|
Size: 7279
Comment:
|
Size: 15009
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Digitize EEG Sensors and Head Shape = | = Digitize EEG sensor locations and head shape = ''Authors: Elizabeth Bock'' <<TableOfContents(2,2)>> |
| Line 5: | Line 9: |
| The connection to the device is made via a serial port connection. This can be accomplished using a direct serial connection or with a USB-to-Serial cable connection to the host computer. {{attachment:polhemus.png|75px"}} |
'''The application is designed for two receivers (i.e. the stylus and reference) and one transmitter. ''' The connection to the device is made via a ''serial port connection''. This can be accomplished using a direct serial connection or with a ''USB-to-Serial'' cable connection to the host computer. == Configuration of the Polhemus == === Fastrak or Isotrak === * These devices operate at 9600 baud, 8 bits, no parity * Configure the unit using the switches at the back of the device: '''11001001<<BR>>'''(9600 baud, 8 bits, parity:none, rs232) * Configure the computer's COM port for the same * Select ''fastrak'' for Unit Type in the GUI settings (see below) === Patriot === * This device operates at 115200 baud, 8 bits, no parity * Configure the unit using the switches at the back of the device (all down) * Configure the computer's COM port for the same * Select ''patriot'' for Unit Type in the GUI settings (see below) {{attachment:polhemus.png||width="200"}} |
| Line 11: | Line 30: |
| {{attachment:typical_polhemus_setup.png|75px}} | {{attachment:typical_polhemus_setup.png||width="300"}} |
| Line 15: | Line 34: |
| The application is designed for two receivers (i.e. the stylus and reference) one transmitter. | '''The application is designed for two receivers (i.e. the stylus and reference) and one transmitter. ''' |
| Line 29: | Line 48: |
| The head coordinate system used in the Digitize application is based on the CTF head coordinate system, see /CoordinateSystems. | The head coordinate system used in the Digitize application is based on the CTF head coordinate system, see CoordinateSystems. == The Digitize GUI == ''' Before starting the GUI, turn on the Polhemus unit and wait until the indicator light is green.''' |
| Line 33: | Line 55: |
| {{attachment:bst_menu_digitize.png|75px}} | {{attachment:bst_menu_digitize.png||width="320"}} |
| Line 39: | Line 61: |
| == The Digitize GUI == {{attachment:gui_noeeg_digitize.png}} Figure 5. Digitize GUI |
{{attachment:panel_digitize.png||width="400"}} Figure 5. Digitize GUI |
| Line 44: | Line 67: |
| * Three anatomical landmarks for computing the head coordinate system | * Three head localization coils (HPI) for computing the head coordinate system * Three anatomical landmarks for MEG/MRI co-registration |
| Line 51: | Line 75: |
| * Two operators, which one will point the stylus at the desired location and one will click the ‘Collect Point’ button on the lower panel of the GUI. * Points are collected in order from Nasion, Left , Right, EEG (if desired), then head shape. |
* Two operators, where one will point the stylus at the desired location and one will click the ‘Collect Point’ button on the lower panel of the GUI. * Points are collected in order from HPI-N, HPI-L, HPI-R, Nasion, Left , Right, EEG (if desired), then head shape. |
| Line 64: | Line 88: |
| {{attachment:gui_editsettings_digitize.png}} Figure 6. Edit Settings {{attachment:edit_settings_digitize.png}} Figure 7. Edit settings dialog |
{{attachment:gui_editsettings_digitize.png||width="400"}} Figure 6. Edit Settings {{attachment:edit_settings_unittype.png||width="250"}} Figure 7. Edit settings dialog |
| Line 70: | Line 98: |
| '''Baud rate''' – this the baud rate of the connection. The default is 9600. '''Byte count''' – this is the number of bytes that will be collected for each point and sent to the host computer. '''Number of times to collect the three fiducials''' – these collections will occur at the beginning of the session and will be averaged together to compute the transformation to the head coordinate system. |
'''Unit Name''' – this will select the settings for the specific device. Fastrak and Isotrak will both use the name ''fastrak''. '''Digitize MEG HPI coils''' - this will allow for collection of the positions of the HPI coils. This is important for CTF users since the data collected from the MEG system is saved in HPI coil coordinate system. Users collecting only EEG sensors or just head shape will not collect these points. '''Number of times to collect the three fiducials''' – these collections will occur at the beginning of the session and will be averaged together to compute the transformation to the HPI or head coordinate system. |
| Line 79: | Line 107: |
| {{attachment:eeg_settings_digitize.png}} Figure 8. Edit EEG settings | {{attachment:eeg_settings_digitize.png||width="400"}} Figure 8. Edit EEG settings |
| Line 87: | Line 117: |
| {{attachment:eegmontage_example_digitize.png}} Figure 9. Example EEG montage | {{attachment:eegmontage_example_digitize.png||width="150"}} . Figure 9. Example EEG montage |
| Line 93: | Line 125: |
| {{attachment:example_eeg_digitize.png}} Figure 10. Example EEG setups | {{attachment:example_eeg_digitize.png||width="640"}} Figure 10. Example EEG setups |
| Line 96: | Line 130: |
| === Collect the required number of HPI coils === * When collecting more than one set of fiducials, an error message is displayed if the difference between the sets is > 5mm * Once collected, the head coordinate system is computed and the points are plotted on the 3D point display overlayed on the scalp surface. Unless the user changes the anatomy for the brainstorm Digitize subject, the default anatomy is used. It may not match exactly the subject being digitized, but should be a good quality control image to ensure the points are positioned where expected. * Note that the 3D figure is a standard Brainstorm window, where you can use all the mouse operations (rotation, zoom, popup menu) and the keyboard shortcuts (predefined view, screen capture...) introduced in the previous tutorials. * The coordinates are displayed in the coordinate list in cm * The points are saved in the brainstorm channel file as HPI Head Points * The collection automatically moves to Anatomical Fiducials {{attachment:HPI_quality_check.png||width="300"}} Figure 11. Collect coils |
|
| Line 97: | Line 143: |
| * When collecting more than one set of fiducials, an error message is displayed if the difference between the sets is > 2mm * Once collected, the head coordinate system is computed and the points are plotted on the 3D point display |
* When collecting more than one set of fiducials, an error message is displayed if the difference between the sets is > 5mm * If the MEG HPI coils are not used, then the head coordinate system is computed with the anatomical fiducials and the points are plotted on the 3D point display. Unless the user changes the anatomy for the brainstorm Digitize subject, the default anatomy is used. It may not match exactly the subject being digitized, but should be a good quality control image to ensure the points are positioned where expected. * Note that the 3D figure is a standard Brainstorm window, where you can use all the mouse operations (rotation, zoom, popup menu) and the keyboard shortcuts (predefined view, screen capture...) introduced in the previous tutorials. |
| Line 103: | Line 150: |
| {{attachment:error_fiducials_digitize.png}} Figure 11. Collect fiducials, error message | {{attachment:HPI_FIDS_quality_check.png||width="300"}} Figure 12. Collect fiducials |
| Line 111: | Line 160: |
| {{attachment:eeg_collection_digitize.png}} Figure 12. Collect EEG |
|
| Line 114: | Line 161: |
| * In order to take full advantage of the MEG/MRI co-registration algorithms in Brainstorm, it is recommended to collect approximately 100-150 points from the head: 5-10 points from the boney part of the nose, 10-15 points across the eyebrows, then an even coverage of the scalp from the eyebrows to the inion. It is NOT recommended to collect points below the inion on the neck, since the shape of the neck can change between lying in the MRI scanner and sitting during point collection. Even coverage is more important that dense coverage. See Figure 13 as an example. | |
| Line 118: | Line 166: |
| * Once complete, the operator can save the points to the CTF *.pos file with the ‘Save as POS’ button or select ‘Save and exit’ from the File menu {{attachment:headshape_collection_digitize.png}} Figure 13. Collect head shape |
* Once complete, the operator can save the points to a text file in a variety of formats. {{attachment:final_shape.png||width="540"}} Figure 13. Collect head shape === Save === * The points can be saved as a POS file for import into CTF datasets or your Brainstorm database. Select 'Save as...' and confirm the file name and location. * When exiting the program, select File > Save and Exit to ensure the points are saved in the brainstorm database. |
| Line 123: | Line 177: |
| All points are automatically saved in the brainstorm channel file | All points are automatically saved in a brainstorm channel file |
| Line 128: | Line 182: |
| {{attachment:bst_saveas_digitize.png}} Figure 14. Export point to text file | {{attachment:bst_saveas_digitize.PNG||width="300"}} Figure 14. Export points to text file == How to use this file with CTF systems == The procedure we recommend is the following: * Export the file in the format "EEG+headshape: Polhemus (*.ds)" * Copy the .pos file to all the CTF datasets recorder for this subject (.ds folder) * When importing the CTF datasets in Brainstorm, the file will be automatically loaded. == Troubleshooting == There are several things that can affect the quality and reproducibility of the measurements from the Polhemus Fastrak: === Device communication === Problem appears to be with the communication between the device and Brainstorm: 1. Power on the device 1. Restart Matlab 1. Start brainstorm 1. Start Digitize 1. At the Matlab command prompt, type this: s = instrfind() If you do not have Matlab, you can open a command prompt from the Brainstorm File menu BEFORE starting Digitize. The command, s = instrfind() should be typed into the window, click execute and see the output in the terminal window.: {{attachment:start_command_window.jpg||width="340"}} Figure 15: Start command window from Brainstorm {{attachment:serial_start_digitize.jpg||width="340"}} Figure 16: Serial connection settings (Fastrak) 1. From the Digitize->File->Edit settings (Figure 7), confirm the information in 's' matches your settings 1. Confirm the information in 's' matches your COM port settings of your computer (this info is in the Device Manager) 1. Press the button of the stylus, then execute s = instrfind() again. You should see that there are bytes available (94 bytes for Fastrak, 120 bytes for Patriot). {{attachment:serial_collectpoint_digitize.jpg||width="340"}} Figure 17: Serial connection after one point collected (Fastrak) === Accuracy === * It is very sensitive to it's environment. It can take many repeated tests, moving the phantom around the room, to find the quietest spot where the errors between measurements are a minimum. You can do this simply with Matlab to collect points so that you skip the Brainstorm step which transforms the anatomical points to headspace. * In our experience, there is an ideal distance and relationship between the mounted receiver and the transmitter. For instance, when measuring a short person or child, the setup may not work and you may need to use a booster on the chair. Try different heights and distances. For our setup, this is approximately like Figure 2 (~40cm up and ~30cm forward from the transmitter to the mounted receiver). * The orientation of the stylus should always be perpendicular to the scalp surface. There is some compensation for the orientation of the stylus as part of the brainstorm application, but errors can be introduced if the collection is not consistent. * Be very careful not to move the mounted receiver during collection (or between collections when comparing the two). * As for the orientation of the transmitter, this may be trial and error depending on the settings of your device. Care should be taken to ensure the orientation of coordinate system is correct after point collection. == Additional documentation == * Forum: [[http://neuroimage.usc.edu/forums/showthread.php?1663|CTF coordinates and .pos files]] * Forum: [[http://neuroimage.usc.edu/forums/showthread.php?1769|Stylus orientation issues]] * Forum: [[http://neuroimage.usc.edu/forums/showthread.php?2100-co-registration-evaluation-and-Polhemus-vs-Vit-E|Polhemus headshape vs. vitamin E markers]] * Forum: [[http://neuroimage.usc.edu/forums/showthread.php?2111-Another-Head-Digitizer-Port-Problem|Debugging the Polhemus driver]] * Forum: [[https://neuroimage.usc.edu/forums/t/inaccurate-eeg-digitization-electrode-position/35319|Emitter position]] * Forum: [[https://neuroimage.usc.edu/forums/t/stylus-button-not-working-on-fastrak-digitizer/39204|Stylus button not working]] <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/TutDigitize)>> |
Digitize EEG sensor locations and head shape
Authors: Elizabeth Bock
Contents
Getting Started
The Brainstorm Digitize application allows for a connection to a Polhemus device and collection of digitized head points and/or EEG sensor locations.
The application is designed for two receivers (i.e. the stylus and reference) and one transmitter.
The connection to the device is made via a serial port connection. This can be accomplished using a direct serial connection or with a USB-to-Serial cable connection to the host computer.
Configuration of the Polhemus
Fastrak or Isotrak
- These devices operate at 9600 baud, 8 bits, no parity
Configure the unit using the switches at the back of the device: 11001001
(9600 baud, 8 bits, parity:none, rs232)- Configure the computer's COM port for the same
Select fastrak for Unit Type in the GUI settings (see below)
Patriot
- This device operates at 115200 baud, 8 bits, no parity
- Configure the unit using the switches at the back of the device (all down)
- Configure the computer's COM port for the same
Select patriot for Unit Type in the GUI settings (see below)
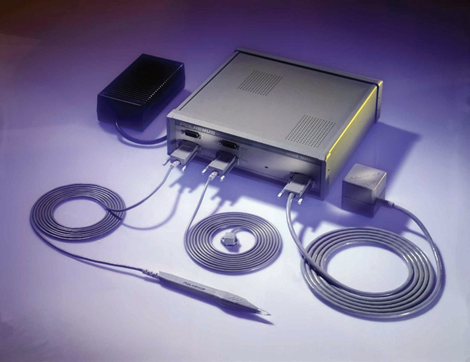

Figure 1. Polhemus system with two receivers and one transmitter
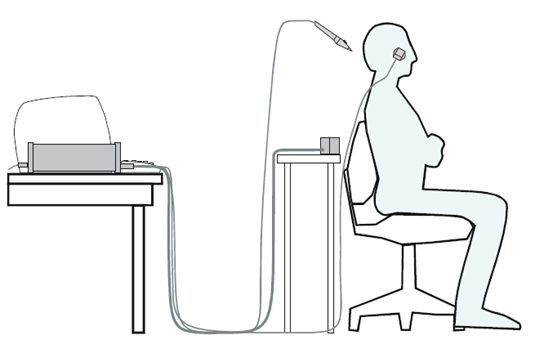
Figure 2. Typical Polhemus setup
The application is designed for two receivers (i.e. the stylus and reference) and one transmitter.
The stylus is used by the operator to point to the desired position for collection
The reference is fixed to the subject’s head, typically with glasses or headband. While the subject’s head can move during the collection, the reference sensor should not move from the starting position relative to the head.
The transmitter is fixed behind the subject, typically to a non-metal chair or table. The orientation is shown here, with X+ pointing up and Y+ point to the right.

Figure 3. Transmitter orientation
****Care should be taken when setting up this system to ensure accurate collection. Thorough testing should be performed using known head coordinates (phantom) to determine accuracy.
The head coordinate system used in the Digitize application is based on the CTF head coordinate system, see CoordinateSystems.
The Digitize GUI
Before starting the GUI, turn on the Polhemus unit and wait until the indicator light is green.
To start the Digitize application, start Brainstorm. Then select ‘Digitize’ from the File Menu.
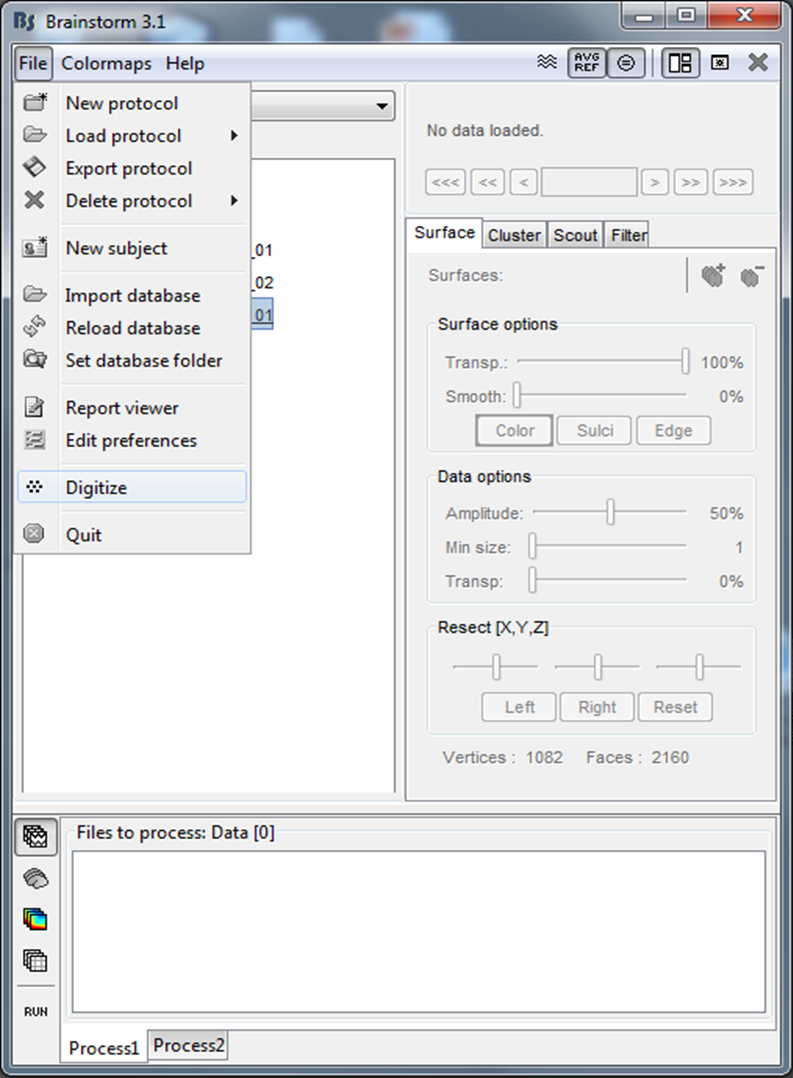
Figure 4. Starting Digitize
The Brainstorm GUI will be hidden during the use of the Digitize application and will reappear after exiting Digitize.
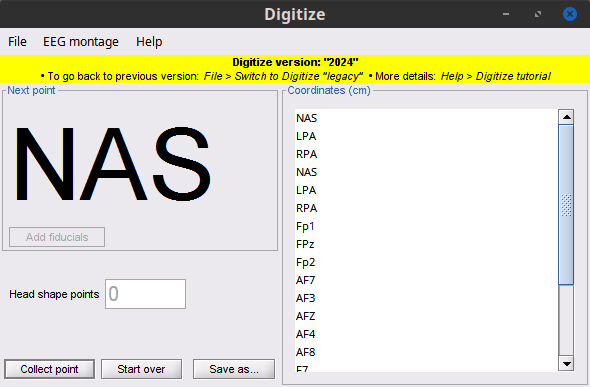
Figure 5. Digitize GUI
This application allows you collect:
- Three head localization coils (HPI) for computing the head coordinate system
- Three anatomical landmarks for MEG/MRI co-registration
- EEG sensor locations
- Scalp points for head shape
The collection can performed by:
- One operator who will press the button on the stylus to collect a point
- Two operators, where one will point the stylus at the desired location and one will click the ‘Collect Point’ button on the lower panel of the GUI.
- Points are collected in order from HPI-N, HPI-L, HPI-R, Nasion, Left , Right, EEG (if desired), then head shape.
If an error is made during collection:
- The last point can be removed by clicking “Delete Last Point” on the lower panel of the GUI
- The entire session can be restarted by selecting ‘Start Over’ from the File menu
When complete the coordinates can be save in a variety of formats including CTF (*.pos), megDraw (*.eeg) or tab-delimited ASCII text.
Edit Settings
Settings for the serial connection and point collection can be adjusted using the ‘Edit settings’ menu option from the Digitize File Menu and may vary for each user.
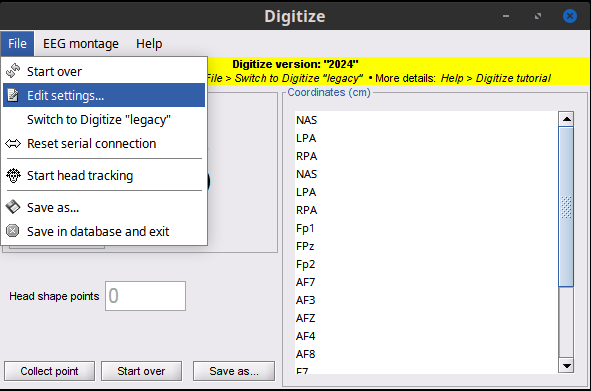
Figure 6. Edit Settings
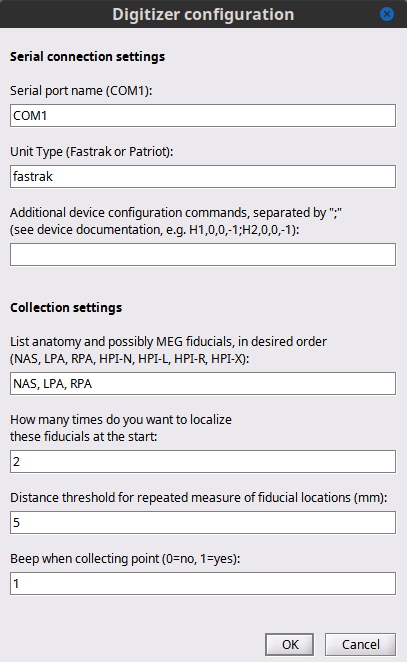

Figure 7. Edit settings dialog
Serial port name – this is the name of the serial port that is connected to the Polhemus, for example COM1 (Windows), ttyS1 (Linux).
Unit Name – this will select the settings for the specific device. Fastrak and Isotrak will both use the name fastrak.
Digitize MEG HPI coils - this will allow for collection of the positions of the HPI coils. This is important for CTF users since the data collected from the MEG system is saved in HPI coil coordinate system. Users collecting only EEG sensors or just head shape will not collect these points.
Number of times to collect the three fiducials – these collections will occur at the beginning of the session and will be averaged together to compute the transformation to the HPI or head coordinate system.
Beep – this will generate a beep sound with each point collection (this is often helpful to realize a button press on the stylus when there is one operator).
EEG Montages
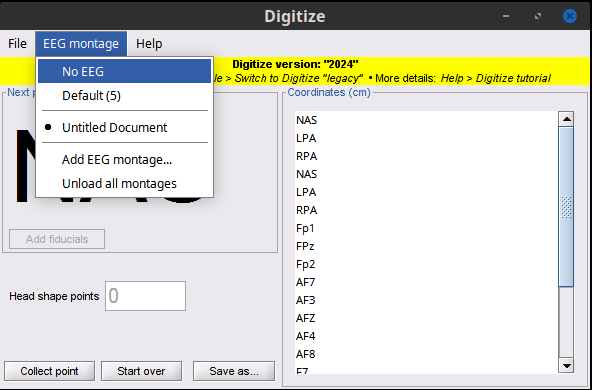
Figure 8. Edit EEG settings
There are three options for EEG sensors:
- No EGG: No collection and therefore no labels
- Default: This is entered as a number and the default labels are used: EEG01, EEG02, EEG03, etc.
- Custom Montage: A custom number of points is collected and the specified labels are used. This should be defined in a tab delimited plain text file as shown.
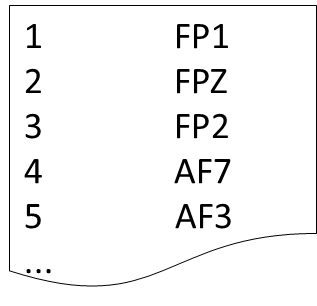
- Figure 9. Example EEG montage
EEG settings should be adjusted for each session. Custom montages that are imported are stored in the brainstorm preferences and are available on subsequent sessions.
The Coordinates list on the left panel of the GUI displays the labels of the points which have been selected for collection. This is vary depending on the number of times the fiducials are collected and the number of EEG sensors to be defined. The head shape points are not listed since this will vary depending on the individual need.
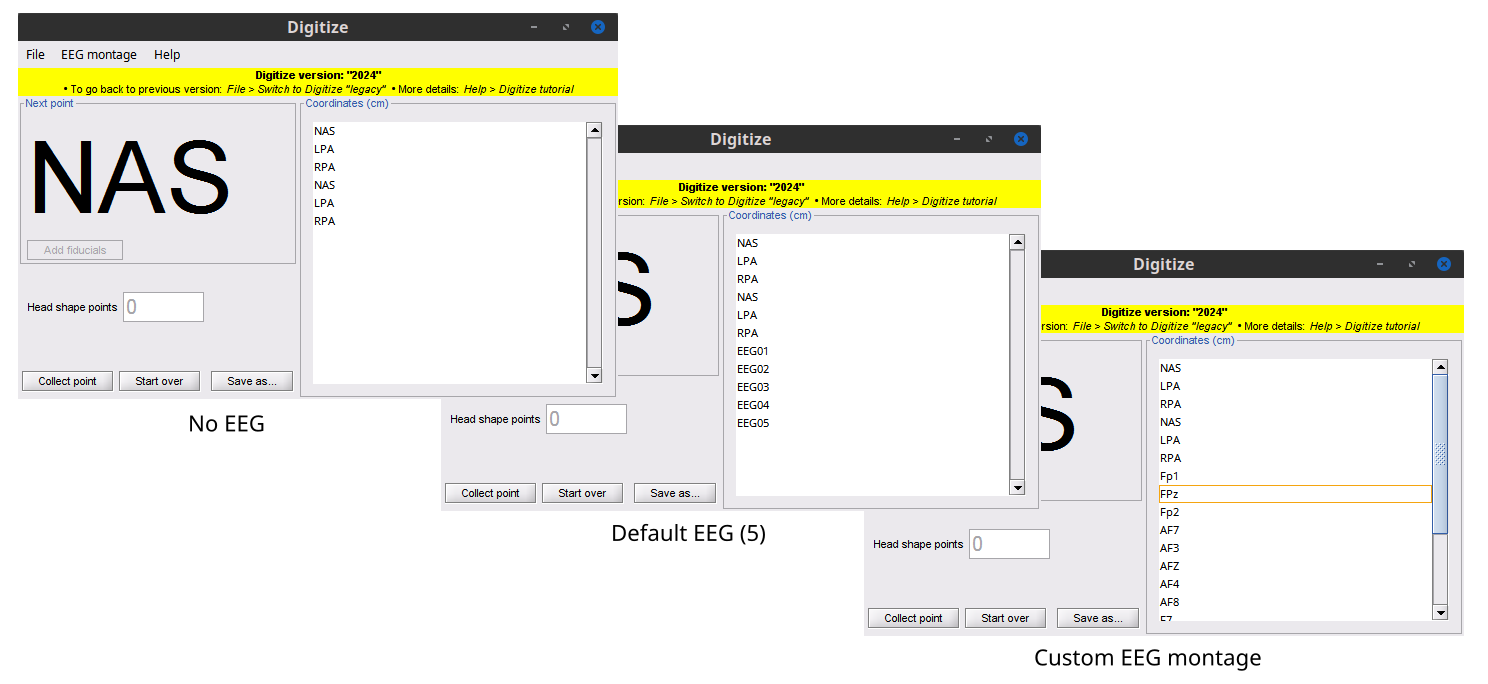
Figure 10. Example EEG setups
Collection
Collect the required number of HPI coils
When collecting more than one set of fiducials, an error message is displayed if the difference between the sets is > 5mm
- Once collected, the head coordinate system is computed and the points are plotted on the 3D point display overlayed on the scalp surface. Unless the user changes the anatomy for the brainstorm Digitize subject, the default anatomy is used. It may not match exactly the subject being digitized, but should be a good quality control image to ensure the points are positioned where expected.
- Note that the 3D figure is a standard Brainstorm window, where you can use all the mouse operations (rotation, zoom, popup menu) and the keyboard shortcuts (predefined view, screen capture...) introduced in the previous tutorials.
- The coordinates are displayed in the coordinate list in cm
- The points are saved in the brainstorm channel file as HPI Head Points
- The collection automatically moves to Anatomical Fiducials
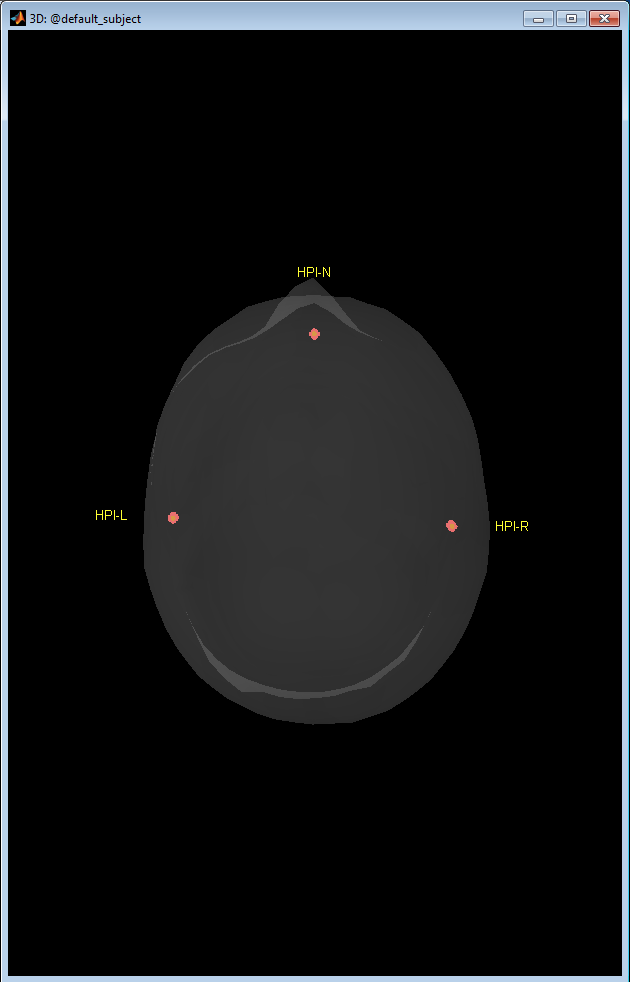
Figure 11. Collect coils
Collect the required number of Anatomical Fiducials
When collecting more than one set of fiducials, an error message is displayed if the difference between the sets is > 5mm
- If the MEG HPI coils are not used, then the head coordinate system is computed with the anatomical fiducials and the points are plotted on the 3D point display. Unless the user changes the anatomy for the brainstorm Digitize subject, the default anatomy is used. It may not match exactly the subject being digitized, but should be a good quality control image to ensure the points are positioned where expected.
- Note that the 3D figure is a standard Brainstorm window, where you can use all the mouse operations (rotation, zoom, popup menu) and the keyboard shortcuts (predefined view, screen capture...) introduced in the previous tutorials.
- The coordinates are displayed in the coordinate list in cm
- The points are saved in the brainstorm channel file as CARDINAL Head Points
- The collection automatically moves to EEG (if desired) or head shape
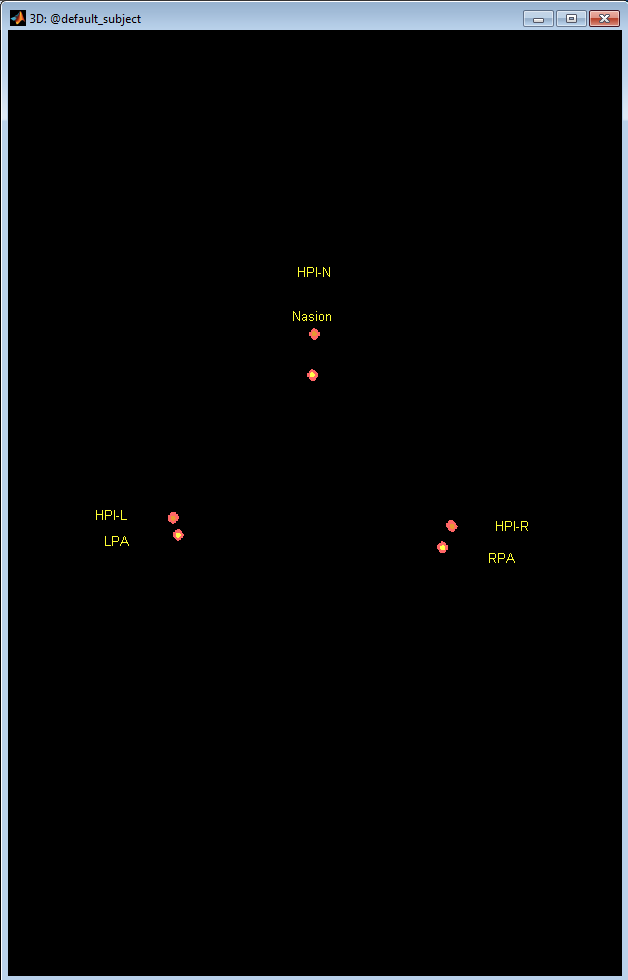
Figure 12. Collect fiducials
Collect the required number of EEG points
- Each point is displayed on the 3D points display
- The counter is incremented each time a point is collected and therefore indicates which point needs to be collected next
- The coordinates are displayed in the coordinate list in cm The points are saved in the brainstorm channel file as both EEG Channel Locations and EEG Head Points
- Once complete, the collection automatically moves to the head shape collection
Collect the desired number of head shape points
- In order to take full advantage of the MEG/MRI co-registration algorithms in Brainstorm, it is recommended to collect approximately 100-150 points from the head: 5-10 points from the boney part of the nose, 10-15 points across the eyebrows, then an even coverage of the scalp from the eyebrows to the inion. It is NOT recommended to collect points below the inion on the neck, since the shape of the neck can change between lying in the MRI scanner and sitting during point collection. Even coverage is more important that dense coverage. See Figure 13 as an example.
- Each point is displayed on the 3D points display
- The counter is incremented each time a point is collected and therefore indicates which point will to be collected next
- The coordinates are displayed in the coordinate list in cm
- The points are saved in the brainstorm channel file as EXTRA Head Points
- Once complete, the operator can save the points to a text file in a variety of formats.
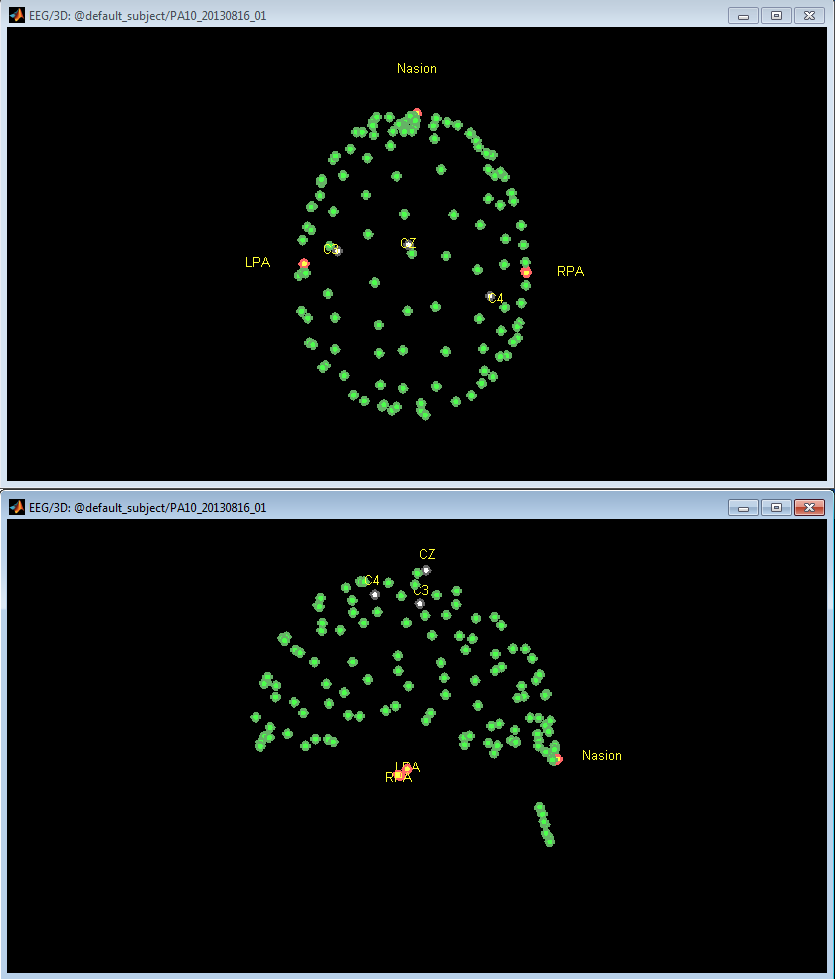
Figure 13. Collect head shape
Save
- The points can be saved as a POS file for import into CTF datasets or your Brainstorm database. Select 'Save as...' and confirm the file name and location.
When exiting the program, select File > Save and Exit to ensure the points are saved in the brainstorm database.
Export points to text file
All points are automatically saved in a brainstorm channel file
- After exiting from the Digitize application, the points can be written to a text file in a variety of formats.
Right click on the channel file just created, select File > Export to file, select the type of file to save.
Figure 14. Export points to text file
How to use this file with CTF systems
The procedure we recommend is the following:
- Export the file in the format "EEG+headshape: Polhemus (*.ds)"
- Copy the .pos file to all the CTF datasets recorder for this subject (.ds folder)
- When importing the CTF datasets in Brainstorm, the file will be automatically loaded.
Troubleshooting
There are several things that can affect the quality and reproducibility of the measurements from the Polhemus Fastrak:
Device communication
Problem appears to be with the communication between the device and Brainstorm:
- Power on the device
- Restart Matlab
- Start brainstorm
- Start Digitize
- At the Matlab command prompt, type this: s = instrfind()
If you do not have Matlab, you can open a command prompt from the Brainstorm File menu BEFORE starting Digitize. The command, s = instrfind() should be typed into the window, click execute and see the output in the terminal window.:
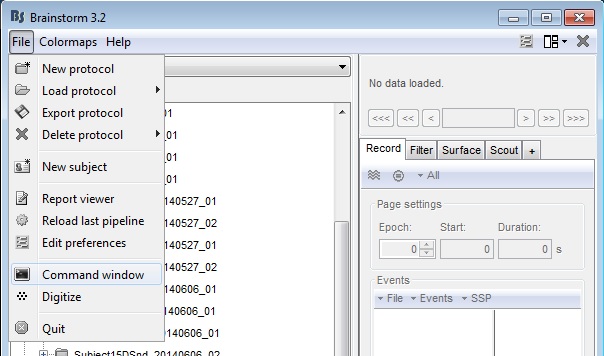
Figure 15: Start command window from Brainstorm
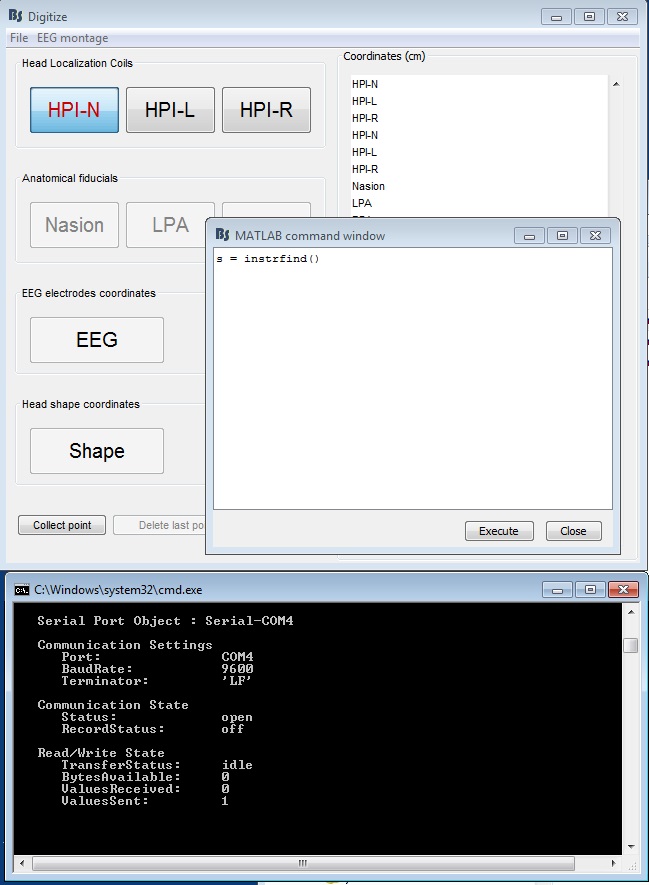
Figure 16: Serial connection settings (Fastrak)
From the Digitize->File->Edit settings (Figure 7), confirm the information in 's' matches your settings
- Confirm the information in 's' matches your COM port settings of your computer (this info is in the Device Manager)
- Press the button of the stylus, then execute s = instrfind() again. You should see that there are bytes available (94 bytes for Fastrak, 120 bytes for Patriot).
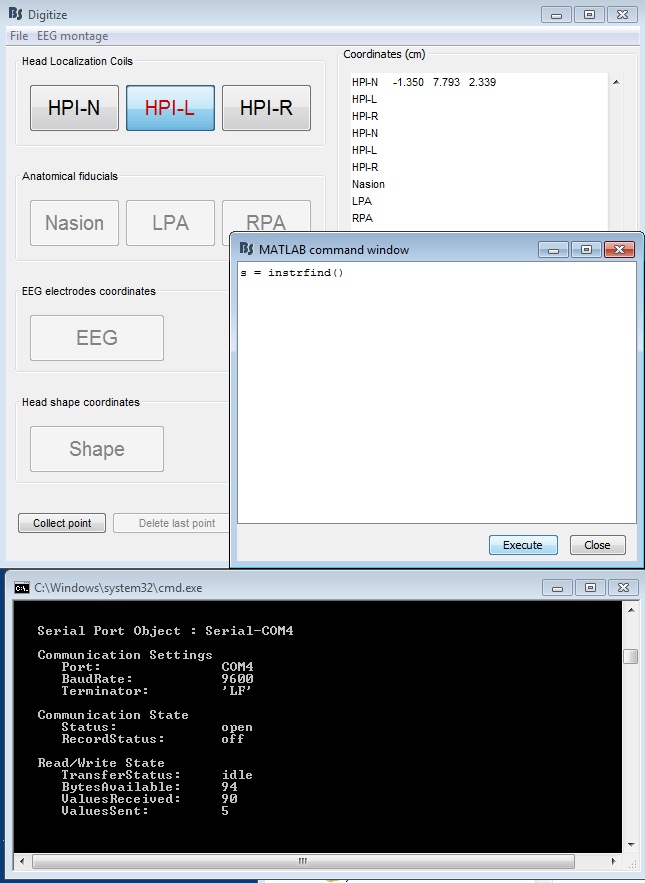
Figure 17: Serial connection after one point collected (Fastrak)
Accuracy
- It is very sensitive to it's environment. It can take many repeated tests, moving the phantom around the room, to find the quietest spot where the errors between measurements are a minimum. You can do this simply with Matlab to collect points so that you skip the Brainstorm step which transforms the anatomical points to headspace.
- In our experience, there is an ideal distance and relationship between the mounted receiver and the transmitter. For instance, when measuring a short person or child, the setup may not work and you may need to use a booster on the chair. Try different heights and distances. For our setup, this is approximately like Figure 2 (~40cm up and ~30cm forward from the transmitter to the mounted receiver).
- The orientation of the stylus should always be perpendicular to the scalp surface. There is some compensation for the orientation of the stylus as part of the brainstorm application, but errors can be introduced if the collection is not consistent.
- Be very careful not to move the mounted receiver during collection (or between collections when comparing the two).
- As for the orientation of the transmitter, this may be trial and error depending on the settings of your device. Care should be taken to ensure the orientation of coordinate system is correct after point collection.
Additional documentation
Forum: Stylus orientation issues
Forum: Emitter position
Forum: Stylus button not working
