorial Channel file & MRI registration: using three anatomical fiducials NAS/LPA/RPA and/or using the digitized head shape. In this dataset we do not have access to the positions of the fiducials in the MRI, however, the digitized head shapes (same coordinate system as the MEG) are available in the files *_headshape.pos. Therefore, we defined automatically approximate positions for the fiducials NAS/LPA/RPA in the MRI using standard MNI coordinates, with the expectation that this approximate registration would later be refined using the head shape. This is done in the next section.
Another solution could have been to first mark manually the positions of the anatomical landmarks using the MRI viewer in Brainstorm. The menu File > Batch MRI fiducials opens the MRI viewer for each subject, asks for marking the NAS/LPA/RPA landmarks, and creates a file fiducials.m in the FreeSurfer folder. When importing the FreeSurfer folders, these fiducials are used by default.
TODO: The BIDS-MEG standards are still in development and do not include yet a final solution for saving these anatomical fiducials in the BIDS data structure. When this feature is implemented, we will add the corresponding fields in this example dataset and will update the import function to read the correct fiducials positions automatically.
Refine the MEG-MRI registration
- At this point, the registration MEG/MRI is based only on the three approximate anatomical landmarks NAS/LPA/RPA (standard positions from the MNI template) and must improved. All the MRI scans were anonymized (defaced) so all the head points below the nasion cannot be used. We will try to refine this registration using the additional head points that were digitized above the nasion.
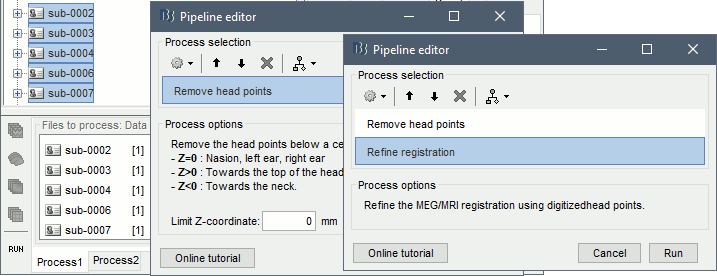
- Drag and drop the rest recordings for all the subjects (exluding sub-emptyroom) into the Process1 box, and click on [Run].
Select process: Import > Import anatomy > Remove head points, Z=0
Add process: Import > Channel file > Refine registration, then run the execution.
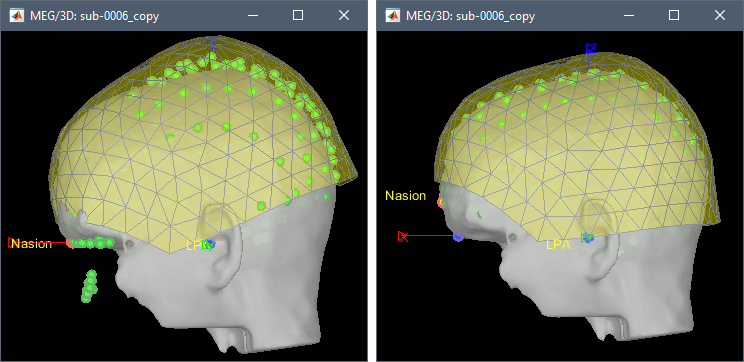
Check the results for one subject: right-click on CTF channels > MRI registration > Check.
Pre-processing
- Switch to the functional view of the protocol (second button above the database explorer).
- Drag and drop all the subjects (including sub-emptyroom) into the Process1 box, and click on [Run].
Select process: Import > Import recordings > Convert to continuous (CTF): Continuous
Add process: Frequency > Power spectrum density (Welch): All file, 4s, 50% overlap, Individual
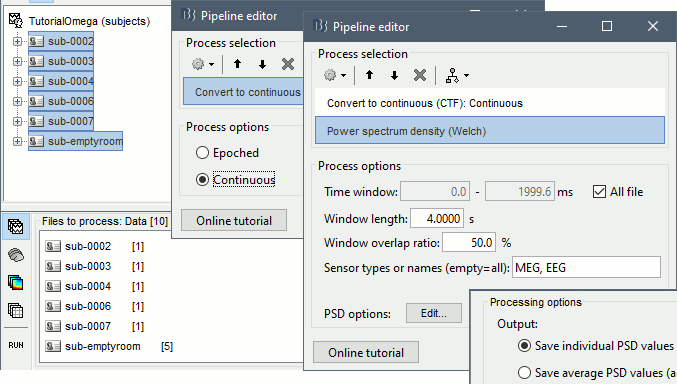
Run, then check the quality of the PSD for all the files, as documented in this tutorial.
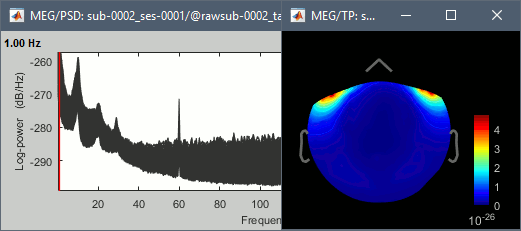
- In Process1, keep the same files selected and click on [Run].
Select process: Pre-process > Notch filter: 60 120 180 240 300 Hz, Process the entire file at once
Add process: Pre-process > Band-pass filter: High-pass filter at 0.3Hz, 60dB, Process entire file.
Add process: Frequency > Power spectrum density (Welch): Same options
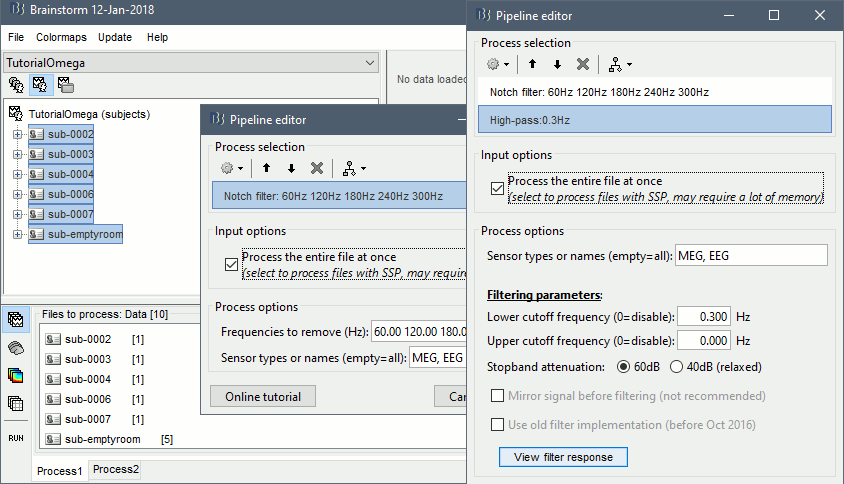
Run, then check the PSD after filtering.
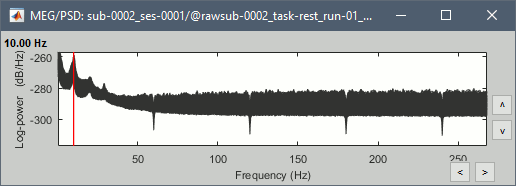
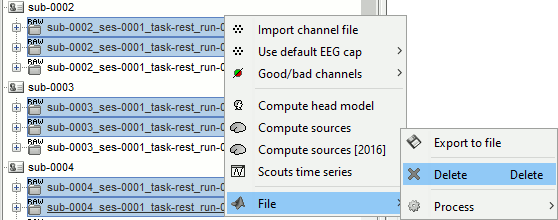
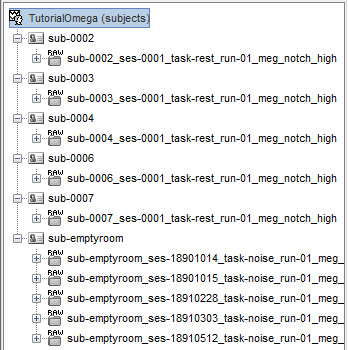
Delete all the folders corresponding to the original recordings (_meg) and the notch filtered data (_meg_notch). Only keep the fully processed files (_meg_notch_high).
Artifact cleaning
We will now run the automatic procedure for cleaning the heartbeats, as described in the introduction tutorials (detection, SSP). The results we want to illustrate here are robust enough, the recordings do not need to be processed any further. If you want to improve the quality of the data with more manual cleaning (blinks, saccades, movements, bad segments), please refer to the introduction tutorials.
- In Process1, select all the subject rest recordings (all the subjects, excluding sub-emptyroom).
Select process: Events > Detect heartbeats: ECG, All file, cardiac
Add process: Artifacts > SSP: Heartbeats: cardiac, MEG, Use existing SSP
Add process: File > Snapshot: Sensor/MRI registration
Add process: File > Snapshot: SSP projectors
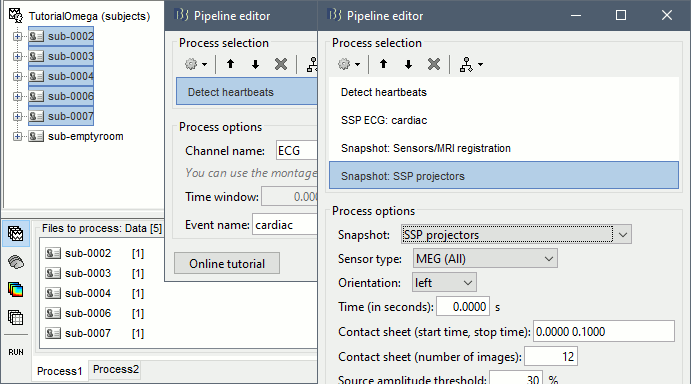
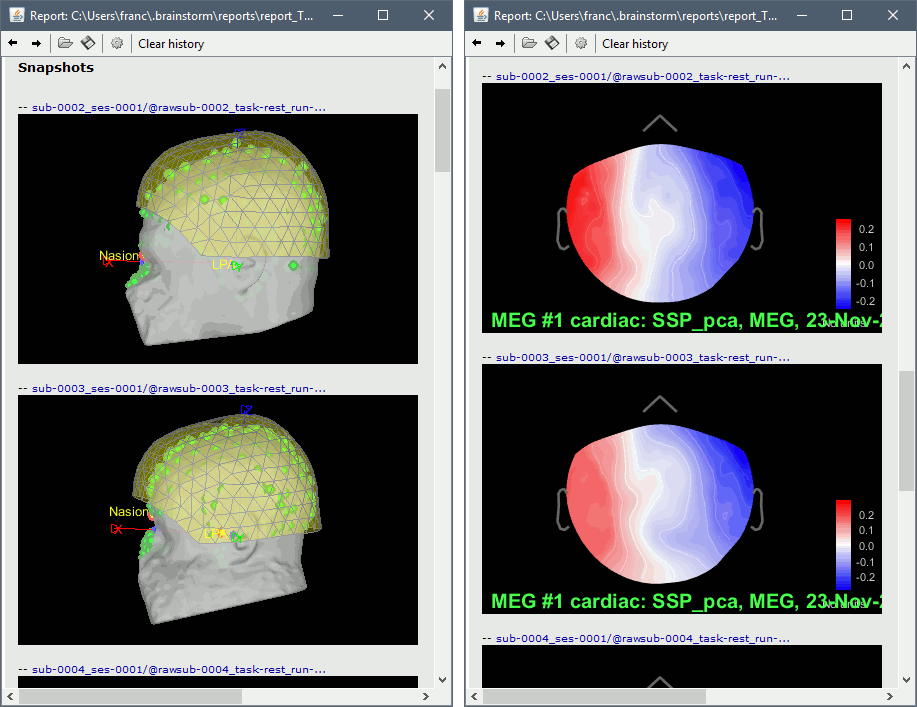
Run, then review the registration and the SSP topographies for all the subjects. Make sure that the subjects' heads are well aligned in the MEG, and that the spatial components that are removed are really representative of the cardiac artifact.
Source estimation
- In Process1, select all the noise recordings (all the recordings in sub-emptyroom).
Run process: Sources > Compute covariance: All file, Noise covariance, Copy to other folders, Copy to other subjects, Match by acquisition date.
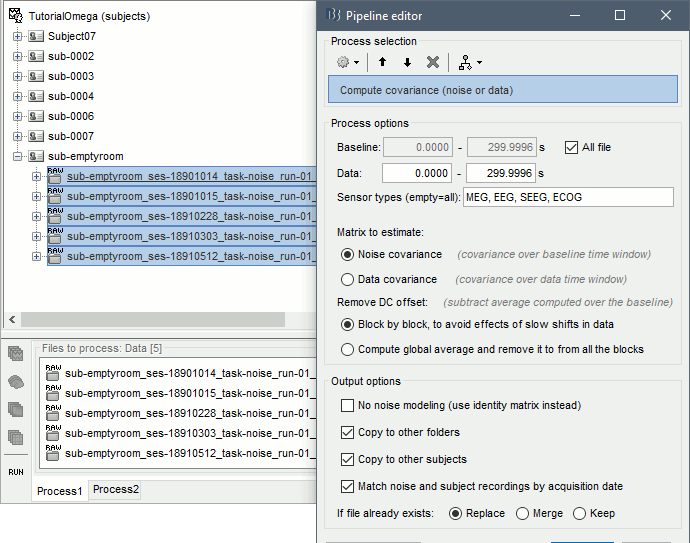
The option Match noise and subject recordings by acquisition date uses the recordings dates read from the *_scans.tsv files in the BIDS dataset to copy in each rest recording session the noise covariance corresponding to its acquisition date. The empty room recordings with the closest acquisition date (in number of days, before or after) is used to estimate the noise covariance for each subject.
- In Process1, select all the subject rest recordings (all the subjects, excluding sub-emptyroom).
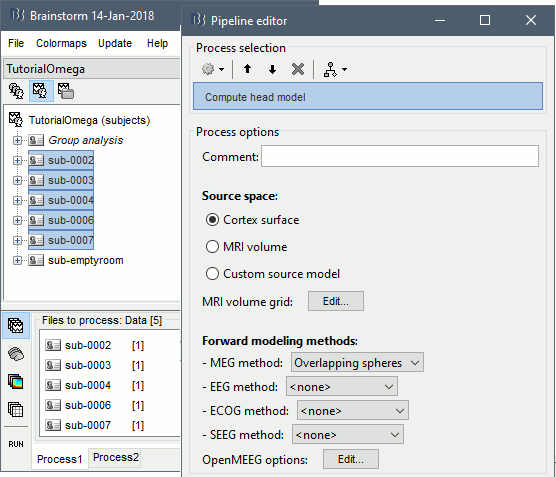
Run process: Sources > Compute head model: Cortex surface, MEG=Overlapping spheres.
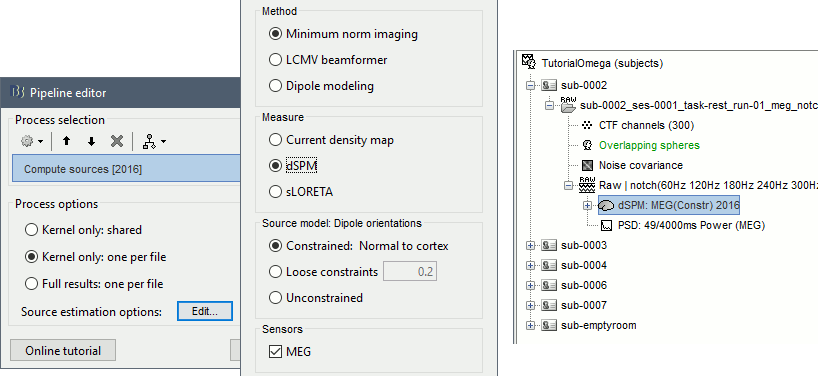
Run process: Sources > Compute sources [2016]: Kernel only, one per file, dSPM, constrained.
Power maps
We have now access to continuous recordings in source space, we can estimate the average power over this resting period, for various frequency bands. We propose here to compute the PSD from the first 100s instead of the full 600s rest recordings, it is faster and leads to very similar results. For more stable estimates, you can compute the the PSD over all the data available instead.
In Process1, select all the subject rest recordings (task-rest), click the [Process sources] button.
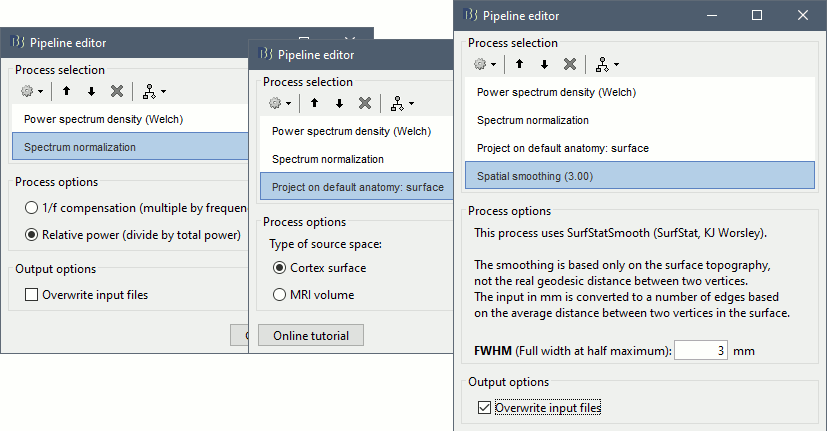
Select process: Frequency > Power spectrum density (Welch): [0,100s], Window=4s, 50% overlap, Group in frequency bands (use the default frequency bands), Save individual PSD values.
Add process: Standardize > Spectrum normalization: Relative power (divide by total power). This normalization is done independently for each source, the output for each frequency band is a percentage of how much this frequency band contributes to the total power of the source signal.
Add process: Sources > Project on default anatomy: Cortex.
Add process: Sources > Spatial smoothing: FWHM=3mm, Overwrite.
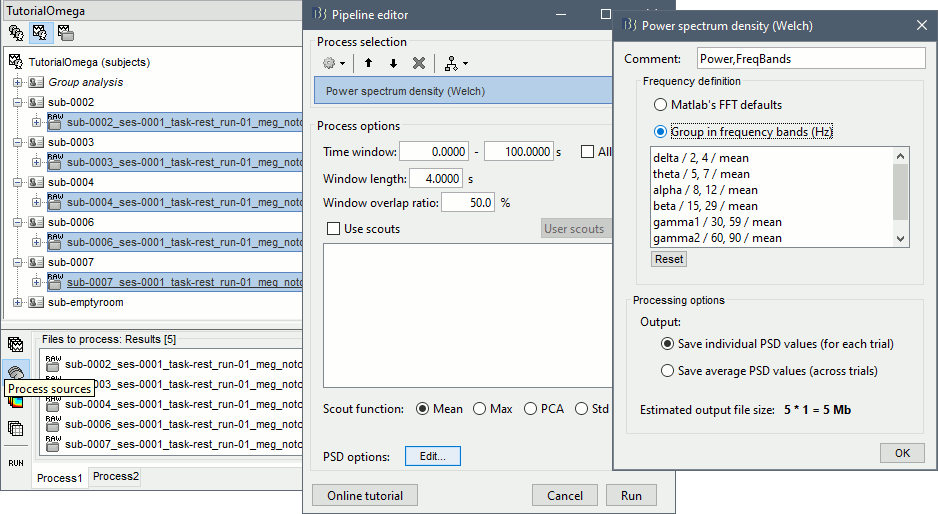
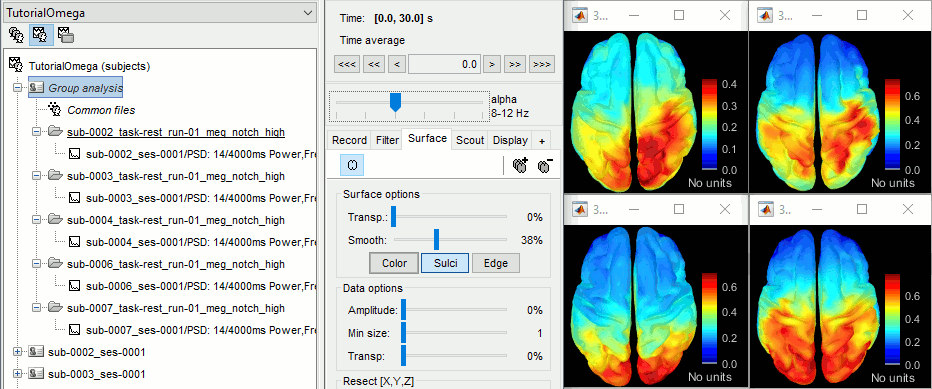
- Execute this pipeline, then open the results for the different subjects and visualize the different frequency bands. They should look similar across subjects. This measure of relative power is very robust and usually do not vary much between subjects or duration of the input signals.
The values in this figure represent the relative power of each source. At each vertex of the cortex surface, the value between 0 and 1 indicates what is the contribution of the current frequency band to the total power in the signal. To obtain comparable colormaps across multiple figures, you can set a fixed maximum (right-click on a figure > Colormap: Timefreq > Maximum: Custom).
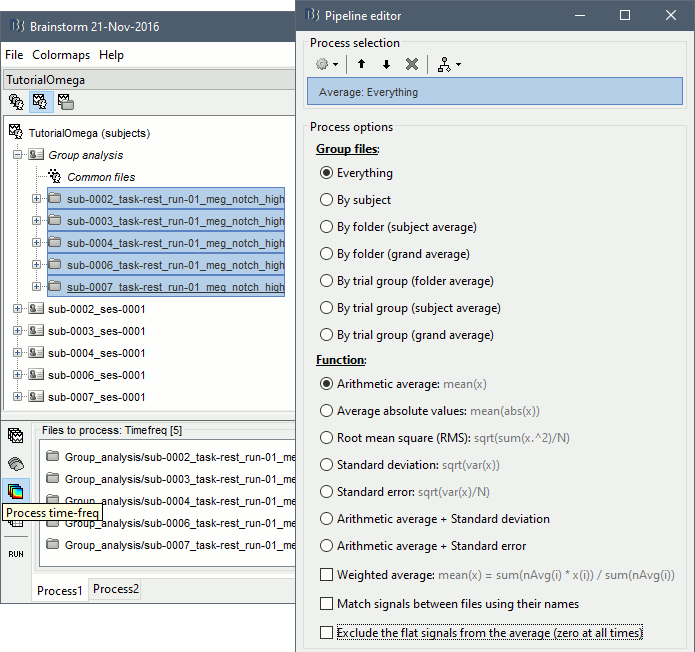
We will now average these files across subjects. In Process1, select all the projected subject averages in the Group analysis folder, and click on the button [Process time-freq].
Run process: Average > Average files: Everything, Arithmetic average, Do not match signals
Double-click on the average to display it. To create the figures below, right-click on the figure > Snapshot > Frequency contact sheet.
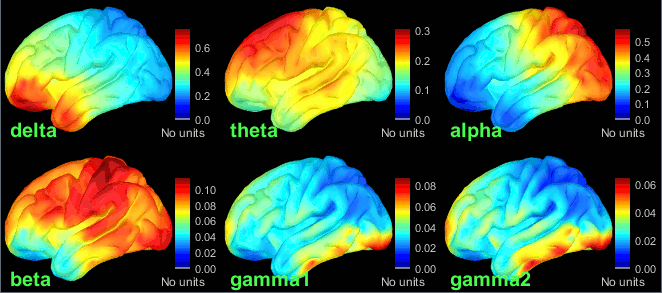
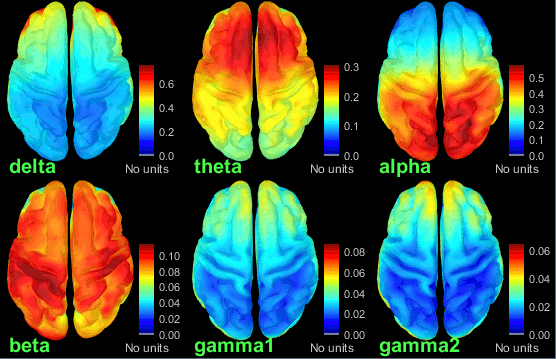
- We can relate these figures with associations commonly reported in the literature:
Theta: Mostly frontal.
Alpha: Parieto-occipital.
Beta: Central regions (motor cortex).
Gamma: Pre-frontal and occipital. However, the occipital sources could be related with muscle activity in the neck. We haven't cleaned the recordings much, we should go back to the recordings and mark as bad all the segments in which we observe a lot of muscle activity.
Delta: Questionable - these source maps showing high values at the poles of the frontal and temporal lobes are typical of eye movements. We should try doing some cleaning of the ocular artifacts (blinks and saccades) and compute these maps again.
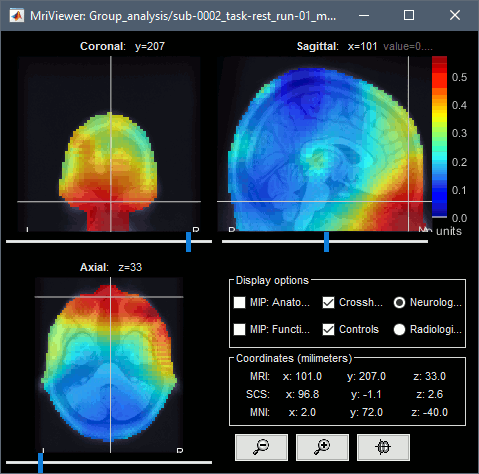
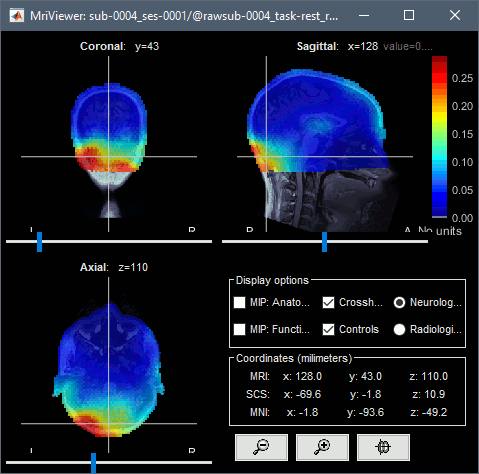
You can compute the same results in the head volume. You just have to use a volume head model on a group grid instead of the one we used here. This is explained in the tutorial Group analysis. This seems to confirm that the power in the delta and gamma bands is mostly coming from non-brain regions (eyes and neck muscles).
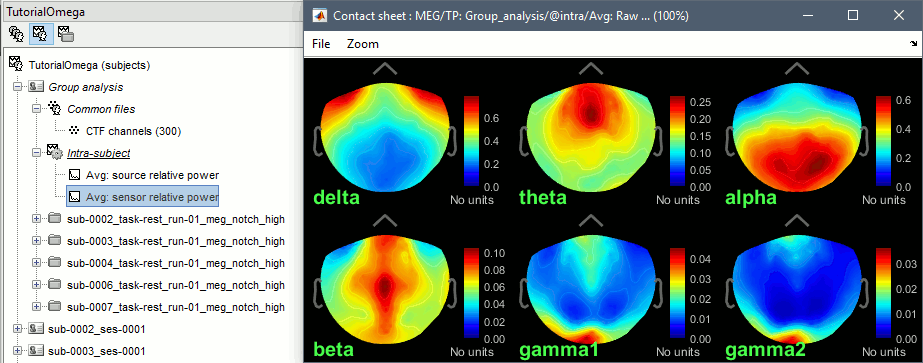
You can produce similar plots at the sensor level: Compute the PSD by frequency band of the rest recordings for each subject, then compute the average across subjects (not as reliable as averaging in source space, as explained here).
Scripting
The following script from the Brainstorm distribution reproduces the analysis presented in this tutorial page: brainstorm3/toolbox/script/tutorial_omega.m
Additional documentation
Articles
BIDS: Gorgolewski, K.J., Auer, T., Calhoun, V.D., Craddock, R.C., Das, S., Duff, E.P., Flandin, G., Ghosh, S.S., Glatard, T., Halchenko, Y.O. and Handwerker, D.A. (2016), The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments, Scientific Data, 3, p.160044.
MEG-BIDS: Niso G, Flandin G, Gramfort, A Henson, RN, Jas M, Litvak V, Moreau JT, Oostenveld R, SchoffelenJ-M, Tadel F, Wexler J, Baillet S (2018), MEG-BIDS, the brain imaging data structure extended to magnetoencephalography, Scientific Data, 5: 180110 EP
EEG-BIDS: Pernet CR, Appelhoff S, Flandin G, Phillips C, Delorme A, Oostenveld R (2018), BIDS-EEG: an extension to the Brain Imaging Data Structure (BIDS) Specification for electroencephalography, PsyArXiv doi:10.31234/osf.io/63a4y
Forum
Normalize PSD based on resting-state before statistical tests: https://neuroimage.usc.edu/forums/t/psd-normalization-based-on-the-resting-psd/37854
