|
Size: 20260
Comment:
|
Size: 21175
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| = Import and process Neuromag raw recordings = This tutorial describes how to process continuous Neuromag MEG recordings. It is based on median nerve stimulation acquired at the Massachusetts General Hospital in 2005 with a Neuromag Vectorview 306 system. The sample dataset contains the results for one subject for both arms: 18 minutes, 300 stimulations per arm. |
= Elekta-Neuromag recordings = This tutorial describes how to process continuous Elekta-Neuromag MEG recordings. It is based on median nerve stimulation acquired at the Massachusetts General Hospital in 2005 with a Neuromag Vectorview 306 system. The sample dataset contains the results for one subject for both arms: 18 minutes, 300 stimulations per arm. |
| Line 15: | Line 15: |
| * Menu File > Create new protocol > "'''!TutorialNeuromag'''", with the following options: * "'''No, use individual anatomy'''", * "'''Yes, use one channel file per subject'''".<<BR>> (we have only one run, it's possible to share the channels positions and head models between different conditions, never select this option if you have more than one run) <<BR>><<BR>> {{attachment:createProtocol.gif||height="239",width="222"}} |
* Menu File > Create new protocol > '''!TutorialNeuromag''', with the following options: * '''No, use individual anatomy''', * '''Yes, use one channel file per subject'''.<<BR>> (we have only one run, it's possible to share the channels positions and head models between different conditions, never select this option if you have more than one run) <<BR>><<BR>> {{attachment:createProtocol.gif||height="372",width="347"}} |
| Line 35: | Line 35: |
| ==== Link the FIF file to the database ==== | === Link the FIF file to the database === |
| Line 42: | Line 42: |
| * Then you have to select from which technical track you want to read the events. <<BR>><<BR>> {{attachment:eventsChannels.gif||height="165",width="375"}} | * Then you have to select from which technical tracks you want to read the events. <<BR>><<BR>> {{attachment:eventsChannels.gif||height="165",width="375"}} |
| Line 49: | Line 49: |
| * Always make sure that the final registration looks good. The yellow surface represents the inner surface of the MEG helmet, and the green dots represent the head points that where saved with the MEG recordings. In this case the registration looks acceptable. If you consider it is not your case, you can try the two other menus: "Edit..." and "Refine using head points" <<BR>><<BR>> {{attachment:checkRegistrationFig.gif}} * The files that appear in the folder ''(Common files)'' will be shared for all the conditions of Subject01. This mechanism allows to compute the forward and inverse models only once for all the conditions when there is only one run (= one channel file = one head position) :<<BR>><<BR>> {{attachment:treeAfterImport1.gif||height="116",width="210"}} |
* Always make sure that the final registration looks good. The yellow surface represents the inner surface of the MEG helmet and the green dots represent the head points that were saved with the MEG recordings. In this case the registration looks acceptable. If you consider it is not the case, you can fix it manually: right-click on the channel file > MRI registration > Edit. <<BR>><<BR>> {{attachment:checkRegistrationFig.gif||height="199",width="218"}} * The files that appear in the folder ''(Common files)'' will be shared for all the conditions of Subject01. This mechanism allows to compute the forward and inverse models only once for all the conditions when there is only one run (= one channel file = one head position).<<BR>><<BR>> {{attachment:treeAfterImport1.gif||height="116",width="210"}} |
| Line 52: | Line 52: |
| ==== Review the recordings ==== | === Review the recordings === |
| Line 64: | Line 64: |
| ==== Evaluate the recordings ==== | === Evaluate the recordings === |
| Line 71: | Line 71: |
| * '''MEG GRAD1''': Only the first axial gradiometer on each chip * '''MEG GRAD2''': Only the second axial gradiometer on each chip |
* '''MEG GRAD2''': Only the first axial gradiometer on each chip (name ends with a "2") * '''MEG GRAD3''': Only the second axial gradiometer on each chip (name ends with a "3") |
| Line 76: | Line 76: |
| ==== Remove: 60Hz and harmonics ==== * __Preliminary note__: This process is very time and resource consuming. If you are not planning to analyze anything specifically around 60Hz, you may want to skip this section and and three hours of computation associated with it it. It is illustrated here as a reference documentation for users who would need it. * The standard procedure illustrated for removing the 60Hz power line noise illustrated in the CTF-based introduction tutorial would be to just run the process "Pre-process > Sinusoid removal" on the continuous file. However, this case is a bit more complicated. * If you try to run this process without selecting the option "Process the entire file at once", you would get the following error. The file already contains some SSP projectors, which require to read the entire file in one block to apply frequency filters.<<BR>><<BR>> {{attachment:sinError.gif||height="259",width="662"}} * But if you try with the option "Process the entire file at once", you would probably crash Matlab or freeze your computer because of a lack of memory. The file is already large and this process requires to create a lot of matrices of the same size in memory. Do not try this unless you have at least 24Gb of memory available on your computer. |
=== Remove: 60Hz and harmonics === * __Preliminary note__: This process is very time and resource consuming. If you are not planning to analyze anything specifically around 60Hz, you may want to skip this section and and the long computation time associated with it. It is described here as a reference for users who would need it. * The standard procedure illustrated for removing the 60Hz power line noise illustrated in the CTF-based introduction tutorial would be simply to run the process "Pre-process > Notch filter" on the continuous file. However, this case is a bit more complicated. * If you try to run this process without selecting the option "Process the entire file at once", you would get the following error. The file already contains some SSP projectors, created by one of the Neuromag progams. Using those projectors requires all the sensors, and running frequency filters requires all the time samples at once. As a consequence, the entire file must be read in one block in order to apply frequency filters on it. <<BR>><<BR>> {{attachment:sinError.gif||height="234",width="658"}} * Alternatively, if you try with the option "Process the entire file at once", you would probably crash Matlab or freeze your computer because of a lack of memory. The file is already large and this process requires to create a lot of matrices of the same size in memory. Do not try this unless you have at least 24Gb of memory available on your computer. |
| Line 84: | Line 84: |
| * Run the process "'''Pre-process > Sinusoid removal'''", configured as follows:<<BR>>(this may take a few hours to complete)<<BR>><<BR>> {{attachment:processSin.gif||height="338",width="350"}} * To evaluate the results of this process, select the last file generated ('''"Raw | clean | sin"''') and run again the process "'''Frequency > Power spectrum density'''".<<BR>><<BR>> {{attachment:processPsd2.gif||height="351",width="396"}} |
* Run the process "'''Pre-process > Notch filter'''", configured as follows:<<BR>>(this may take a while complete...)<<BR>><<BR>> {{attachment:processSin.gif||height="274",width="336"}} * To evaluate the results of this process, select the last file generated ('''"Raw | clean | notch"''') and run again the process "'''Frequency > Power spectrum density'''".<<BR>><<BR>> {{attachment:processPsd2.gif||height="351",width="396"}} |
| Line 88: | Line 88: |
| ==== Heartbeats and eye blinks ==== Signal Space Projection (SSP) is a method in for projecting away stereotyped artifacts (such as eye blinks and heartbeats) out of the recordings. |
=== Heartbeats and eye blinks === Signal Space Projection (SSP) is a method for projecting the recordings away from stereotyped artifacts, such as eye blinks and heartbeats. |
| Line 91: | Line 91: |
| * Double-click on the last continuous file you got to display all the '''MEG '''recordings.<<BR>>("Raw | clean | sin" with the sinusoid removal, or "Link to raw file" without) | * Double-click on the last continuous file you got to display all the '''MEG '''recordings.<<BR>>("Raw | clean | notch" with the notch filter, or "Link to raw file" without) * Right-click on the link > '''ECG '''> Display time series, to look at the heartbeats. |
| Line 93: | Line 94: |
| * Right-click on the link > '''ECG '''> Display time series, to look at the heartbeats. | |
| Line 95: | Line 95: |
| * '''Detect eye blinks:''' Select channel '''EOG 062''', event name "blink".<<BR>><<BR>> {{attachment:processDetectEog.gif||height="224",width="329"}} * '''Detect heartbeats:''' Select channel '''ECG 063''', event name "cardiac".<<BR>><<BR>> {{attachment:processDetectEcg.gif||height="221",width="327"}} |
* '''Detect heartbeats:''' Select channel '''ECG 063''', event name "cardiac".<<BR>><<BR>> {{attachment:processDetectEcg.gif||height="214",width="310"}} * '''Detect eye blinks:''' Select channel '''EOG 062''', event name "blink".<<BR>><<BR>> {{attachment:processDetectEog.gif||height="213",width="309"}} * '''Remove simultaneous:''' To avoid capturing ocular artifacts in the cardiac SSP.<<BR>>Remove events named: "'''cardiac'''"<<BR>>When too close from events: "'''blink'''"<<BR>>Minimum delay between events: '''250ms'''.<<BR>><<BR>> {{attachment:processRemoveSimult.gif||height="234",width="309"}} |
| Line 98: | Line 99: |
| * From the same menu, run the following processes, separately from GRAD and MAG: * '''Compute SSP: Eyeblinks''': Event "blink", sensor types="'''MEG MAG'''".<<BR>>(select components #1 and #2) <<BR>><<BR>> {{attachment:sspEogMag.gif||height="200",width="754"}} * '''Compute SSP: Eyeblinks''': Event "blink", sensor types="'''MEG GRAD'''".<<BR>>(select component #1)<<BR>><<BR>> {{attachment:sspEogGrad.gif||height="217",width="600"}} * '''Compute SSP: Heartbeats''': Event "cardiac", sensor types="'''MEG MAG'''".<<BR>>(select component #1) <<BR>><<BR>> {{attachment:sspEcgMag.gif||height="217",width="600"}} * '''Compute SSP: Heartbeats''': Event "cardiac", sensor types="'''MEG GRAD'''".<<BR>>(select component #2) <<BR>><<BR>> {{attachment:sspEcgGrad.gif||height="217",width="600"}} |
* From the same menu, run the following processes, separately for GRAD and MAG: * '''Compute SSP: Heartbeats''': Event "cardiac", sensor types="'''MEG MAG'''", use existing SSP.<<BR>>(select component #1) <<BR>><<BR>> {{attachment:sspEcgMag.gif||height="218",width="560"}} * '''Compute SSP: Heartbeats''': Event "cardiac", sensor types="'''MEG GRAD'''", use existing SSP.<<BR>>(select component #1) <<BR>><<BR>> {{attachment:sspEcgGrad.gif||height="219",width="561"}} * '''Compute SSP: Eyeblinks''': Event "blink", sensor types="'''MEG MAG'''", use existing SSP.<<BR>>(select component #1) <<BR>><<BR>> {{attachment:sspEogMag.gif||height="218",width="560"}} * '''Compute SSP: Eyeblinks''': Event "blink", sensor types="'''MEG GRAD'''", use existing SSP.<<BR>>(select component #1)<<BR>><<BR>> {{attachment:sspEogGrad.gif||height="218",width="560"}} |
| Line 104: | Line 105: |
| * At this stage, you should also review the entire file (pressing repeatedly F3) to detect any obvious bad segments in the recordings. If you detect any, select the time window to mark as bad, right click on the figure > Reject time segment. | * At this stage, you should also review the entire file (pressing repeatedly F3) to detect any obvious bad segment in the recordings. If you detect any, select the time window to mark as bad, right click on the figure > Reject time segment. |
| Line 108: | Line 109: |
| * Right-click on the raw link > Import in database<<BR>><<BR>> {{attachment:importPopup.gif||height="298",width="193"}} | * Right-click on the raw link > Import in database.<<BR>><<BR>> {{attachment:importPopup.gif||height="298",width="193"}} |
| Line 119: | Line 120: |
| * Rename the condition "'''STI_001__5__'''__" and "'''STI_002'''__'''5'''" into respectively "'''Right'''" and "'''Left'''" (Right-click > Rename, or press F2, or click twice waiting a while before the two clicks). * Rename the groups of trials as well to '''Left''' and '''Right'''.<<BR>><<BR>> {{attachment:rename.gif||height="105",width="225"}} |
* Rename the condition "'''STI_001_5'''" and "'''STI_002''''''_5'''" into respectively "'''Right'''" and "'''Left'''" (Right-click > Rename, or press F2, or click twice waiting a while before the two clicks). * Rename the groups of trials to '''Left''' and '''Right'''.<<BR>><<BR>> {{attachment:rename.gif||height="105",width="225"}} |
| Line 128: | Line 129: |
| ==== Stimulation artifact ==== * We see clearly our evoked responses in those figures, but the stimulation artifact between 0ms and 4ms causes the automatic scaling of the figures to be to high, especially for the right stimulation. This effect will get a lot worse when displaying the sources for those files. To get a better display of these averages, we can cut these peaks at 0ms by replacing the values with linear interpolations. |
=== Stimulation artifact === * We see clearly our evoked responses in those figures, but the stimulation artifact between 0ms and 4ms causes the automatic scaling of the figures to be too high, especially for the right stimulation. This effect would get a lot worse when displaying the sources for those files. To get a better display of these averages, we can cut these peaks at 0ms by replacing the values with linear interpolations. |
| Line 131: | Line 132: |
| * Run the process '''Artifacts > Cut stimulation artifact''': 0-4ms, overwrite<<BR>> {{attachment:processCut.gif||height="239",width="471"}} | * Run the process '''Artifacts > Cut stimulation artifact''': 0-4ms, overwrite<<BR>><<BR>> {{attachment:processCut.gif||height="239",width="471"}} |
| Line 137: | Line 138: |
| ==== Gradiometers & magnetometers ==== | == Gradiometers & magnetometers == |
| Line 139: | Line 140: |
| * The topographies look very similar, because all the displays are re-interpolated on a surface of virtual magnetometers. To display the real values for the different types of sensors, without this magnetic interpolation, use the menus "No magnetic interpolation". * Right-click on the Left average > MEG (magnetometers) > No magnetic interpolation > 2D Sensor cap. <<BR>><<BR>> {{attachment:averageMagNointerp.gif||height="196",width="235"}} * Repeat the same operation for "MEG (gradiometers)". Now it's asking for an additional question: how to group the two gradiometers? <<BR>><<BR>> {{attachment:averageGradNointerp.gif||height="161",width="473"}} * You will not be able to display at the same time the values for the first gradiometer and the second gradiometer. If you select again the same menu, the question is considered to be already answered, it re-using the same response as before. You have to close everything (Close all button in the top-right corner of the Brainstorm window) before being able to make a new choice. * The three options side-by-side at 22ms (first, second, norm): <<BR>><<BR>> {{attachment:averageGradNointerp2.gif||height="165",width="542"}} |
* The topographies look very similar, because all the values are re-interpolated on a surface of virtual magnetometers. To display the real values for the different types of sensors, without this magnetic interpolation, use the menus "No magnetic interpolation". * Right-click on Left > MEG (mag) > No magnetic interpolation > 2D Sensor cap. <<BR>><<BR>> {{attachment:averageMagNointerp.gif||height="196",width="235"}} * Repeat the same operation for "MEG (gradiometers)". You now have three options, because it is not possible to display at the same time the values for the two planar gradiometers: * Grad 2: First gradiometer, sensor names ending with "2" * Grad 3: Second gradiometer, sensor names ending with "3" * Grad norm: Norm of the two gradiometers => sqrt(grad2^2 + grad3^2) * The three options side-by-side at 22ms (grad 2, grad3, norm): <<BR>><<BR>> {{attachment:averageGradNointerp2.gif||height="165",width="542"}} |
| Line 149: | Line 152: |
| * Right-click on the channel file > Compute head model.<<BR>><<BR>> {{attachment:heademodelMenu.gif}} | * Right-click on the channel file > Compute head model.<<BR>><<BR>> {{attachment:headmodelMenu.gif||height="196",width="228"}} |
| Line 154: | Line 157: |
| * Select all the Left and Right trials simultaneously. Right-click > Noise covariance > Compute from recordings.<<BR>><<BR>> {{attachment:noisecovPopup.gif}} * Leave all the default options and click [OK]. <<BR>><<BR>> {{attachment:noisecovOptions.gif}} |
* Select all the Left and Right trials, right-click > Noise covariance > Compute from recordings.<<BR>><<BR>> {{attachment:noisecovPopup.gif||height="188",width="420"}} * Leave all the default options and click [OK]. <<BR>><<BR>> {{attachment:noisecovOptions.gif||height="374",width="382"}} |
| Line 159: | Line 162: |
| * Right-click on the headmodel > Compute sources.<<BR>><<BR>> {{attachment:inversePopup.gif}} * The inverse kernel is saved in a new file in the database.<<BR>><<BR>> {{attachment:inversePopup.gif}} * Select wMNE and all the default options. This will run a source reconstruction grouping the information from the magnetometers and the gradiometers into the same model. This is the recommended procedure, but you can also estimate sources for the two types of sensors separately if needed. <<BR>><<BR>> {{attachment:inverseOptions.gif}} * A shared inversion kernel was created in the ''(Common files)'' folder, and a link node is now visible for each recording file, single trials and averages. <<BR>><<BR>> {{attachment:inverseFiles.gif}} |
* Right-click on the headmodel > Compute sources.<<BR>><<BR>> {{attachment:inversePopup.gif||height="131",width="224"}} * Select wMNE and all the default options. This will run a source reconstruction grouping the information from the magnetometers and the gradiometers into the same model. This is the recommended procedure, but you can also estimate sources for the two types of sensors separately if needed. <<BR>><<BR>> {{attachment:inverseOptions.gif||height="233",width="207"}} * A shared inversion kernel is created in the ''(Common files)'' folder, and a link node is now visible for each recording file, single trials and averages. <<BR>><<BR>> {{attachment:inverseFiles.gif||height="239",width="229"}} |
| Line 165: | Line 167: |
| ==== Scouts ==== You can easily reproduce exactly the same results as the ones presented in the introduction tutorials. Place scouts to capture the activity in the primary and secondary somatosensory areas to track the processing of the electric stimulations in time, at the surface surface of the brain. {{attachment:final.gif||height="419",width="842"}} == Scripting == The operations described in this tutorial can be reproduced from a Matlab script, available in the Brainstorm distribution: '''brainstorm3/toolbox/script/tutorial_neuromag.m ''' |
Elekta-Neuromag recordings
This tutorial describes how to process continuous Elekta-Neuromag MEG recordings. It is based on median nerve stimulation acquired at the Massachusetts General Hospital in 2005 with a Neuromag Vectorview 306 system. The sample dataset contains the results for one subject for both arms: 18 minutes, 300 stimulations per arm.
This document shows what to do step by step, but does not really explain what is happening, the meaning of the different options or processes, the issues or bugs you can encounter, and does not provide an exhaustive description of the software features. Those topics are introduced in the basic tutorials based on CTF recordings; so make sure that you followed all those initial tutorials before going through this one.
The script file tutorial_neuromag.m in the brainstorm3/toolbox/script folder performs exactly the same tasks automatically, without any user interaction. Please have a look at this file if you plan to write scripts to process recordings in .fif format.
Contents
Download and installation
Requirements: You have already followed all the basic tutorials and you have a working copy of Brainstorm installed on your computer.
Go to the Download page of this website and download the file sample_neuromag.zip
- Unzip it in a folder that is not in any of the Brainstorm folders (program folder or database folder). This is really important that you always keep your original data files in a separate folder: the program folder can be deleted when updating the software, and the contents of the database folder is supposed to be manipulated only by the program itself.
- Start Brainstorm (Matlab scripts or stand-alone version)
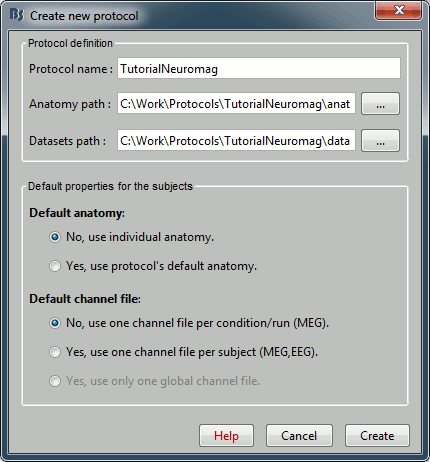
Menu File > Create new protocol > TutorialNeuromag, with the following options:
No, use individual anatomy,
Yes, use one channel file per subject.
(we have only one run, it's possible to share the channels positions and head models between different conditions, never select this option if you have more than one run)
Import the anatomy
Right-click on the TutorialNeuromag folder > New subject > Subject01 (leave all the defaults).
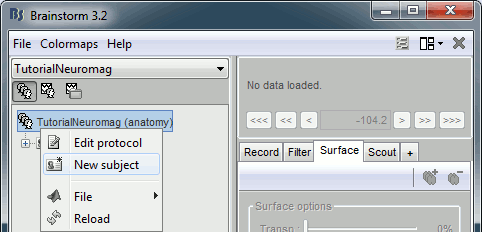
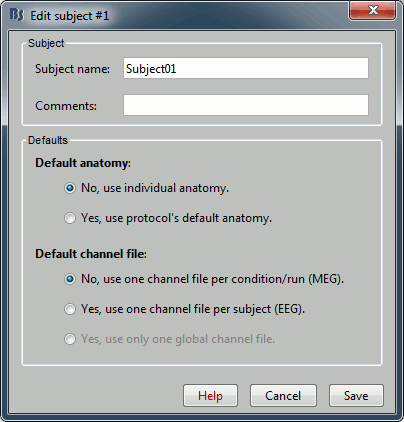
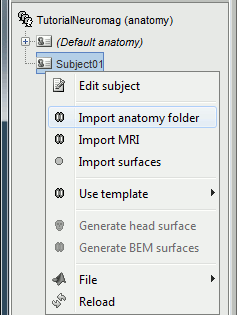
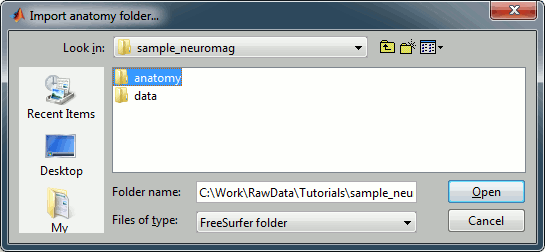
Right-click on the subject node > Import anatomy folder:
Set the file format: "FreeSurfer folder"
Select the folder: sample_neuromag/anatomy
Number of vertices of the cortex surface: 15000 (default value)
- Set the 6 required fiducial points (indicated in MRI coordinates):
- NAS: x=131, y=232, z=123
- LPA: x=48, y=136, z=74
- RPA: x=204, y=131, z=67
- AC: x=127, y=143, z=118
- PC: x=127, y=117, z=111
- IH: x=127, y=120, z=167 (anywhere on the midsagittal plane)
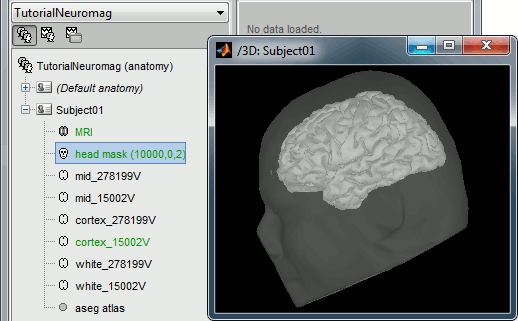
At the end of the process, make sure that the file "cortex_15000V" is selected (downsampled pial surface, that will be used for the source estimation). If it is not, double-click on it to select it as the default cortex surface.
Access the recordings
Link the FIF file to the database
- The sample_neuromag package contains one run for one subject only.
- Switch to the "functional data" view, the middle button in the toolbar above the database explorer.
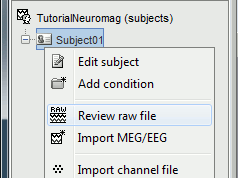
Right click on the Subject01 > Review raw file:
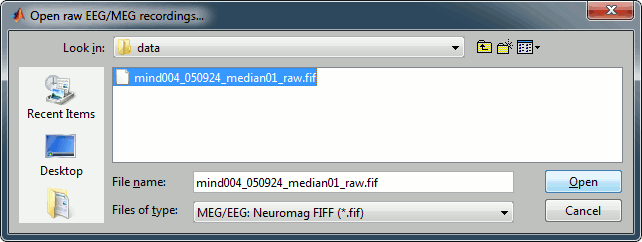
Select the file format: MEG/EEG Neuromag FIFF (*.fif)
Pick the file: sample_neuromag/data/mind004_050924_median01_raw.fif
Select "Event channel" when asked about the events.
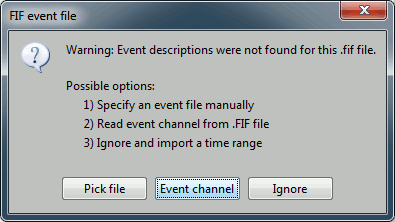
Then you have to select from which technical tracks you want to read the events.
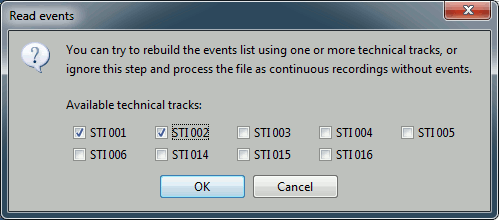
- In this dataset, the tracks of interest are:
STI 001: Contains the onsets of the electric stimulation of the right arm.
STI 002: Contains the onsets of the electric stimulation of the left arm.
- Those are not standard settings, they depend on your acquisition setup. In most cases, you would rather import "events" defined with the Neuromag acquisition software, which are saved in tracks STI 014, STI 101, STI 201...
Select STI 001 and STI 002. The full tracks are read, all the changes of values are detected from those two tracks and a list of events is created.
At the end, you're asked if you want to "Refine the registration with the head points". This operation improves the initial MRI/MEG registration by fitting the head points digitized at the MEG acquisition and the scalp surface. Answer YES.
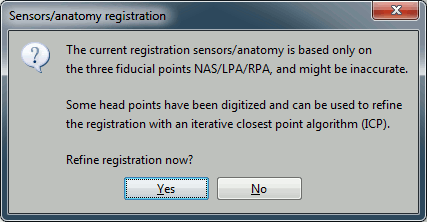
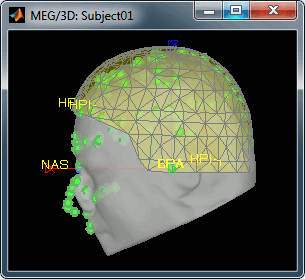
Always make sure that the final registration looks good. The yellow surface represents the inner surface of the MEG helmet and the green dots represent the head points that were saved with the MEG recordings. In this case the registration looks acceptable. If you consider it is not the case, you can fix it manually: right-click on the channel file > MRI registration > Edit.
The files that appear in the folder (Common files) will be shared for all the conditions of Subject01. This mechanism allows to compute the forward and inverse models only once for all the conditions when there is only one run (= one channel file = one head position).
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
Review the recordings
Right-click on the "Link to raw file" to see what types of channels are available:
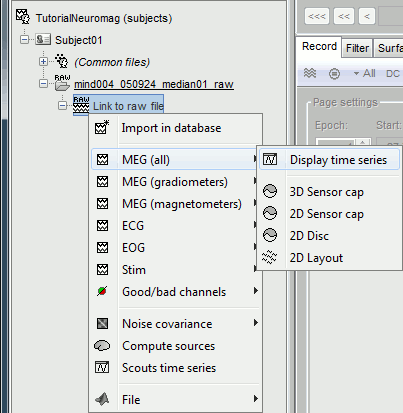
MEG (all): Represents all the MEG sensors in the file (magnetometers and gradiometers)
MEG (magnetometers): 101 magnetometers (sensor names ending with a "1")
MEG (gradiometers): 202 planar gradiometers (sensor names ending with "2" or "3")
ECG: Electro-cardiogram
EOG: Electro-oculogram
Stim: All the stimulation lines
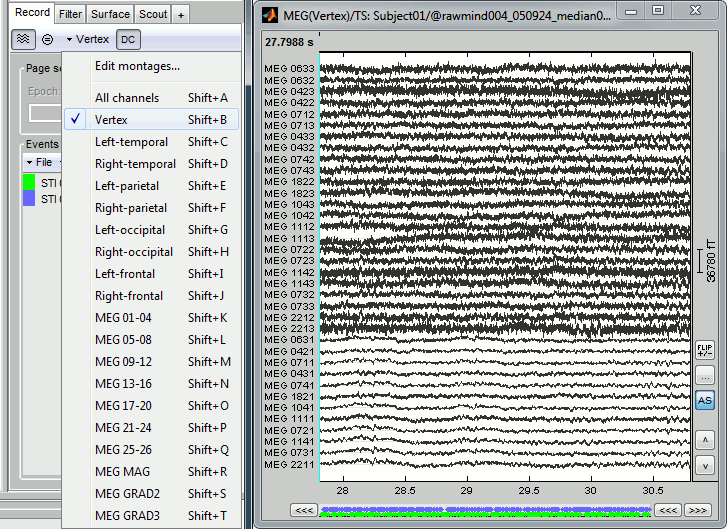
- Double-click on the file to display all the MEG sensors.
In the Record tab, look at the list of the available default display configurations for the Neuromag Vectorview MEG systems. If you would like to use different combinations of sensors, you can create your own combinations with the entry "Edit montages..."
Pre-processing
Evaluate the recordings
- Drag and drop the "Link to raw file" into the Process1 list.
Select the process "Frequency > Power spectrum density", configure it as follows:
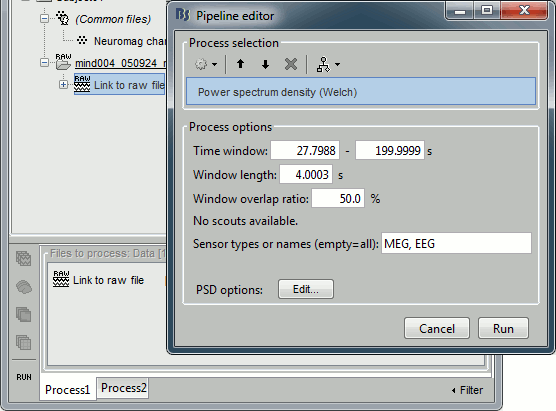
- In all the processes, the types of sensors can be specified using the following convention:
MEG: All the MEG sensors
MEG MAG: All the magnetometers
MEG GRAD: All the gradiometers
MEG GRAD2: Only the first axial gradiometer on each chip (name ends with a "2")
MEG GRAD3: Only the second axial gradiometer on each chip (name ends with a "3")
Double-click on the PSD file to display it.
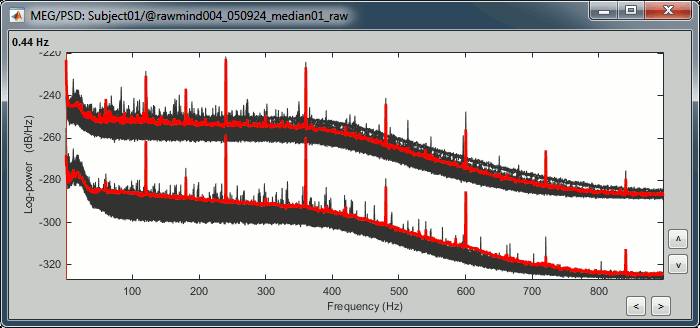
- This power spectrum shows that there is no MEG sensor that is clearly bad, but some sensors show a strong contamination from the power lines (60Hz and harmonics).
Remove: 60Hz and harmonics
Preliminary note: This process is very time and resource consuming. If you are not planning to analyze anything specifically around 60Hz, you may want to skip this section and and the long computation time associated with it. It is described here as a reference for users who would need it.
The standard procedure illustrated for removing the 60Hz power line noise illustrated in the CTF-based introduction tutorial would be simply to run the process "Pre-process > Notch filter" on the continuous file. However, this case is a bit more complicated.
If you try to run this process without selecting the option "Process the entire file at once", you would get the following error. The file already contains some SSP projectors, created by one of the Neuromag progams. Using those projectors requires all the sensors, and running frequency filters requires all the time samples at once. As a consequence, the entire file must be read in one block in order to apply frequency filters on it.
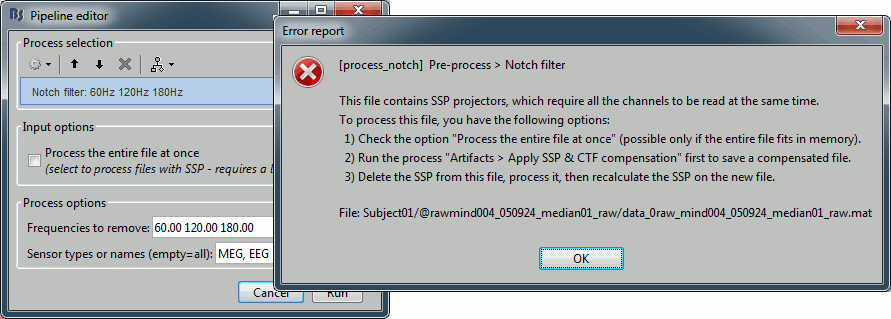
- Alternatively, if you try with the option "Process the entire file at once", you would probably crash Matlab or freeze your computer because of a lack of memory. The file is already large and this process requires to create a lot of matrices of the same size in memory. Do not try this unless you have at least 24Gb of memory available on your computer.
- The solution is to re-write an intermediate file with the SSP projectors applied, that you will be able to process easily after.
Select the "Link to raw file" in Process1 and run process "Artifacts > Apply SSP and CTF compensation". This creates a new .fif file which is an exact copy of the original .fif file, but with the SSP projectors applied to the recordings values (instead of being just saved as linear operators on the side).
Clear the Process1 list and select the new continuous file "Raw | clean"
Run the process "Pre-process > Notch filter", configured as follows:
(this may take a while complete...)
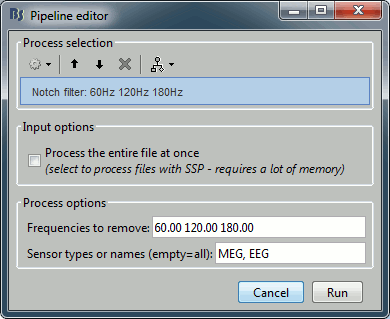
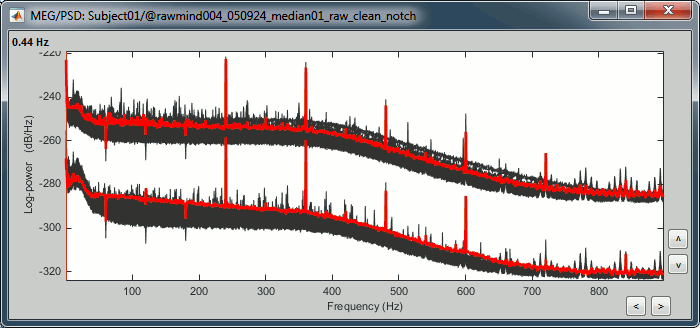
To evaluate the results of this process, select the last file generated ("Raw | clean | notch") and run again the process "Frequency > Power spectrum density".
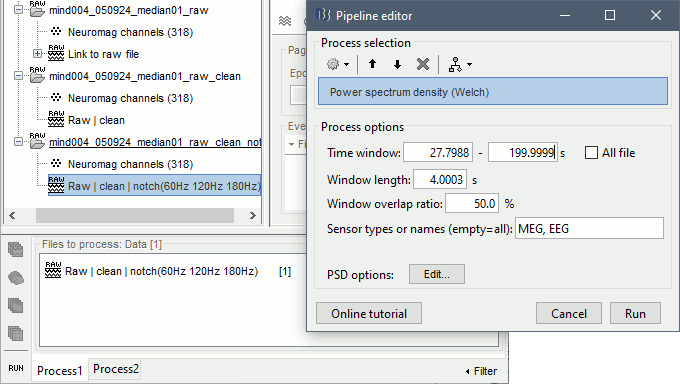
You should observe a significant decrease of the contributions of the removed frequencies (60Hz, 120Hz, 180Hz) compared with the original spectrum.
Heartbeats and eye blinks
Signal Space Projection (SSP) is a method for projecting the recordings away from stereotyped artifacts, such as eye blinks and heartbeats.
Double-click on the last continuous file you got to display all the MEG recordings.
("Raw | clean | notch" with the notch filter, or "Link to raw file" without)Right-click on the link > ECG > Display time series, to look at the heartbeats.
Right-click on the link > EOG > Display time series, to look at the eye movements.
- From the SSP menu in the Record tab, run the automatic detections:
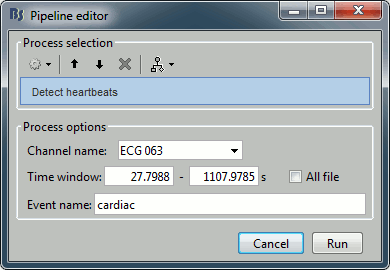
Detect heartbeats: Select channel ECG 063, event name "cardiac".
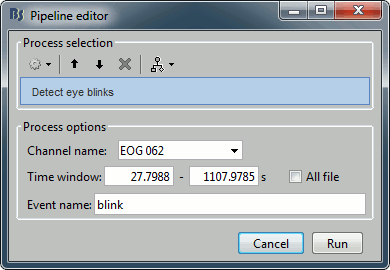
Detect eye blinks: Select channel EOG 062, event name "blink".
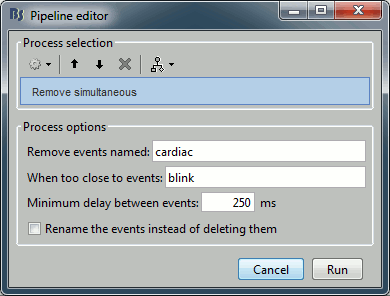
Remove simultaneous: To avoid capturing ocular artifacts in the cardiac SSP.
Remove events named: "cardiac"
When too close from events: "blink"
Minimum delay between events: 250ms.
Review the traces of ECG/EOG channels and make sure the events detected make sense.
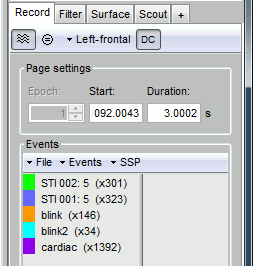
- From the same menu, run the following processes, separately for GRAD and MAG:
Compute SSP: Heartbeats: Event "cardiac", sensor types="MEG MAG", use existing SSP.
(select component #1)
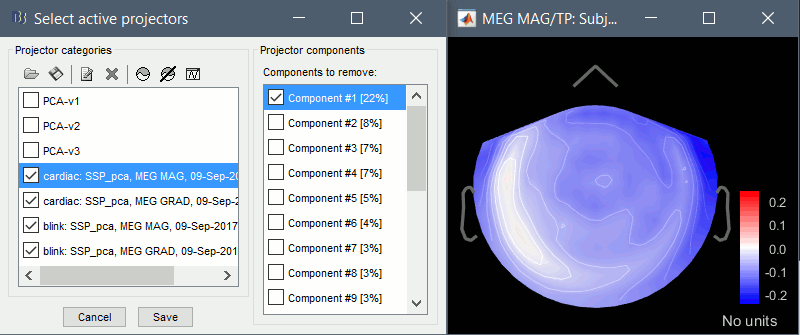
Compute SSP: Heartbeats: Event "cardiac", sensor types="MEG GRAD", use existing SSP.
(select component #1)
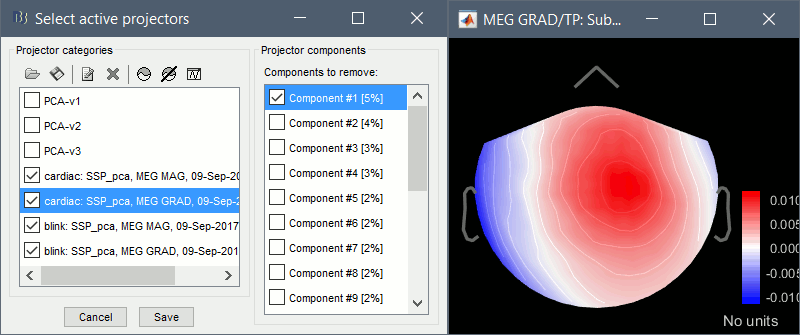
Compute SSP: Eyeblinks: Event "blink", sensor types="MEG MAG", use existing SSP.
(select component #1)
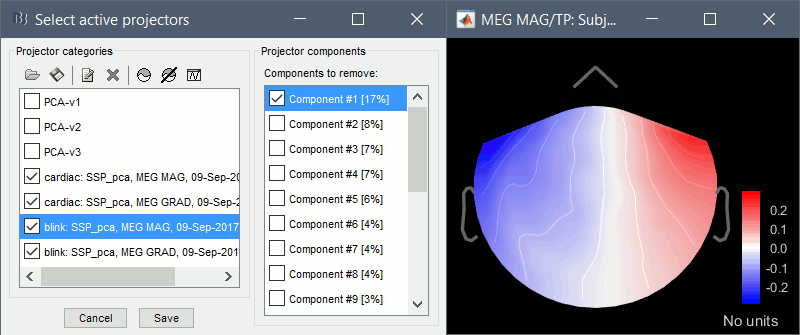
Compute SSP: Eyeblinks: Event "blink", sensor types="MEG GRAD", use existing SSP.
(select component #1)
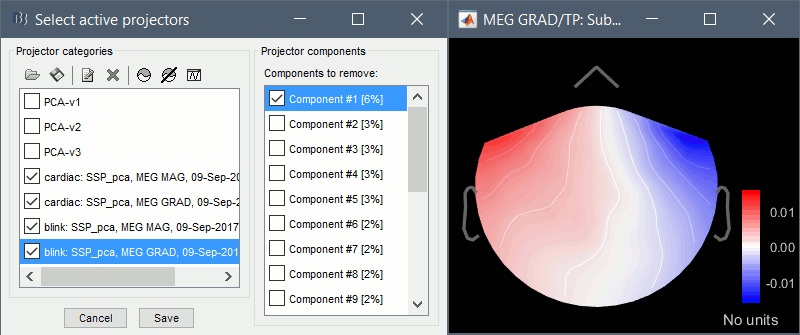
For more information regarding this SSP method, refer to this tutorial:
?Detect and remove artifacts.At this stage, you should also review the entire file (pressing repeatedly F3) to detect any obvious bad segment in the recordings. If you detect any, select the time window to mark as bad, right click on the figure > Reject time segment.
Epoching and averaging
Import the recordings
Right-click on the raw link > Import in database.
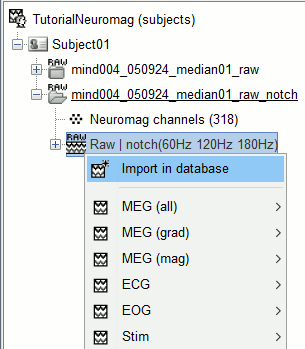
The following figure appears, that asks how to import those recordings in the Brainstorm database.
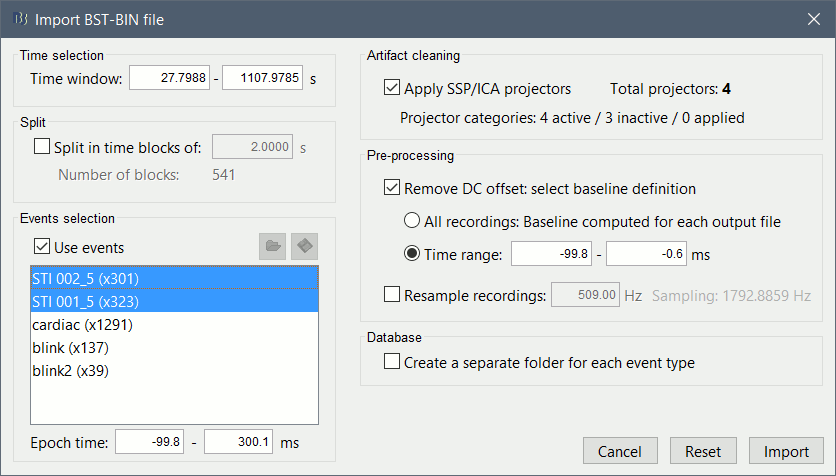
Time window: Time range of interest, keep all the time definition.
Split: Useful to import continuous recordings without events. We do not need this here.
Events selection: Check the "Use events" option, and select both STI 001 and STI 002. The values in the parenthesis represent the number of occurrences of each event.
Epoch time: Time instants that will be extracted before an after each event, to create the epochs that will be saved in the database. Set it to [-100, +300] ms
Use Signal Space Projections: Save and use the SSP vectors created by MaxFilter, MNE and Brainstorm. Keep this option selected.
Remove DC Offset: Check this option, and select: Time range: [-100, 0] ms.
For each epoch, this will: compute the average of each channel over the baseline (pre-stimulus interval: -100ms to 0ms), then subtract it from the channel at all the times in [-100,+300]ms.Resample recordings: Keep this unchecked
Create new conditions for epochs: If selected, a new condition is created for each event type (here, it will create two conditions "STI 001" and "STI 002"). If not selected, all the epochs from all the selected events are saved in the same folder.
- Click on [Import] and wait.
Rename the condition "STI_001_5" and "STI_002_5" into respectively "Right" and "Left" (Right-click > Rename, or press F2, or click twice waiting a while before the two clicks).
Rename the groups of trials to Left and Right.

At this stage, you should review the trials one after the other (press F3 to jump to the next file), separately for the magnetometers and the gradiometers, to make sure that you don't have any bad trial that have been imported. If you find a bad trial: right-click on the file or on the figure > Reject trial.
Averaging
- Drag and drop all the left and right trials to the Process1 tab.
Run the process Average > Average files > By condition:
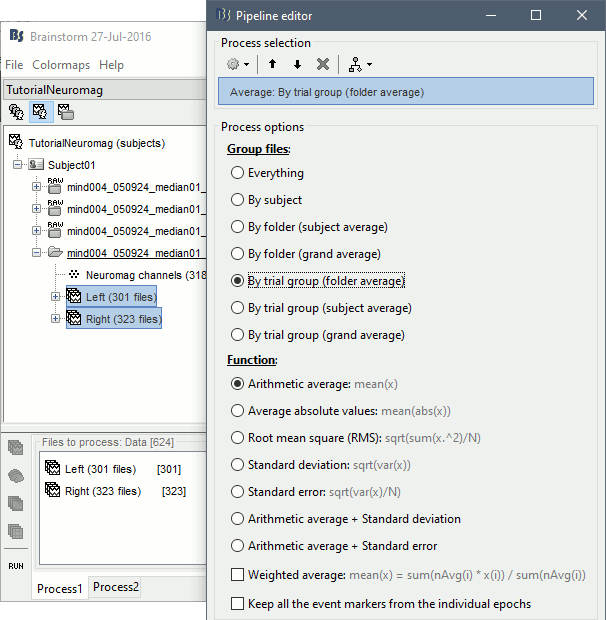
Double-click on the Left and Right averages to display all the MEG sensors:
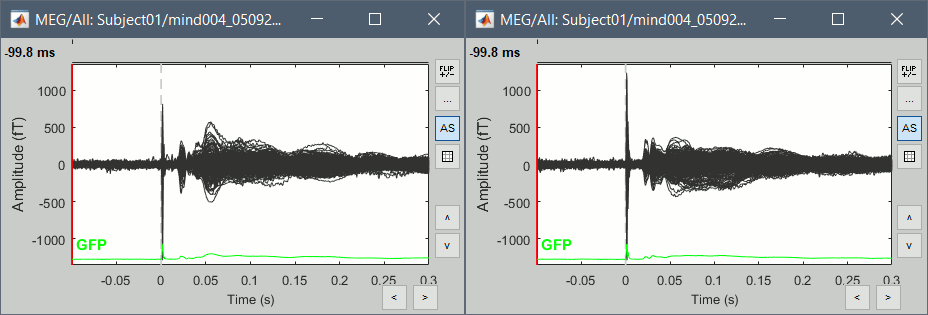
Stimulation artifact
- We see clearly our evoked responses in those figures, but the stimulation artifact between 0ms and 4ms causes the automatic scaling of the figures to be too high, especially for the right stimulation. This effect would get a lot worse when displaying the sources for those files. To get a better display of these averages, we can cut these peaks at 0ms by replacing the values with linear interpolations.
- Select the averages for the Left and Right condition in the Process1 tab.
Run the process Artifacts > Cut stimulation artifact: 0-4ms, overwrite
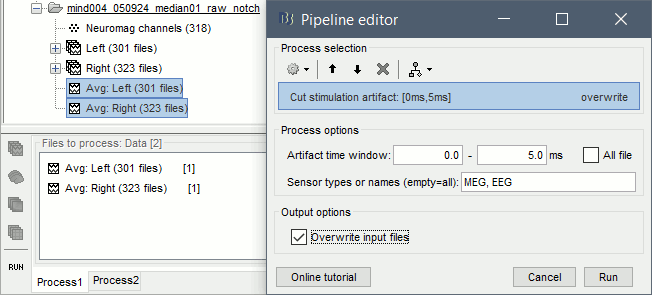
Now you can display the Left and Right averages correctly: Right-click on the file > MEG (All) > Display time series.
Display the 2D topographies for each average: Right-click on the file > MEG (All) > 2D Sensor cap (or press CTRL+T from the time series figures).
You can also add a low-pass filter at 100Hz for the visualization.
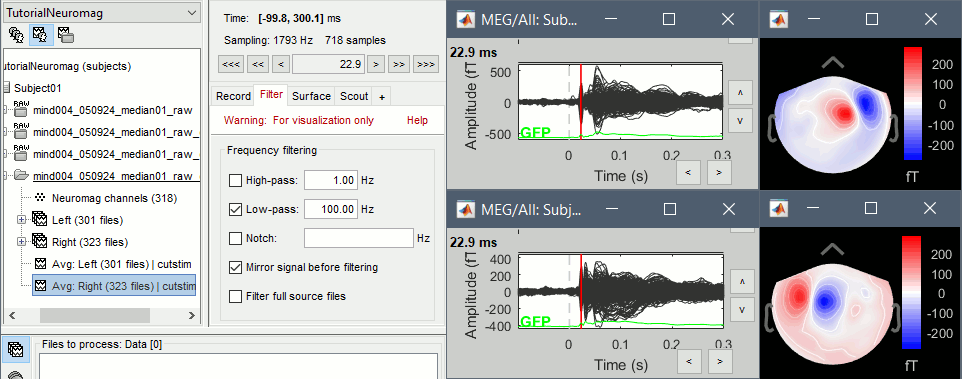
- Everything looks ok, the time series figures show a clean signal with a very high SNR, and the topographies at the first response peak (about 22ms) show an activity on the left side for the Right condition, and on the right side for the Left condition.
Gradiometers & magnetometers
You can also display the MEG MAG (top) and MEG GRAD (bottom) sensors separately:
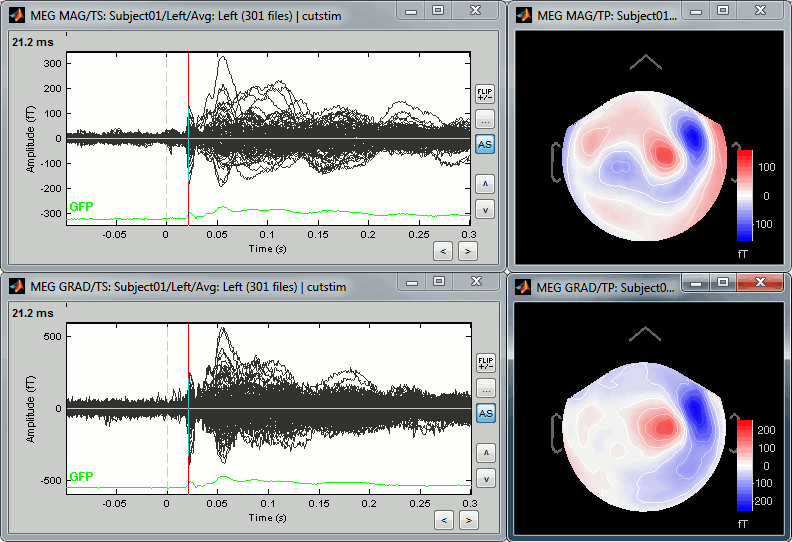
- The topographies look very similar, because all the values are re-interpolated on a surface of virtual magnetometers. To display the real values for the different types of sensors, without this magnetic interpolation, use the menus "No magnetic interpolation".
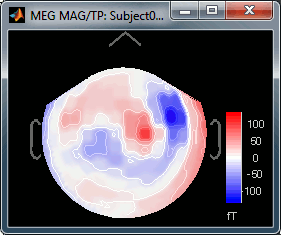
Right-click on Left > MEG (mag) > No magnetic interpolation > 2D Sensor cap.
- Repeat the same operation for "MEG (gradiometers)". You now have three options, because it is not possible to display at the same time the values for the two planar gradiometers:
- Grad 2: First gradiometer, sensor names ending with "2"
- Grad 3: Second gradiometer, sensor names ending with "3"
Grad norm: Norm of the two gradiometers => sqrt(grad22 + grad32)
The three options side-by-side at 22ms (grad 2, grad3, norm):
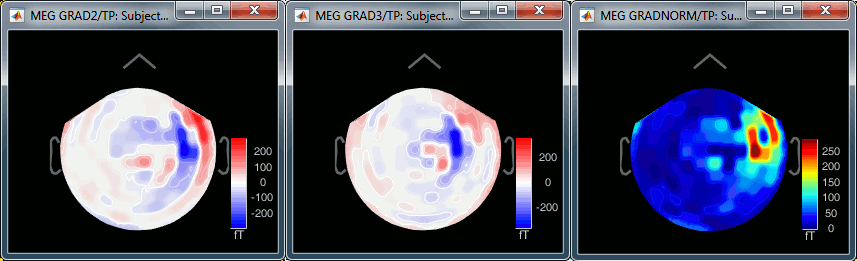
Source estimation
We need now to calculate a source model for these recordings, using a noise covariance matrix calculated from the pre-stimulation baselines. This process is not detailed a lot here because it is very similar to what is shown in the CTF-based introduction tutorials.
Head model
Right-click on the channel file > Compute head model.
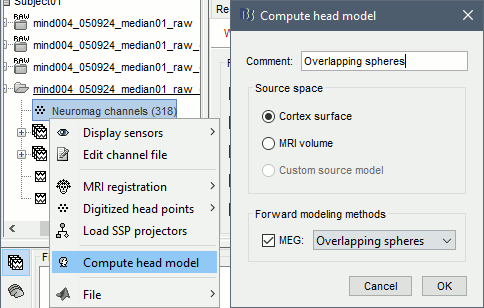
Use the overlapping spheres model and keep all of the options at their default values.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
For more information: ?Head model tutorial.
Noise covariance
Select all the Left and Right trials, right-click > Noise covariance > Compute from recordings.
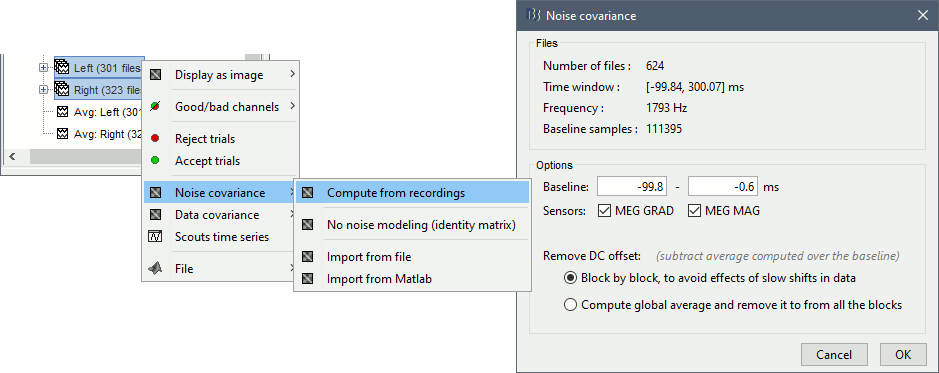
For more information: ?Noise covariance tutorial.
Inverse model
Right-click on the headmodel > Compute sources.
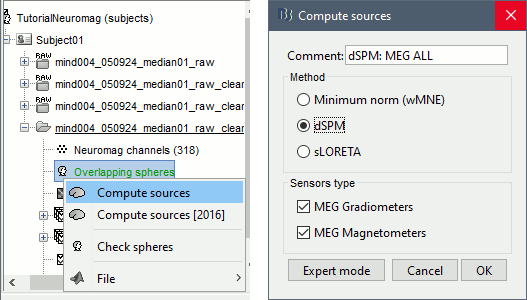
Select wMNE and all the default options. This will run a source reconstruction grouping the information from the magnetometers and the gradiometers into the same model. This is the recommended procedure, but you can also estimate sources for the two types of sensors separately if needed.
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
A shared inversion kernel is created in the (Common files) folder, and a link node is now visible for each recording file, single trials and averages.
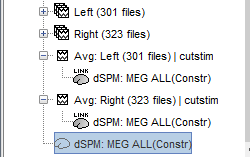
For more information: ?Source estimation tutorial.
Scouts
You can easily reproduce exactly the same results as the ones presented in the introduction tutorials. Place scouts to capture the activity in the primary and secondary somatosensory areas to track the processing of the electric stimulations in time, at the surface surface of the brain.
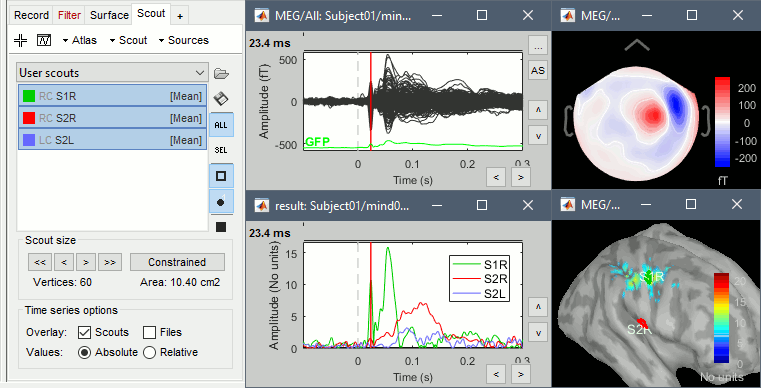
Scripting
The operations described in this tutorial can be reproduced from a Matlab script, available in the Brainstorm distribution: brainstorm3/toolbox/script/tutorial_neuromag.m
Feedback
