|
Size: 9196
Comment:
|
Size: 21680
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| The aim of this tutorial is to reproduce in the Brainstorm environment the analysis described in the SPM tutorial "[[ftp://ftp.mrc-cbu.cam.ac.uk/personal/rik.henson/wakemandg_hensonrn/Publications/SPM12_manual_chapter.pdf|Multimodal, Multisubject data fusion]]". The data processed here consists in simulateneous MEG/EEG recordings of 16 subjects performing simple visual task on a large number of famous, unfamiliar and scrambled faces. The analysis is split in two tutorial pages: the present tutorial describes the detailed analysis of one single subject and another one that the describes the batch processing and [[Tutorials/VisualGroup|group analysis of the 16 subjects]]. Note that the operations used here are not detailed, the goal of this tutorial is not to teach Brainstorm to a new inexperienced user. For in depth explanations of the interface and the theory, please refer to the introduction tutorials. |
The aim of this tutorial is to reproduce in the Brainstorm environment the analysis described in the SPM tutorial "[[ftp://ftp.mrc-cbu.cam.ac.uk/personal/rik.henson/wakemandg_hensonrn/Publications/SPM12_manual_chapter.pdf|Multimodal, Multisubject data fusion]]". The data processed here consists in simulateneous MEG/EEG recordings of 19 subjects performing simple visual task on a large number of famous, unfamiliar and scrambled faces. The analysis is split in two tutorial pages: the present tutorial describes the detailed analysis of one single subject and another one that the describes the batch processing and [[Tutorials/VisualGroup|group analysis of the 19 subjects]]. Note that the operations used here are not detailed, the goal of this tutorial is not to teach Brainstorm to a new inexperienced user. For in depth explanations of the interface and the theory, please refer to the [[http://neuroimage.usc.edu/brainstorm/Tutorials#Get_started|introduction tutorials]]. |
| Line 19: | Line 19: |
| * 16 subjects * 6 runs (sessions) of approximately 10mins for each subject |
* 19 subjects (three are excluded from the group analysis for various reasons) * 6 acquisition runs (sessions) of approximately 10mins for each subject |
| Line 31: | Line 31: |
| * MEG data have been "cleaned" using Signal-Space Separation as implemented in MaxFilter 2.1. * A Polhemus digitizer was used to digitise three fiducial points and a large number of other points across the scalp, which can be used to coregister the M/EEG data with the structural MRI image. * The distribution contains 3 sub-directories of empty-room recordings of 3-5mins acquired at roughly the same time of year (spring 2009) as the 16 subjects. The sub-directory names are Year (first 2 digits), Month (second 2 digits) and Day (third 2 digits). Inside each are 2 raw *.fif files: one for which basic SSS has been applied by maxfilter in a similar manner to the subject data above, and one (*-noSSS.fif) for which SSS has not been applied (though the data have been passed through maxfilter just to convert to float format). |
* MEG data have been "cleaned" using Signal-Space Separation as implemented in MaxFilter 2.2. * A Polhemus device was used to digitize three fiducial points and a large number of other points across the scalp, which can be used to coregister the M/EEG data with the structural MRI image. * Stimulation triggers: The triggers related with the visual presetation are saved in the STI101 channel, with the following event codes (bit 3 = face, bit 4 = unfamiliar, bit 5 = scrambled): * Famous faces: 5 (00101), 6 (00110), 7 (00111) * Unfamiliar faces: 13 (01101), 14 (01110), 15 (01111) * Scrambled images: 17 (10001), 18 (10010), 19 (10011) * Delays between the trigger in STI101 and the actual presentation of stimulus: '''34.5ms''' * The distribution includes MEG noise recordings acquired at a similar period, processed with MaxFilter 2.2 in a way similar to the subject recordings. |
| Line 40: | Line 45: |
| * The data is hosted on this FTP site (use an FTP client such as FileZilla, not your web browser): <<BR>>ftp://ftp.mrc-cbu.cam.ac.uk/personal/rik.henson/wakemandg_hensonrn/ * Download only the following folders (about 75Gb): * '''EmptyRoom''': MEG empty room measurements. * '''SubXX/MEEG''': MEG and EEG recordings in FIF format and triggers definition files. * '''Publications''': Reference publications related with this dataset. * '''README.TXT''': License and dataset description. * The FreeSurfer segmentations of the T1 images are not part of this package. You can either process them by yourself, or download the result of the segmentation from the Brainstorm website. <<BR>>Go to the [[http://neuroimage.usc.edu/bst/download.php|Download]] page, and download the file: '''sample_group_anat.zip'''<<BR>>Unzip this file in the same folder where you downloaded all the datasets. |
* The data is hosted on the OpenfMRI website: https://openfmri.org/dataset/ds000117/ * Download all the files available for dowload from this website (approximately 160Gb):<<BR>>ds117_metadata.tgz, ds117_sub001_raw.tgz, ..., ds117_sub019_raw.tgz * Unzip all the .tgz files in the same folder. * The FreeSurfer segmentations of the T1 images are not part of this package. You can either process them by yourself, or download the result of the segmentation from the Brainstorm website. <<BR>>Go to the [[http://neuroimage.usc.edu/bst/download.php|Download]] page, and download the file: '''sample_group_freesurfer.zip'''<<BR>>Unzip this file in the same folder where you downloaded all the datasets. * We will use the following files from this distribution: * '''/anatomy/subXXX/''': Segmentation folders generated with FreeSurfer. * '''/emptyroom/090707_raw_st.fif''': MEG empty room measurements, processed with MaxFilter. * '''/ds117/subXXX/MEG/*_sss.fif''': MEG and EEG recordings, processed with MaxFilter. * '''/README''': License and dataset description. |
| Line 55: | Line 62: |
| === FreeSurfer === This page explains how to import and process subject '''#002''' only. Subject #001 will be later excluded from the EEG group analysis because the position of the electrodes is incorrect, so it was not the best example. |
|
| Line 56: | Line 66: |
| * Right-click on the TutorialAuditory folder > New subject > '''Sub01''' | * Right-click on the TutorialAuditory folder > New subject > '''sub002''' |
| Line 60: | Line 70: |
| * Select the folder: '''Anatomy/Sub01''' (from sample_group_anat.zip) | * Select the folder: '''anatomy/freesurfer/sub002''' (from sample_group_freesurfer.zip) |
| Line 62: | Line 72: |
| * The two sets of fiducials we usually have to define interactively are here set automatically. | * The two sets of fiducials we usually have to define interactively are here automatically set. |
| Line 64: | Line 74: |
| * '''AC/PC/IH''': Identified automatically using the SPM affine registration with an MNI template. | * '''AC/PC/IH''': Automatically identified using the SPM affine registration with an MNI template. |
| Line 69: | Line 79: |
| * All the anatomical atlases [[Tutorials/LabelFreeSurfer|generated by FreeSurfer]] were imported automatically: the surface-based cortical atlases and the atlas of sub-cortical regions (ASEG). <<BR>><<BR>> {{attachment:anatomy_atlas.gif||height="211",width="550"}} | * All the anatomical atlases [[Tutorials/LabelFreeSurfer|generated by FreeSurfer]] were automatically imported: the surface-based cortical atlases and the atlas of sub-cortical regions (ASEG). <<BR>><<BR>> {{attachment:anatomy_atlas.gif||height="211",width="550"}} === BEM layers === Later in the tutorial, we will need to compute a [[Tutorials/TutBem|BEM forward model]] to estimate the brain sources from the EEG recordings. For this, we will need some layers defining the separation between the different tissues of the head (scalp, inner skull, outer skull). * Right-click on the subject folder > '''Generate BEM surfaces''': The number of vertices to use for each layer depend on your computing power and the accuracy you expect. You can try for instance with '''1082 vertices '''(scalp) and '''642 vertices '''(outer skull and innser skull). <<BR>><<BR>> {{attachment:anatomy_bem.gif}} |
| Line 73: | Line 88: |
| We need to attach the continuous .fif files containing the recordings to the database. |
|
| Line 76: | Line 93: |
| * Select all the FIF files in: Sub01'''/MEEG''' <<BR>><<BR>>SCREEN CAPTURE * Events: SCREEN CAPTURES * Refine registration now? '''NO''' * The head points that are available in the FIF files contain all the points that were digitized during the MEG acquisition, including the ones corresponding to the parts of the face that have been removed from the MRI. If we run the fitting algorithm, all the points around the nose will not match any close points on the head surface, leading to a wrong result. * If the registration based on the thre fiducials NAS/LPA/RPA, you can refine the registration manually. Right-click on the channel file > MRI registration > MEG : Edit... <<BR>><<BR>> {{attachment:registration.gif||height="222",width="251"}} * === Multiple runs and head position === * The two AEF runs 01 and 02 were acquired successively, the position of the subject's head in the MEG helmet was estimated twice, once at the beginning of each run. The subject might have moved between the two runs. To evaluate visually the displacement between the two runs, select at the same time all the channel files you want to compare (the ones for run 01 and 02), right-click > Display sensors > MEG. . {{http://neuroimage.usc.edu/brainstorm/Tutorials/Auditory?action=AttachFile&do=get&target=raw3.gif|raw3.gif|height="220",width="441",class="attachment"}} * Typically, we would like to group the trials coming from multiple runs by experimental conditions. However, because of the subject's movements between runs, it's not possible to directly compare the sensor values between runs because they probably do not capture the brain activity coming from the same regions of the brain. * You have three options if you consider grouping information from multiple runs: * '''Method 1''': Process all the runs separately and average between runs at the source level: The more accurate option, but requires a lot more work, computation time and storage. * '''Method 2''': Ignore movements between runs: This can be acceptable for commodity if the displacements are really minimal, less accurate but much faster to process and easier to manipulate. * '''Method 3''': Co-register properly the runs using the process Standardize > Co-register MEG runs: Can be a good option for displacements under 2cm. Warning: This method has not be been fully evaluated on our side, to use at your own risk. Also, it does not work correctly if you have different SSP projectors calculated for multiple runs. * In this tutorial, we will illustrate only method 1: runs are not co-registered. <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/VisualSingle)>> |
* Select the '''first''' FIF files in: '''Sub001/MEEG''' <<BR>><<BR>> {{attachment:review_raw.gif||height="202",width="542"}} * Events:''' Ignore'''<<BR>>We will read events for the stimulus channel and define trial definitions later.<<BR>><<BR>> {{attachment:review_ignore.gif||height="186",width="330"}} * Refine registration now? '''NO''' <<BR>>The head points that are available in the FIF files contain all the points that were digitized during the MEG acquisition, including the ones corresponding to the parts of the face that have been removed from the MRI. If we run the fitting algorithm, all the points around the nose will not match any close points on the head surface, leading to a wrong result. We will first remove the face points and then run the registration manually. === Channel classification === A few non-EEG channels are mixed in with the EEG channels, we need to change this before applying any operation on the EEG channels. * Right-click on the channel file > '''Edit channel file'''. Double-click on a cell to edit it. * Change the type of '''EEG062''' to '''EOG '''(electrooculogram). * Change the type of '''EEG063 '''to '''ECG '''(electrocardiogram). * Change the type of '''EEG061''' and '''EEG064''' to '''NOSIG'''. Close the window and save the modifications. <<BR>><<BR>> {{attachment:channel_edit.gif||height="252",width="561"}} === MRI registration === At this point, the registration MEG/MRI is based only the three fiducial points NAS/LPA/RPA. All the anatomical scans were anonymized (defaced) and for some subjects the nasion could not be defined properly. We will try to refine this registration using the additional head points that were digitized (only the points above the nasion). * Right-click on the channel file > '''Digitized head points > Remove points below nasion'''.<<BR>><<BR>> {{attachment:channel_remove.gif||height="194",width="329"}} * Right-click on the channel file > '''MRI registration > Refine using head points'''.<<BR>><<BR>> {{attachment:channel_refine.gif||height="173",width="294"}} * MEG/MRI registration, before (left) and after (right) this automatic registration procedure: <<BR>><<BR>> {{attachment:registration.gif||height="209",width="236"}} {{attachment:registration_final.gif||height="208",width="237"}} * Right-click on the channel file > MRI registration > EEG: Edit...<<BR>>Click on ['''Project electrodes on surface''']<<BR>><<BR>> {{attachment:channel_project.gif||height="207",width="477"}} * Close all the windows (use the [X] button at the top-right corner of the Brainstorm window). === Import event markers === We need to read the stimulus markers from the STI channels. The following tasks can be done in an interactive way with menus in the Record tab, as in the introduction tutorials. We will here illustrate how to do this with the pipeline editor, it will be easier to batch it for all the runs and all the subjects. * In Process1, select the "Link to raw file", click on [Run]. * Select process '''Events > Read from channel''', Channel: '''STI101''', Detection mode: '''Bit'''.<<BR>>Do not execute the process, we will add other processes to classify the markers.<<BR>><<BR>> {{attachment:raw_read_events.gif||height="290",width="499"}} * We want to create three categories of events: * Famous faces: 5 (00101), 6 (00110), 7 (00111) => Bit 3 only * Unfamiliar faces: 13 (01101), 14 (01110), 15 (01111) => Bit 3 and 4 * Scrambled images: 17 (10001), 18 (10010), 19 (10011) => Bit 5 only * We will start by creating the category "Unfamiliar" (combination of events "3" and "4") and remove the remove the initial events. Then we just have to rename the renaming "3" in "Famous", and all the "5" in "Scrambled". * Add process '''Events > Group by name''': "'''Unfamiliar=3,4'''", Delay=0, '''Delete original events''' * Add process '''Events > Rename event''': 3 => Famous * Add process '''Events > Rename event''': 5 => Scrambled<<BR>><<BR>> {{attachment:events_merge.gif||height="402",width="532"}} * Add process '''Events > Add time offset''' to correct for the presentation delays:<<BR>>Event names: "'''Famous, Unfamiliar, Scrambled'''", Time offset = '''34.5ms'''<<BR>><<BR>> {{attachment:events_offset.gif||height="355",width="348"}} * Finally run the script. Double-click on the recordings to make sure the labels were detected correctly. You can delete the unwanted events that are left in the recordings (1,2,9,13): <<BR>><<BR>> {{attachment:events_display.gif||height="194",width="595"}} == Pre-processing == === Spectral evaluation === * Keep the "Link to raw file" in Process1. * Run process '''Frequency > Power spectrum density (Welch)''' with the options illustrated below. <<BR>><<BR>> {{attachment:psd_process.gif||height="314",width="669"}} * Right-click on the PSD file > Power spectrum. <<BR>><<BR>> {{attachment:psd_plot.gif||height="315",width="546"}} * Observations: * Three groups of sensors, from top to bottom: EEG, MEG gradiometers, MEG magnetometers. * Power lines: '''50 '''Hz and harmonics * Alpha peak around 10 Hz * Artifacts due to Elekta electronics: '''293'''Hz, '''307'''Hz, '''314'''Hz, '''321'''Hz, '''328'''Hz. * Suspected bad EEG channels: '''EEG016''' * Close all the windows. === Remove line noise === * Keep the "Link to raw file" in Process1. * Select process '''Pre-process > Notch filter''' to remove the line noise (50-200Hz).<<BR>>Add immediately after the process "'''Frequency > Power spectrum density (Welch)'''" <<BR>><<BR>> {{attachment:notch_process.gif||height="262",width="562"}} * Double-click on the PSD for the new continuous file to evaluate the quality of the correction. <<BR>><<BR>> {{attachment:notch_result.gif||height="215",width="566"}} * Close all the windows (use the [X] button at the top-right corner of the Brainstorm window). === EEG reference and bad channels === * Right-click on link to the processed file ("Raw | notch(50Hz ...") > '''EEG > Display time series'''. * Select channel '''EEG016''' and mark it as '''bad''' (using the popup menu or pressing the Delete key). <<BR>><<BR>> {{attachment:channel_bad.gif||height="214",width="587"}} * In the Record tab, menu '''Artifacts > Re-reference EEG''' > "AVERAGE". <<BR>><<BR>> {{attachment:channel_ref.gif||height="270",width="535"}} * At the end, the window "select active projectors" is open to show the new re-referencing projector. Just close this window. To get it back, use the menu Artifacts > Select active projectors. == Artifact correction with SSP == === Artifact detection === * Empty the Process1 list (right-click > Clear list). * Drag and drop the continuous processed file ("Raw | notch(50Hz...)") to the Process1 list. * Run process''' Events > Detect heartbeats''': Channel name='''EEG063''', All file, Event name=cardiac <<BR>><<BR>> {{attachment:detect_cardiac.gif||height="230",width="310"}} * In many of the other tutorials, you will have detected the blink category and used the SSP process to remove the blink artifact. In this experiment, we are particularly interested in the subject's response to seeing the stimulus. Therefore we will mark eyeblinks and other eye movements as BAD. * Run process '''Artifacts > Detect events above threshold''': Event name='''blink_BAD''', Channel='''EEG062''', All file, Maximum threshold='''100''', Threshold units='''uV''', Filter='''[0.30,20.00]Hz''', '''Use absolute value of signal'''.<<BR>><<BR>> {{attachment:detect_blinks.gif||height="455",width="285"}} * Inspect visually the two new cateogries: cardiac and blink_BAD.<<BR>><<BR>> {{attachment:detect_display.gif||height="237",width="655"}} * Close all the windows (using the [X] button). === Heartbeats === * Keep the "Raw | notch" file selected in Process1. * Select process '''Artifacts > SSP: Heartbeats''' > Sensor type: '''MEG MAG''' * Add process '''Artifacts > SSP: Heartbeats''' > Sensor type: '''MEG GRAD''' - Run the execution <<BR>><<BR>> {{attachment:ssp_ecg_process.gif||height="225",width="293"}} * Double-click on the continuous file to show all the MEG sensors. <<BR>>In the Record tab, select sensors "Left-temporal". * Menu Artifacts > Select active projectors. * In category '''cardiac/MEG MAG''': Select '''component #1''' and view topography. * In category '''cardiac/MEG GRAD''': Select '''component #1 '''and view topography. <<BR>><<BR>> {{attachment:ssp_ecg_topo.gif||height="175",width="604"}} * Make sure that selecting the two components removes the cardiac artifact. Then click '''[Save]'''. === Additional bad segments === * Process1>Run: Select the process "'''Events > Detect other artifacts'''". This should be done separately for MEG and EEG to avoid confusion about which sensors are involved in the artifact. <<BR>><<BR>> {{attachment:detect_other.gif||height="253",width="486"}} * Display the MEG sensors. Review the segments that were tagged as artifact, determine if each event represents an artifact and then mark the time of the artifact as BAD. This can be done by selecting the time window around the artifact, Right-click > '''Reject time segment'''. Note that this detection process marks 1-second segments but the artifact can be shorter.<<BR>><<BR>> {{attachment:artifacts_mark_bad.png||width="640"}} * Once all the events in the two categories are reviewed and bad segments are marked, the two categories (1-7Hz and 40-240Hz) can be deleted. * Do this detection and review again for the EEG. === SQUID jumps === MEG signals recorded with Elekta-Neuromag systems frequently contain SQUID jumps ([[http://neuroimage.usc.edu/brainstorm/Tutorials/BadSegments?highlight=(squid)#Elekta-Neuromag_SQUID_jumps|more information]]). These sharp steps followed by a change of baseline value are usually easy to identify visually but more complicated to detect automatically. The process "Detect other artifacts" usually detects most of them in the category "1-7Hz". If you observe that some are skipped, you can try re-running it with a high sensitivity. Tt is important to review all the sensors and all the time in each recording to be sure these events are marked as bad segments. . {{attachment:detect_jumps.gif||height="214",width="547"}} == Epoching and averaging == === Import epochs === * Keep the "Raw | notch" file selected in Process1. * Select process: '''Import recordings > Import MEG/EEG: Events''' (do not run immediately)<<BR>>Event names "Famous, Unfamiliar, Scrambled", All file, Epoch time=[-300,1000]ms * Add process: '''Pre-process > Remove DC offset''': Baseline=[-300,-0.9]ms - Run exectuion.<<BR>><<BR>> {{attachment:import_epochs.gif||height="399",width="647"}} === Average === * In Process1, select all the imported trials. * Run process: '''Average > Average files''': '''By trial groups (folder average)''' <<BR>><<BR>> {{attachment:average_process.gif||height="504",width="478"}} === Review EEG ERP === * EEG evoked response (famous, scrambled, unfamiliar): <<BR>><<BR>> {{attachment:average_topo.gif||height="392",width="684"}} * Open the [[Tutorials/ChannelClusters|Cluster tab]] and create a cluster with the channel '''EEG065''' (button [NEW IND]).<<BR>><<BR>> {{attachment:cluster_create.gif||height="166",width="569"}} * Select the cluster then select three average files, right-click > '''Clusters time series''' (Overlay:Files). <<BR>><<BR>> {{attachment:cluster_display.gif||height="220",width="658"}} * Basic observations for EEG065 (right parieto-occiptal electrode): * Around 170ms (N170): greater negative deflection for Famous than Scrambled faces. * After 250ms: difference between Famous and Unfamiliar faces. == Empty room recordings == === MEG === For this dataset, one empty room recording will be used for all subjects. * Create a new subject: '''emptyroom''' * Create a link to the raw file: '''sample_group/emptyroom/090707_raw_st.fif''' * Apply the same '''notch filter''' as for the data * Compute the noise covariance. Right-click on''' Raw | notch > Noise covariance > Compute from recordings''' * Copy the noise covariance to Sub001. Right-click on the '''Noise covariance > Copy to other subjects '''<<BR>><<BR>> {{attachment:emptyroom.png||height="180",width="300"}} === EEG === Compute a EEG noise covariance for EACH RUN using the EEG recordings and merge it with the existing noise covariance, which contains the noise covariance for MEG. * Expand the folder run_01_sss_notch to see the imported files. Select all imported trials, Right-click >''' Noise covariance > Compute from recordings''', Time window=[-300,0]ms, Sensor types=EEG, select '''Merge'''. * Repeat for runs 2-6. == Source estimation == === Head Model === * Switch back to the "Functional" view. Drag and drop the 6 imported trial folders in Process1. * Run > '''Sources > Compute head model''': Overlapping spheres (MEG) and OpenMEEG BEM (EEG). It will take quite a long time to compute the BEM models using the default values. === Source Model === * Drop all imported files into the Process1 box * MEG: Run > '''Sources > Compute sources''', Sensor types=MEG, Leave all the default options to calculate a wMNE solution with constrained dipole orientation. Click Run. * EEG: Run > '''Sources > Compute sources''', Sensor types=EEG, Leave all the default options to calculate a wMNE solution with constrained dipole orientation. == Other subjects == * sub001: Bad registration * sub006: Lots of blinks * sub016: Lots of SQUID jumps == Scripting == Corresponding script in the Brainstorm distribution:<<BR>> '''TODO: Coming soon.''' . <<EmbedContent(http://neuroimage.usc.edu/bst/get_feedback.php?Tutorials/VisualSingle)>> |
MEG visual tutorial: Single subject
Authors: Francois Tadel, Elizabeth Bock.
The aim of this tutorial is to reproduce in the Brainstorm environment the analysis described in the SPM tutorial "Multimodal, Multisubject data fusion". The data processed here consists in simulateneous MEG/EEG recordings of 19 subjects performing simple visual task on a large number of famous, unfamiliar and scrambled faces.
The analysis is split in two tutorial pages: the present tutorial describes the detailed analysis of one single subject and another one that the describes the batch processing and group analysis of the 19 subjects.
Note that the operations used here are not detailed, the goal of this tutorial is not to teach Brainstorm to a new inexperienced user. For in depth explanations of the interface and the theory, please refer to the introduction tutorials.
Contents
License
These data are provided freely for research purposes only (as part of their Award of the BioMag2010 Data Competition). If you wish to publish any of these data, please acknowledge Daniel Wakeman and Richard Henson. The best single reference is: Wakeman DG, Henson RN, A multi-subject, multi-modal human neuroimaging dataset, Scientific Data (2015)
Any questions, please contact: rik.henson@mrc-cbu.cam.ac.uk
Presentation of the experiment
Experiment
- 19 subjects (three are excluded from the group analysis for various reasons)
- 6 acquisition runs (sessions) of approximately 10mins for each subject
- Presentation of series of images: familiar faces, unfamiliar faces, phase-scrambled faces
- The subject has to judge the left-right symmetry of each stimulus
- Total of nearly 300 trials in total for each of the 3 conditions
MEG acquisition
Acquisition at 1100Hz with an Elekta-Neuromag VectorView system (simultaneous MEG+EEG).
- Recorded channels (404):
- 102 magnetometers
- 204 planar gradiometers
- 70 EEG electrodes recorded with a nose reference.
MEG data have been "cleaned" using Signal-Space Separation as implemented in MaxFilter 2.2.
- A Polhemus device was used to digitize three fiducial points and a large number of other points across the scalp, which can be used to coregister the M/EEG data with the structural MRI image.
- Stimulation triggers: The triggers related with the visual presetation are saved in the STI101 channel, with the following event codes (bit 3 = face, bit 4 = unfamiliar, bit 5 = scrambled):
- Famous faces: 5 (00101), 6 (00110), 7 (00111)
- Unfamiliar faces: 13 (01101), 14 (01110), 15 (01111)
- Scrambled images: 17 (10001), 18 (10010), 19 (10011)
Delays between the trigger in STI101 and the actual presentation of stimulus: 34.5ms
The distribution includes MEG noise recordings acquired at a similar period, processed with MaxFilter 2.2 in a way similar to the subject recordings.
Subject anatomy
- MRI data acquired on a 3T Siemens TIM Trio: 1x1x1mm T1-weighted structural MRI
Processed with FreeSurfer 5.3
Download and installation
The data is hosted on the OpenfMRI website: https://openfmri.org/dataset/ds000117/
Download all the files available for dowload from this website (approximately 160Gb):
ds117_metadata.tgz, ds117_sub001_raw.tgz, ..., ds117_sub019_raw.tgz- Unzip all the .tgz files in the same folder.
The FreeSurfer segmentations of the T1 images are not part of this package. You can either process them by yourself, or download the result of the segmentation from the Brainstorm website.
Go to the Download page, and download the file: sample_group_freesurfer.zip
Unzip this file in the same folder where you downloaded all the datasets.- We will use the following files from this distribution:
/anatomy/subXXX/: Segmentation folders generated with FreeSurfer.
/emptyroom/090707_raw_st.fif: MEG empty room measurements, processed with MaxFilter.
/ds117/subXXX/MEG/*_sss.fif: MEG and EEG recordings, processed with MaxFilter.
/README: License and dataset description.
- Reminder: Do not put the downloaded files in the Brainstorm folders (program or database folders).
Start Brainstorm (Matlab scripts or stand-alone version). For help, see the Installation page.
Select the menu File > Create new protocol. Name it "TutorialVisual" and select the options:
"No, use individual anatomy",
"No, use one channel file per condition".
Import the anatomy
FreeSurfer
This page explains how to import and process subject #002 only. Subject #001 will be later excluded from the EEG group analysis because the position of the electrodes is incorrect, so it was not the best example.
- Switch to the "anatomy" view.
Right-click on the TutorialAuditory folder > New subject > sub002
- Leave the default options you set for the protocol
Right-click on the subject node > Import anatomy folder:
Set the file format: "FreeSurfer folder"
Select the folder: anatomy/freesurfer/sub002 (from sample_group_freesurfer.zip)
- Number of vertices of the cortex surface: 15000 (default value)
- The two sets of fiducials we usually have to define interactively are here automatically set.
NAS/LPA/RPA: The file Anatomy/Sub01/fiducials.m contains the definition of the nasion, left and right ears. The anatomical points used by the authors are the same as the ones we recommend in the Brainstorm coordinates systems page.
AC/PC/IH: Automatically identified using the SPM affine registration with an MNI template.
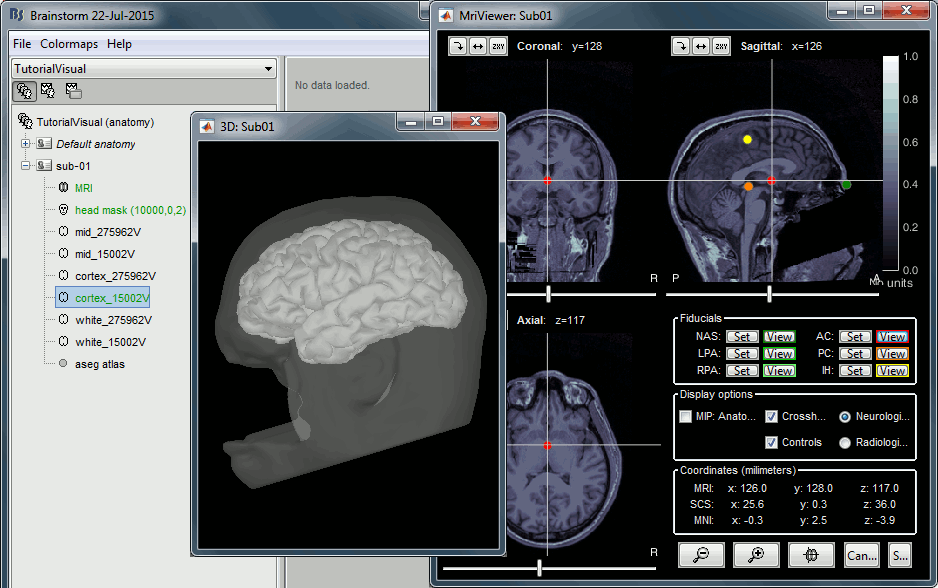
If you want to double-check that all these points were correctly marked after importing the anatomy, right-click on the MRI > Edit MRI.
At the end of the process, make sure that the file "cortex_15000V" is selected (downsampled pial surface, that will be used for the source estimation). If it is not, double-click on it to select it as the default cortex surface. Do not worry about the big holes in the head surface, parts of MRI have been remove voluntarily for anonymization purposes.
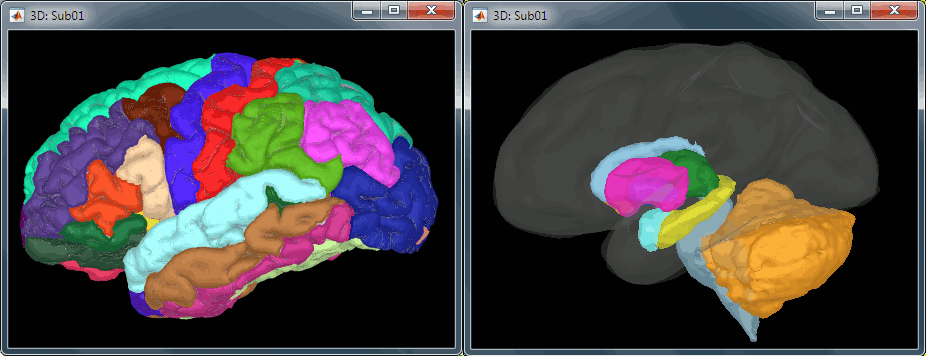
All the anatomical atlases generated by FreeSurfer were automatically imported: the surface-based cortical atlases and the atlas of sub-cortical regions (ASEG).
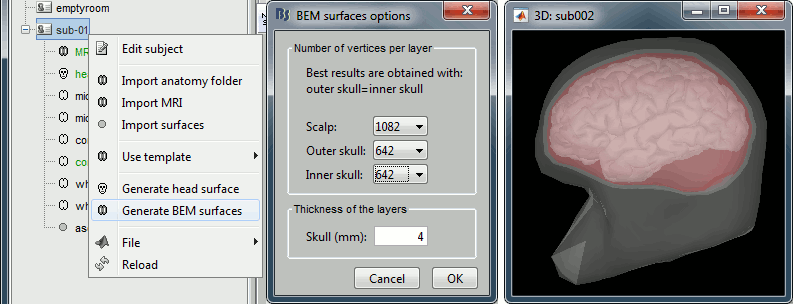
BEM layers
Later in the tutorial, we will need to compute a BEM forward model to estimate the brain sources from the EEG recordings. For this, we will need some layers defining the separation between the different tissues of the head (scalp, inner skull, outer skull).
Right-click on the subject folder > Generate BEM surfaces: The number of vertices to use for each layer depend on your computing power and the accuracy you expect. You can try for instance with 1082 vertices (scalp) and 642 vertices (outer skull and innser skull).
Access the recordings
Link the recordings
We need to attach the continuous .fif files containing the recordings to the database.
- Switch to the "functional data" view.
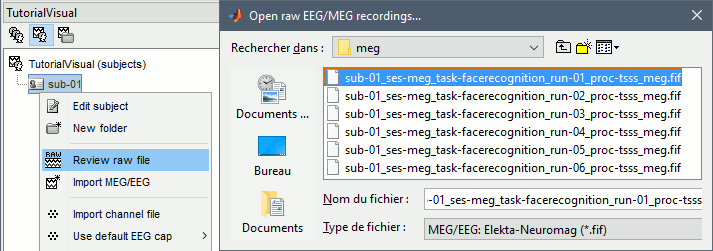
Right-click on the subject folder > Review raw file.
Select the file format: "MEG/EEG: Neuromag FIFF (*.fif)"
Select the first FIF files in: Sub001/MEEG
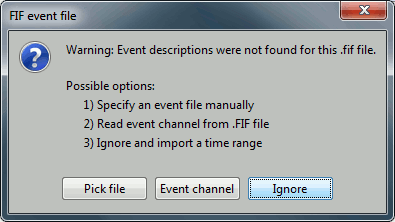
Events: Ignore
We will read events for the stimulus channel and define trial definitions later.
Refine registration now? NO
The head points that are available in the FIF files contain all the points that were digitized during the MEG acquisition, including the ones corresponding to the parts of the face that have been removed from the MRI. If we run the fitting algorithm, all the points around the nose will not match any close points on the head surface, leading to a wrong result. We will first remove the face points and then run the registration manually.
Channel classification
A few non-EEG channels are mixed in with the EEG channels, we need to change this before applying any operation on the EEG channels.
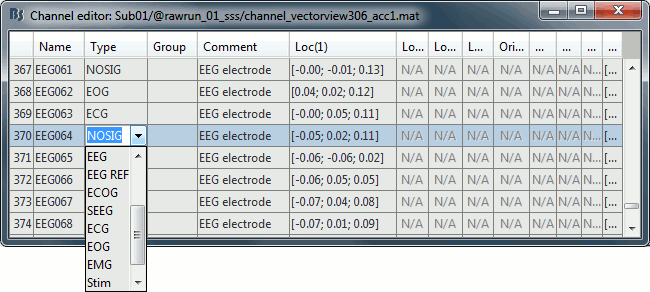
Right-click on the channel file > Edit channel file. Double-click on a cell to edit it.
Change the type of EEG062 to EOG (electrooculogram).
Change the type of EEG063 to ECG (electrocardiogram).
Change the type of EEG061 and EEG064 to NOSIG. Close the window and save the modifications.
MRI registration
At this point, the registration MEG/MRI is based only the three fiducial points NAS/LPA/RPA. All the anatomical scans were anonymized (defaced) and for some subjects the nasion could not be defined properly. We will try to refine this registration using the additional head points that were digitized (only the points above the nasion).
Right-click on the channel file > Digitized head points > Remove points below nasion.
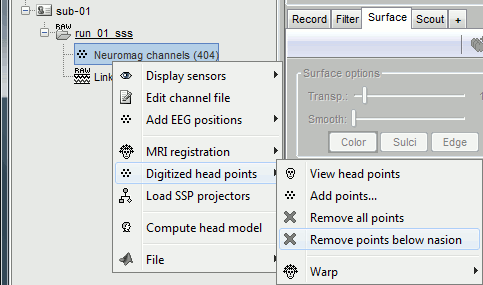
Right-click on the channel file > MRI registration > Refine using head points.
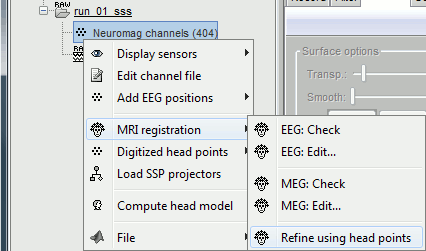
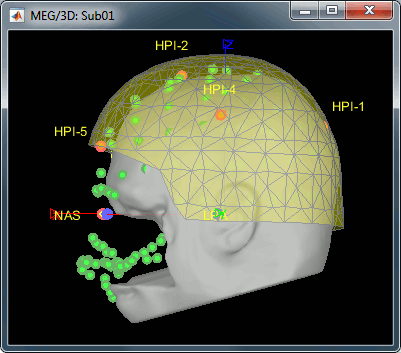
MEG/MRI registration, before (left) and after (right) this automatic registration procedure:
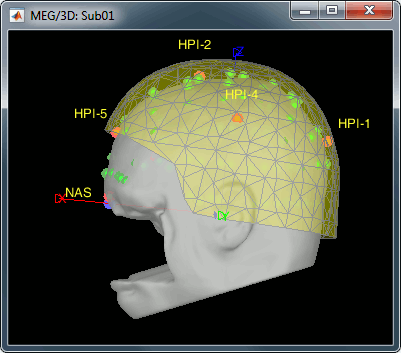
Right-click on the channel file > MRI registration > EEG: Edit...
Click on [Project electrodes on surface]
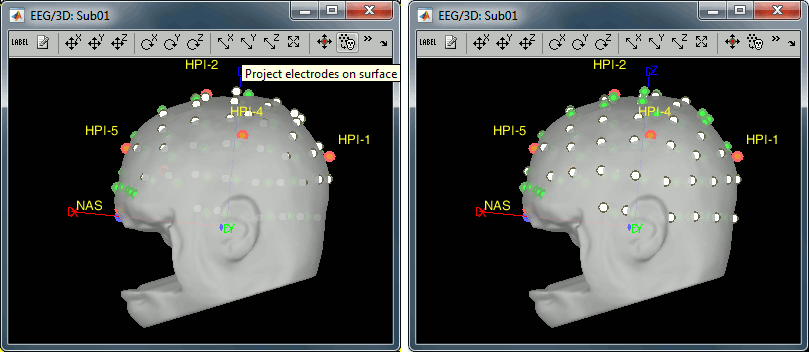
- Close all the windows (use the [X] button at the top-right corner of the Brainstorm window).
Import event markers
We need to read the stimulus markers from the STI channels. The following tasks can be done in an interactive way with menus in the Record tab, as in the introduction tutorials. We will here illustrate how to do this with the pipeline editor, it will be easier to batch it for all the runs and all the subjects.
- In Process1, select the "Link to raw file", click on [Run].
Select process Events > Read from channel, Channel: STI101, Detection mode: Bit.
Do not execute the process, we will add other processes to classify the markers.
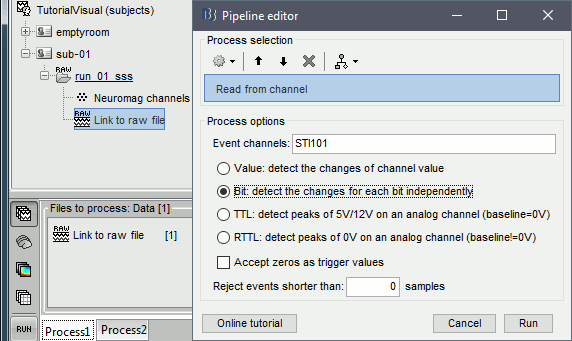
- We want to create three categories of events:
Famous faces: 5 (00101), 6 (00110), 7 (00111) => Bit 3 only
Unfamiliar faces: 13 (01101), 14 (01110), 15 (01111) => Bit 3 and 4
Scrambled images: 17 (10001), 18 (10010), 19 (10011) => Bit 5 only
- We will start by creating the category "Unfamiliar" (combination of events "3" and "4") and remove the remove the initial events. Then we just have to rename the renaming "3" in "Famous", and all the "5" in "Scrambled".
Add process Events > Group by name: "Unfamiliar=3,4", Delay=0, Delete original events
Add process Events > Rename event: 3 => Famous
Add process Events > Rename event: 5 => Scrambled
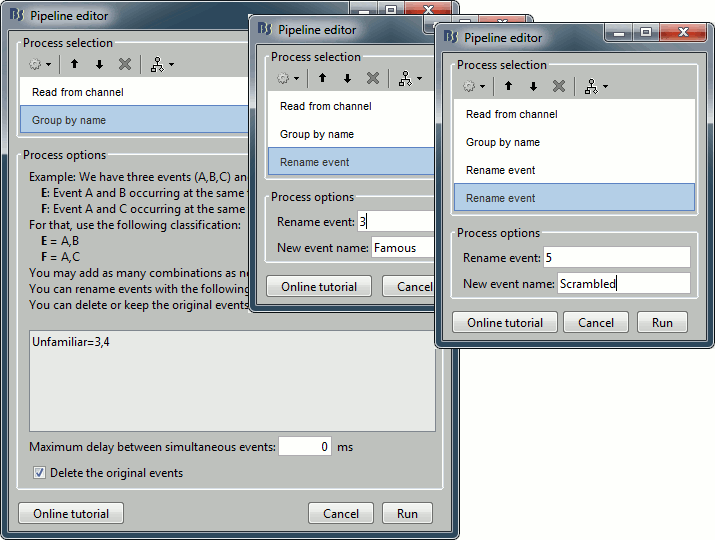
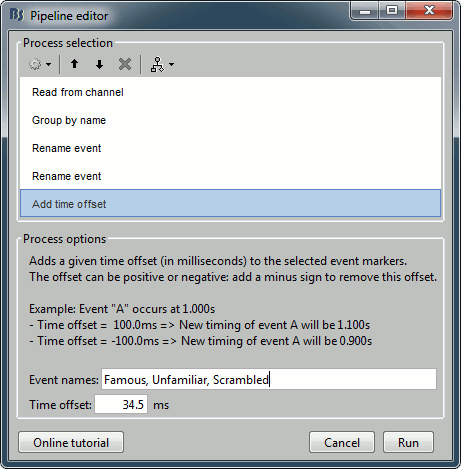
Add process Events > Add time offset to correct for the presentation delays:
Event names: "Famous, Unfamiliar, Scrambled", Time offset = 34.5ms
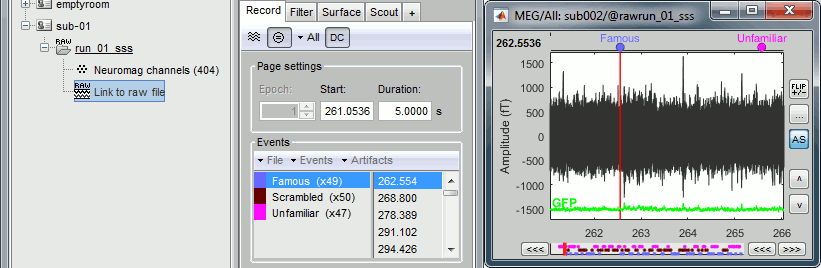
Finally run the script. Double-click on the recordings to make sure the labels were detected correctly. You can delete the unwanted events that are left in the recordings (1,2,9,13):
Pre-processing
Spectral evaluation
- Keep the "Link to raw file" in Process1.
Run process Frequency > Power spectrum density (Welch) with the options illustrated below.
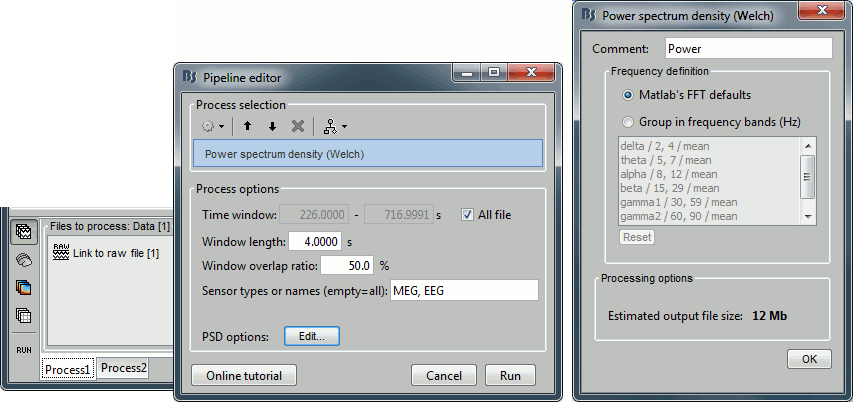
Right-click on the PSD file > Power spectrum.
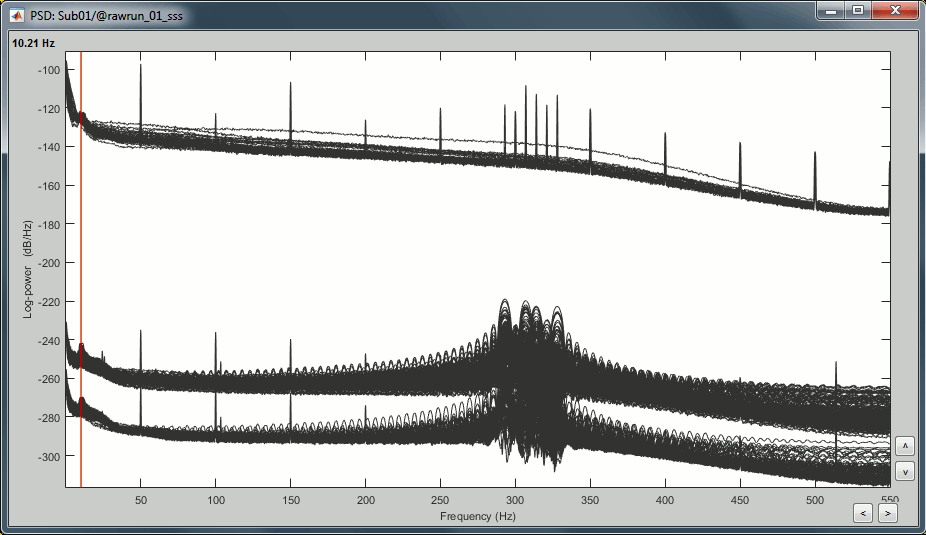
- Observations:
- Three groups of sensors, from top to bottom: EEG, MEG gradiometers, MEG magnetometers.
Power lines: 50 Hz and harmonics
- Alpha peak around 10 Hz
Artifacts due to Elekta electronics: 293Hz, 307Hz, 314Hz, 321Hz, 328Hz.
Suspected bad EEG channels: EEG016
- Close all the windows.
Remove line noise
- Keep the "Link to raw file" in Process1.
Select process Pre-process > Notch filter to remove the line noise (50-200Hz).
Add immediately after the process "Frequency > Power spectrum density (Welch)"
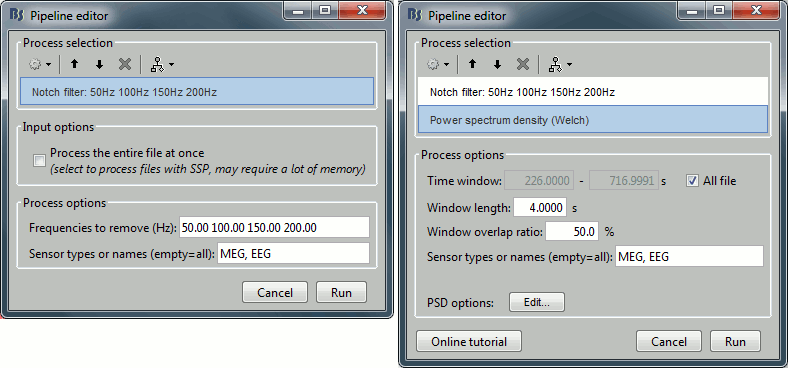
Double-click on the PSD for the new continuous file to evaluate the quality of the correction.
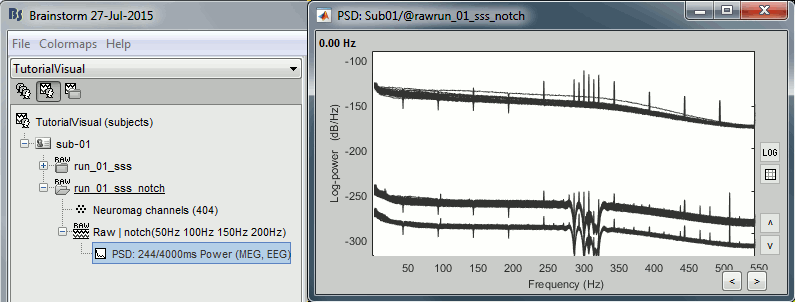
- Close all the windows (use the [X] button at the top-right corner of the Brainstorm window).
EEG reference and bad channels
Right-click on link to the processed file ("Raw | notch(50Hz ...") > EEG > Display time series.
Select channel EEG016 and mark it as bad (using the popup menu or pressing the Delete key).
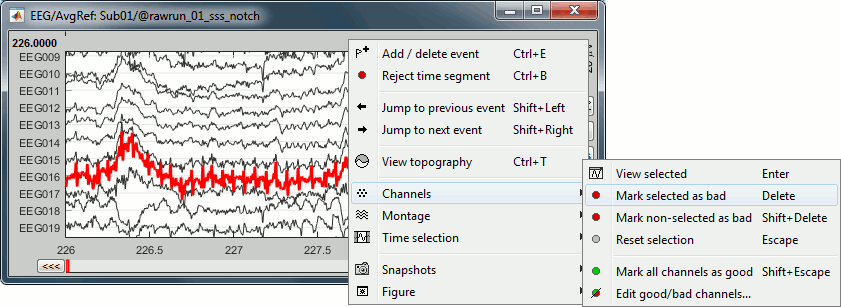
In the Record tab, menu Artifacts > Re-reference EEG > "AVERAGE".
At the end, the window "select active projectors" is open to show the new re-referencing projector. Just close this window. To get it back, use the menu Artifacts > Select active projectors.
Artifact correction with SSP
Artifact detection
Empty the Process1 list (right-click > Clear list).
- Drag and drop the continuous processed file ("Raw | notch(50Hz...)") to the Process1 list.
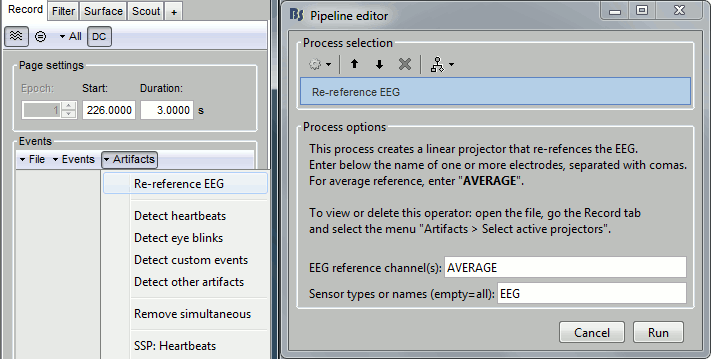
Run process Events > Detect heartbeats: Channel name=EEG063, All file, Event name=cardiac
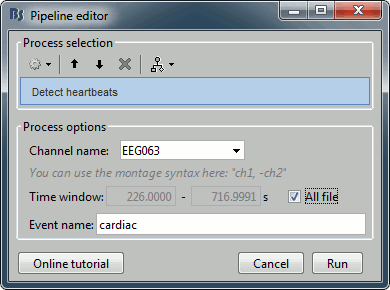
- In many of the other tutorials, you will have detected the blink category and used the SSP process to remove the blink artifact. In this experiment, we are particularly interested in the subject's response to seeing the stimulus. Therefore we will mark eyeblinks and other eye movements as BAD.
Run process Artifacts > Detect events above threshold: Event name=blink_BAD, Channel=EEG062, All file, Maximum threshold=100, Threshold units=uV, Filter=[0.30,20.00]Hz, Use absolute value of signal.
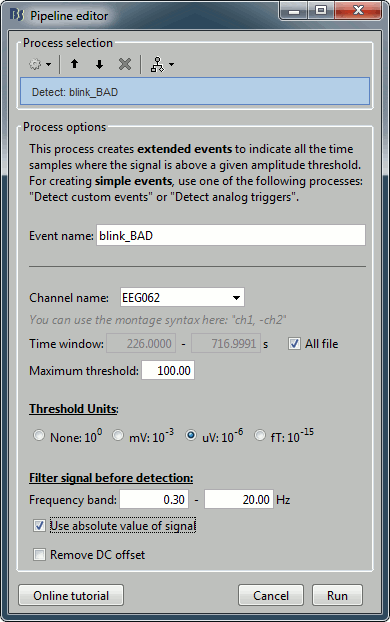
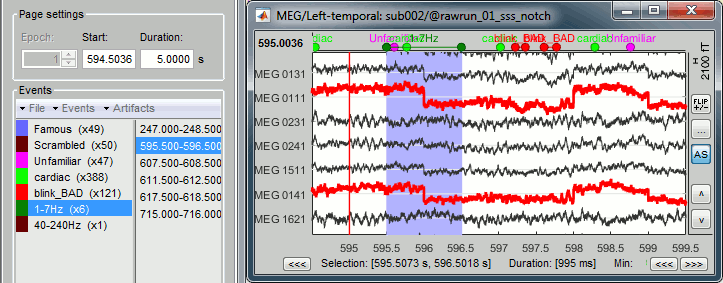
Inspect visually the two new cateogries: cardiac and blink_BAD.
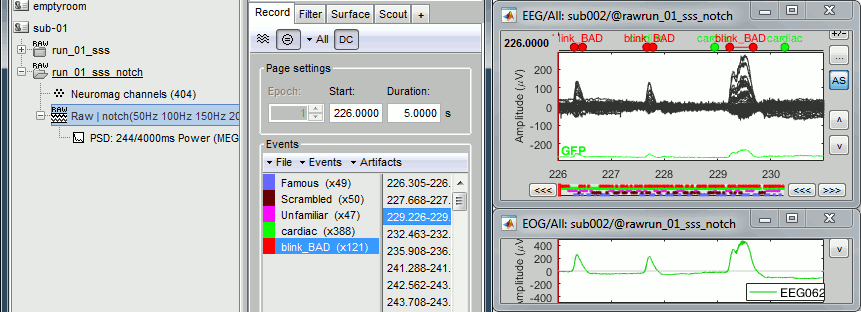
- Close all the windows (using the [X] button).
Heartbeats
- Keep the "Raw | notch" file selected in Process1.
Select process Artifacts > SSP: Heartbeats > Sensor type: MEG MAG
Add process Artifacts > SSP: Heartbeats > Sensor type: MEG GRAD - Run the execution
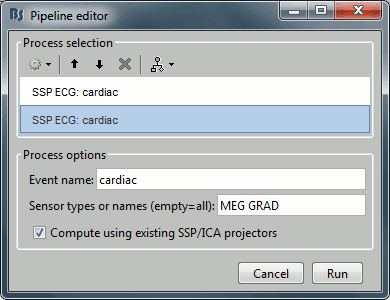
Double-click on the continuous file to show all the MEG sensors.
In the Record tab, select sensors "Left-temporal".Menu Artifacts > Select active projectors.
In category cardiac/MEG MAG: Select component #1 and view topography.
In category cardiac/MEG GRAD: Select component #1 and view topography.
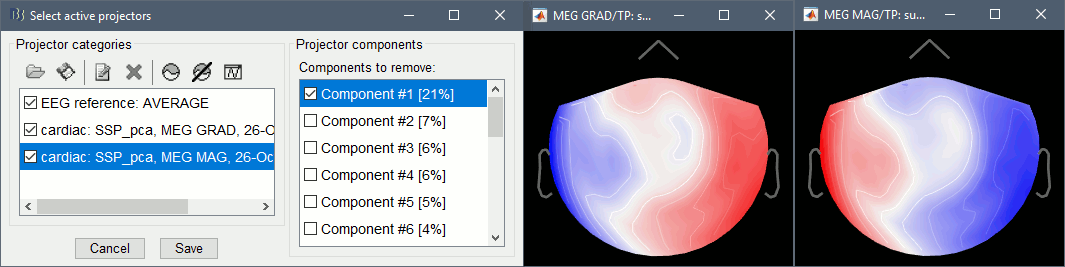
Make sure that selecting the two components removes the cardiac artifact. Then click [Save].
Additional bad segments
Process1>Run: Select the process "Events > Detect other artifacts". This should be done separately for MEG and EEG to avoid confusion about which sensors are involved in the artifact.
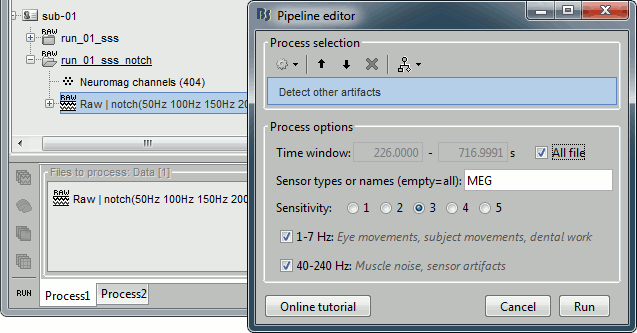
Display the MEG sensors. Review the segments that were tagged as artifact, determine if each event represents an artifact and then mark the time of the artifact as BAD. This can be done by selecting the time window around the artifact, Right-click > Reject time segment. Note that this detection process marks 1-second segments but the artifact can be shorter.
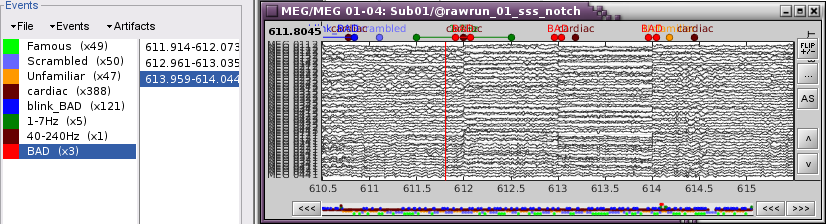
- Once all the events in the two categories are reviewed and bad segments are marked, the two categories (1-7Hz and 40-240Hz) can be deleted.
- Do this detection and review again for the EEG.
SQUID jumps
MEG signals recorded with Elekta-Neuromag systems frequently contain SQUID jumps (more information). These sharp steps followed by a change of baseline value are usually easy to identify visually but more complicated to detect automatically.
The process "Detect other artifacts" usually detects most of them in the category "1-7Hz". If you observe that some are skipped, you can try re-running it with a high sensitivity. Tt is important to review all the sensors and all the time in each recording to be sure these events are marked as bad segments.
Epoching and averaging
Import epochs
- Keep the "Raw | notch" file selected in Process1.
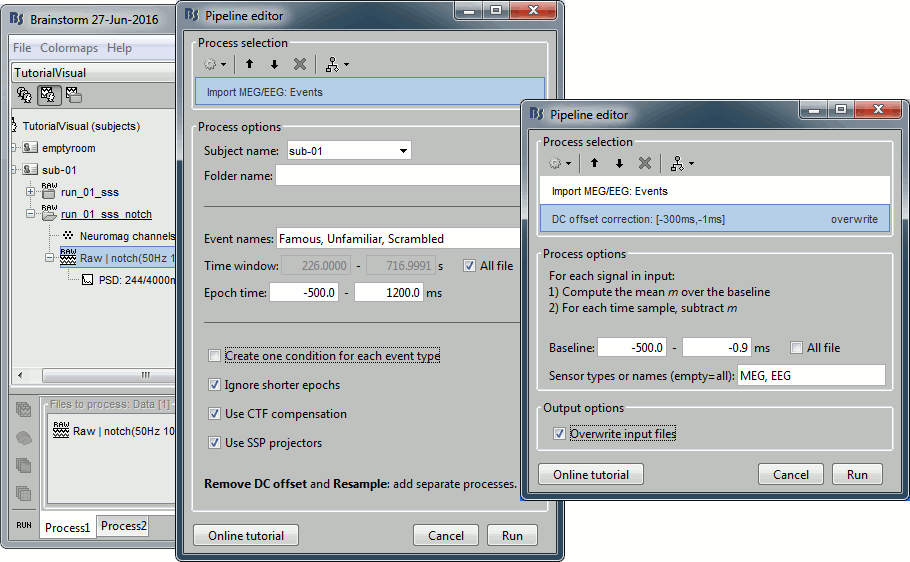
Select process: Import recordings > Import MEG/EEG: Events (do not run immediately)
Event names "Famous, Unfamiliar, Scrambled", All file, Epoch time=[-300,1000]msAdd process: Pre-process > Remove DC offset: Baseline=[-300,-0.9]ms - Run exectuion.
Average
- In Process1, select all the imported trials.
Run process: Average > Average files: By trial groups (folder average)
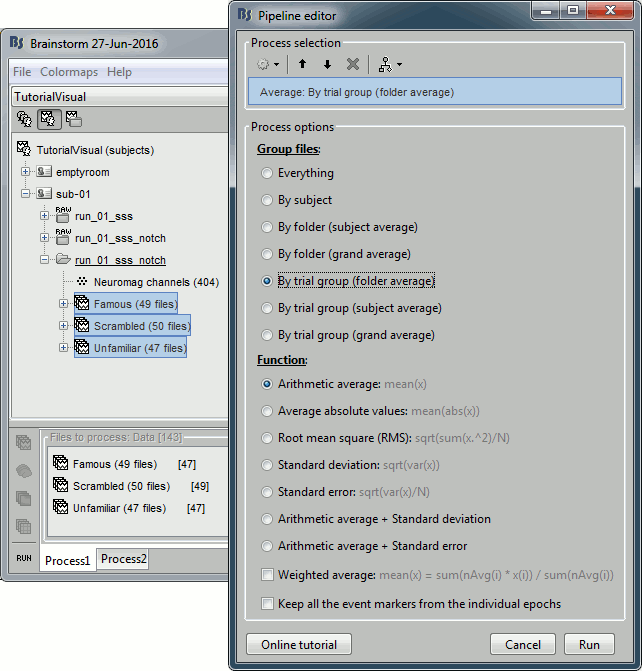
Review EEG ERP
EEG evoked response (famous, scrambled, unfamiliar):
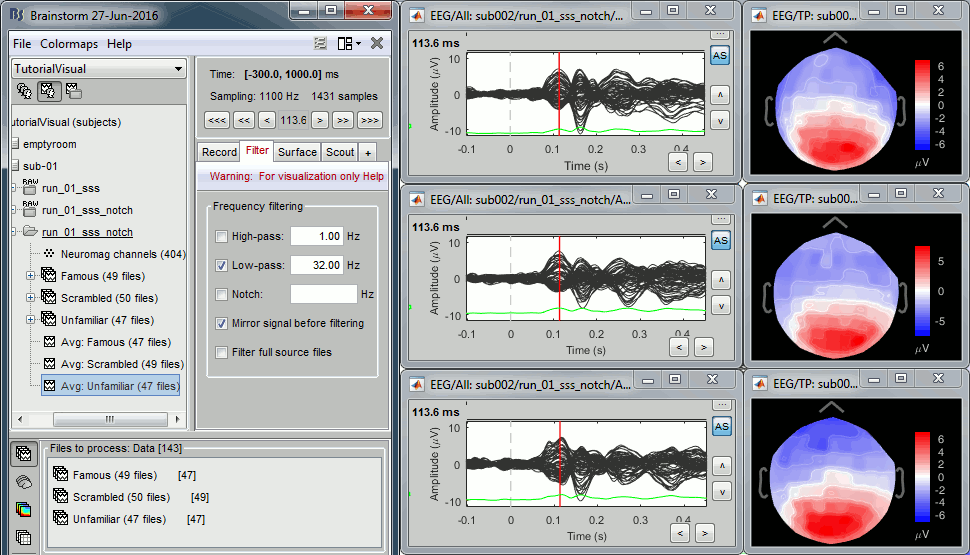
Open the Cluster tab and create a cluster with the channel EEG065 (button [NEW IND]).
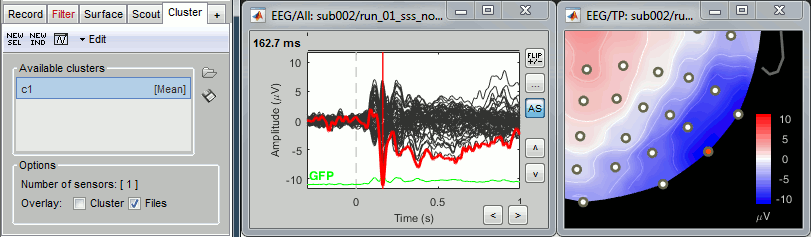
Select the cluster then select three average files, right-click > Clusters time series (Overlay:Files).
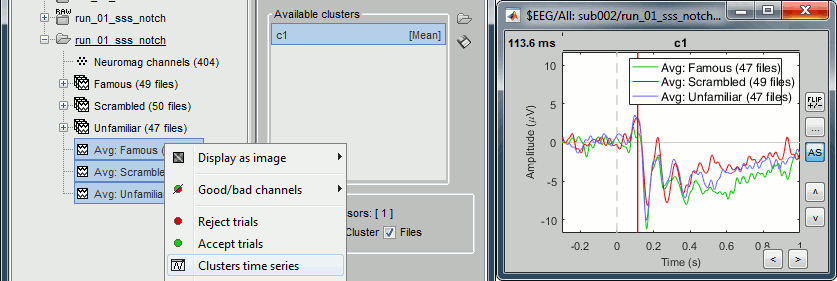
- Basic observations for EEG065 (right parieto-occiptal electrode):
- Around 170ms (N170): greater negative deflection for Famous than Scrambled faces.
- After 250ms: difference between Famous and Unfamiliar faces.
Empty room recordings
MEG
For this dataset, one empty room recording will be used for all subjects.
Create a new subject: emptyroom
Create a link to the raw file: sample_group/emptyroom/090707_raw_st.fif
Apply the same notch filter as for the data
Compute the noise covariance. Right-click on Raw | notch > Noise covariance > Compute from recordings
Copy the noise covariance to Sub001. Right-click on the Noise covariance > Copy to other subjects
![[ATTACH] [ATTACH]](/moin_static1911/brainstorm1/img/attach.png)
EEG
Compute a EEG noise covariance for EACH RUN using the EEG recordings and merge it with the existing noise covariance, which contains the noise covariance for MEG.
Expand the folder run_01_sss_notch to see the imported files. Select all imported trials, Right-click > Noise covariance > Compute from recordings, Time window=[-300,0]ms, Sensor types=EEG, select Merge.
- Repeat for runs 2-6.
Source estimation
Head Model
- Switch back to the "Functional" view. Drag and drop the 6 imported trial folders in Process1.
Run > Sources > Compute head model: Overlapping spheres (MEG) and OpenMEEG BEM (EEG). It will take quite a long time to compute the BEM models using the default values.
Source Model
- Drop all imported files into the Process1 box
MEG: Run > Sources > Compute sources, Sensor types=MEG, Leave all the default options to calculate a wMNE solution with constrained dipole orientation. Click Run.
EEG: Run > Sources > Compute sources, Sensor types=EEG, Leave all the default options to calculate a wMNE solution with constrained dipole orientation.
Other subjects
- sub001: Bad registration
- sub006: Lots of blinks
- sub016: Lots of SQUID jumps
Scripting
Corresponding script in the Brainstorm distribution:
TODO: Coming soon.