|
Size: 8192
Comment:
|
Size: 11592
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| ## page was renamed from brainsuiteBDP ## page was renamed from brainsuite |
'''[TUTORIAL UNDER CONSTRUCTION: NOT READY FOR PUBLIC USE]''' ---- |
| Line 6: | Line 8: |
| '''[TUTORIAL UNDER WRITING: NOT READY FOR PUBLIC USE]''' | In this tutorial, we describe the estimation of realistic conductivity tensors of living brain tissues using the [[http://brainsuite.org/|BrainSuite software]]. These results are used in FEM forward modeling, as described in the tutorials: [[https://neuroimage.usc.edu/brainstorm/Tutorials/Duneuro#DUNEuro_options:_Advanced|FEM with DUNEuro]] and [[https://neuroimage.usc.edu/brainstorm/Tutorials/FemMedianNerve#FEM_tensors|FEM median nerve example]]. |
| Line 8: | Line 10: |
| Describe brainsuite here : here we will describe the process of the brain tissues anisotrpy estimation and the different functions that brainstorm offers. | The realistic tensors are estimated from the Diffusion-Weighted Images (DWI): Brainstorm calls the BrainSuite software to compute the diffusion tensors on each brain MRI voxel (DTI), then Effective Medium Approach (EMA) is applied to estimate the conductivity tensors for each element of a tetrahedral FEM mesh. This is particularly interesting for the modeling the anisotropy of the white matter. |
| Line 10: | Line 12: |
| This tutorial explains how to use Brainsuite to estimate the anisotropy of the brain tissues. refer to this page [[https://neuroimage.usc.edu/brainstorm/Tutorials/SegBrainSuite?highlight=(Anand)|https://neuroimage.usc.edu/brainstorm/Tutorials/SegBrainSuite?highlight=%28anand%29]] The realistic tensors are estimated from the Diffusion-Weighted Images (DWI). For this purpose, Brainstorm calls internally the BrainSuite Diffusion Pipeline to compute the diffusion tensors on each brain voxel. Afterward, the Effective Medium Approach is applied to convert the diffusion tensors to the conductivity tensors. The following section shows the users how to do it from the graphical interface. Only the NIfTI is supported. All the diffusion data, including the DWI file and direction and the value of the gradient files, respectively the *.nii, the *.bval and the *.bvec are required. Ideally, these files should have the same name and saved in the same folder. |
BrainSuite is also used for other purposes in Brainstorm, particularly the T1 MRI segmentation, as documented in this tutorial: [[Tutorials/SegBrainSuite|MRI segmentation: BrainSuite]]. |
| Line 22: | Line 16: |
| == Requirement == * You have already followed all the introduction tutorials * You have a working copy of Brainstorm installed on your computer |
== Download and installation == ==== Requirements ==== * You have already followed all the introduction tutorials. * You have a working copy of Brainstorm installed on your computer. * For the DWI data, only the NIfTI files (.nii) are supported. |
| Line 26: | Line 22: |
| == Brainsuite Installation == 1. Download the latest version of BrainSuite from http://www.brainsuite.org/download. |
==== Install Brainsuite ==== 1. Download the latest version of BrainSuite from http://forums.brainsuite.org/download/. |
| Line 29: | Line 26: |
| 1. Note that you will be using BrainSuite Diffusion Pipeline(BDP), so you need to install a compatible [[http://www.mathworks.com/products/compiler/mcr|MATLAB Compiler Runtime]](last version). 1. Start BrainSuite to check if the installation (It's not required to open BrainSuite to run this tutorial). 1. The BrainSuite installation folder should be informed in the Brainstorm preferences |
|
| Line 33: | Line 27: |
| 1. You will be using BrainSuite Diffusion Pipeline (BDP), so you need to install a compatible [[https://www.mathworks.com/products/compiler/matlab-runtime.html|MATLAB Runtime]] (2019b for BrainSuite 21a). | |
| Line 34: | Line 29: |
| ---- /!\ '''Edit conflict - other version:''' ---- {{attachment:brainsuiteInstalation.JPG||height="250",width="650"}} |
1. In Brainstorm, menu File > Edit preferences > Enter the BrainSuite installation folder:<<BR>><<BR>> {{attachment:brainsuiteInstall.gif}} |
| Line 37: | Line 31: |
| ==== Download the dataset ==== * Download the files: [[http://brainsuite.org/WebTutorialData/BrainSuiteTutorialSVReg_Sept16.zip|MRI T1w]] and [[http://brainsuite.org/WebTutorialData/DWI_Feb15.zip|MRI DWI]] (from the [[http://brainsuite.org/tutorials/dtiexercise/|BrainSuite diffusion tutorial]]). |
|
| Line 38: | Line 34: |
| ---- /!\ '''Edit conflict - your version:''' ---- {{attachment:brainsuiteInstalation.JPG||height="250",width="650"}} |
* Unzip it outside of any of the Brainstorm folders (program folder or database folder). * Start Brainstorm (Matlab scripts or stand-alone version) * Select the menu File > Create new protocol. Name it "'''TutorialTensors'''" and select: * No, use individual anatomy * No, use one channel file per condition |
| Line 41: | Line 40: |
| == Import the anatomy == === T1 MRI === * Switch to the "anatomical data" view, the left button in the toolbar above the database explorer. * Right-click on the TutorialFem folder > New subject > '''Subject01''' * Keep the default options you set for the protocol. |
|
| Line 42: | Line 46: |
| ---- /!\ '''End of edit conflict''' ---- | * Right-click on the subject node > '''Import MRI''': * Set the file format: '''All MRI files (subject space)''' |
| Line 44: | Line 49: |
| == Dataset == In this tutorial, we use the Brainsuite dataset example available on the Brainsuite tutorial webpage |
* Select the T1 file: BrainSuiteTutorialSVReg/'''2523412.nii.gz''' |
| Line 47: | Line 51: |
| http://brainsuite.org/tutorials/. The T1w of the subject can be download from this link [[http://brainsuite.org/WebTutorialData/BrainSuiteTutorialSVReg_Sept16.zip|BrainSuiteTutorialSVReg_Sept16.zip]]. The DWI from this link [[http://brainsuite.org/WebTutorialData/DWI_Feb15.zip|DWI_Feb15.zip]] | * Click on the link "'''Click here to compute MNI normalization'''": option "'''maff8'''". This estimates an affine transformation to the [[https://neuroimage.usc.edu/brainstorm/CoordinateSystems#MNI_coordinates|MNI space]] and sets default positions for the anatomical fiducials. The NAS/LPA/RPA fiducials are needed for defining the Brainstorm [[CoordinateSystems|subject coordinate system]], in which the surfaces and FEM meshes are stored. <<BR>><<BR>> {{attachment:importT1.gif}} |
| Line 49: | Line 53: |
| The first link contains the T1 MRI, with the name '2523412.nii.gz'. as in this figure, | === Diffusion imaging === This computes the This requires BrainSuite to be installed on your computer, with the bdp program available in the system path. |
| Line 51: | Line 56: |
| {{attachment:vierMriFolder.JPG||height="130",width="580"}} | * Right-click on Subject01''' '''> '''Convert DWI to DTI''' |
| Line 53: | Line 58: |
| The second file is the DWI and should contain at least three files | * Select the DWI file: DWI/'''2523412.dwi.nii.gz''' |
| Line 55: | Line 60: |
| {{attachment:vierDWIFolder.JPG||height="150",width="580"}} | * The associated text files '''*.bvec''' (orientation of the gradient) and '''*.bval''' (value of the gradient) must be in the same folder, with the same file name. Theses files are created from for the DWI acquisition. If you don't have them, ask the person who programmed your DWI sequence and get the files that are specific to your use case. |
| Line 57: | Line 62: |
| Where the *.bval is a text file that contains the value of the gradient, and the *.bvec is also a text file that contains the orientation of the gradient. The nii.gz file is the NifTi file of the DWI where the images are stored. | * The process can take up to 30min. At the end, a new file '''DTI-EIG''' appears in the database (DTI=diffusion tensors images, EIG=eigenvalue). This file contains 12 volumes, ie. 12 values for each voxel. From 1 to 9: components of the three eigenvectors; from 10 to 12: the values of their norm to the eigenvalue. <<BR>><<BR>> {{attachment:importDTI.gif}} |
| Line 59: | Line 64: |
| == Realistic condctivity tensors == First, you need to create a new subject in your protocol, let call it the 'BrainSuiteSubject'. Then import the T1 MRI of the subject and set the fiducials points as explained in the previous tutorial. |
== FEM mesh == The FEM approach requires a segmentation of the head volume in different tissues, represented as hexahedral or tetrahedral 3D meshes. The methods available within Brainstorm are listed in the tutorial [[https://neuroimage.usc.edu/brainstorm/Tutorials/FemMesh|FEM mesh generation]]. Here we illustrate only the use of '''Brain2mesh''': this is not the most accurate solution for MRI segmentation but it is probably the fastest solution to obtain a tetrahedral mesh of the head with 5 tissues (gray matter, white matter, CSF, skull, skin). For more accurate results, we recommend using SimNIBS, as illustrated in the tutorial [[https://neuroimage.usc.edu/brainstorm/Tutorials/FemMedianNerve#FEM_mesh_with_SimNIBS|FEM median nerve example]]. |
| Line 62: | Line 67: |
| {{attachment:mri.JPG||height="100",width="250"}} === Diffusion tensor generation (DTI) from DWI === Right-click on the subject and then select the item "Convert DWI to DTI". ---- /!\ '''Edit conflict - other version:''' ---- Then follow the popup windows by selecting the DWI, bval, and bval. If these files are in the same folder, Brainstorm will detect them automatically, otherwise, the user will be asked to browse the files one by one (as it's the case in this tutorial). ---- /!\ '''Edit conflict - your version:''' ---- Then follow the popup windows by selecting the DWI, bval, and bval. If these files are in the same folder, Brainstorm will detect them automatically, otherwise, the user will be asked to browse the files one by one (as it's the case in this tutorial). ---- /!\ '''End of edit conflict''' ---- In this step, Brainstorm calls the Brainsuite internally, and the diffusion tensors are computed. At the end of this process, a new node will appear in the Brainstorm database with the name 'DTI-EIT'. This name refers to, DTI: diffusion tensors images, and EIG for eigenvalue, since the eigenvalues and eigenvectors are computed at voxel and stored in Brainstorm database. {{attachment:mriAndDTI-EIG.JPG||height="100",width="250"}} |
* Right-click on the T1 MRI > MRI segmentation > '''Generate FEM mesh''' > '''Brain2mesh'''. * After less than 15 minutes, you will obtain a new FEM mesh in the database. <<BR>><<BR>>{{attachment:femMesh.gif}} |
| Line 81: | Line 71: |
| The Effective Medium Approach is applied to convert the diffusion tensors to the conductivity tensors. This step requires the FEM mesh of the head model. You can generate the FEM head model from the MRI as explained on this page (add link). | The Effective Medium Approach is applied to convert the diffusion tensors to the conductivity tensors. |
| Line 83: | Line 73: |
| ---- /!\ '''Edit conflict - other version:''' ---- | www.pnas.org/content/98/20/11697 |
| Line 85: | Line 75: |
| For the following, we used the SimNibs FEM mesh generation. The following figure shows the FEM mesh obtained from using the T1 MRI. Note that this mesh is obtained only from the T1, the use of the T2 is highly recommended if it's available. | ==== FEM mesh head model ==== {{attachment:Mri&femMeshView.JPG|Mri&femMeshView.JPG|width="260",height="300"}} {{attachment:femMeshView.JPG||width="280",height="300"}} |
| Line 87: | Line 78: |
| ---- /!\ '''Edit conflict - your version:''' ---- | Note that this mesh is obtained only from the T1, the use of the T2 is highly recommended if it's available, as recommended in the [[https://neuroimage.usc.edu/brainstorm/Tutorials/FemMesh|FEM mesh tutorial]]. |
| Line 89: | Line 80: |
| For the following, we used the SimNibs FEM mesh generation. The following figure shows the FEM mesh obtained from using the T1 MRI. Note that this mesh is obtained only from the T1, the use of the T2 is highly recommended if it's available. | ==== Computation of FEM mesh tensors ==== Once the FEM mesh and the DTI tensors are available in the Brainstorm database, the next step for the FEM tensors can be performed by the following: |
| Line 91: | Line 83: |
| ---- /!\ '''End of edit conflict''' ---- | - Right-click on the FEM mesh - Compute FEM tensors |
| Line 93: | Line 85: |
| The FEM head model to use for tensors should be selected and highlighted with the green color (double click on the FEM mesh node to select it) | {{attachment:menuGenerateFemTensors.png||width="250",height="280"}} |
| Line 95: | Line 87: |
| When this is done, then right-click on the subject > Convert DWI to DTI, | Brainstorm checks the available tissues in the FEM head model and displays the following panel |
| Line 97: | Line 89: |
| {{attachment:FEMConductivitiesIsoPanel.JPG||width="250",height="220"}} | |
| Line 98: | Line 91: |
| ---- /!\ '''Edit conflict - other version:''' ---- Brainstorm will load the available tissue in the FEM head model and the following windows appear. |
This panel lists the tissues available in the FEM head model and assigns a default value of the conductivity for each compartment. Users can change these values to their own if needed. |
| Line 101: | Line 93: |
| Select the WM anisotropy and kee all the other tissues as isotropic. | DTI values can be used to generate conductivity tensors for the white matter (and in some cases for the grey matter). Please, note that the DWI can be used only for the brain tissues and not for the outers compartments (skull and skin) |
| Line 103: | Line 95: |
| The process of conversion from DWI to Conductivity tensors uses the EMA, furthermore, brainstorm proposes the option to use the adaptative EMA with the volume constraint option [ref]. In this example, we select the EMA with the VC. | In this tutorial (and in most cases) we select the white matter. Select the WM anisotropy and keep all the other tissues as isotropic, then these additional options appear asking for the method to use. |
| Line 105: | Line 97: |
| The process will take around 10 min, and then the FEM tensors are computed and stored in the FEM structure. [explain how it is organized and how to use it outside brainstorm ] | {{attachment:FEMConductivitiesAnisoPanel.JPG||width="250",height="300"}} |
| Line 107: | Line 99: |
| ---- /!\ '''Edit conflict - your version:''' ---- Brainstorm will load the available tissue in the FEM head model and the following windows appear. |
The available methods are: |
| Line 110: | Line 101: |
| Select the WM anisotropy and kee all the other tissues as isotropic. | - Effective Medium approach (EMA) |
| Line 112: | Line 103: |
| The process of conversion from DWI to Conductivity tensors uses the EMA, furthermore, brainstorm proposes the option to use the adaptative EMA with the volume constraint option [ref]. In this example, we select the EMA with the VC. | - Effective Medium approach with volume constraints (EMA + VC) |
| Line 114: | Line 105: |
| The process will take around 10 min, and then the FEM tensors are computed and stored in the FEM structure. [explain how it is organized and how to use it outside brainstorm ] | - Simulated or the artificial anisotropy |
| Line 116: | Line 107: |
| ---- /!\ '''End of edit conflict''' ---- | Only the two first methods require the DTI. More information about these methods can be found on these references [ref1][ref2] and in our main paper [link] |
| Line 118: | Line 109: |
| === Display the tensors === == Artificial/simulated conductivity tensors == |
In this tutorial, we use the method "EMA + VC", where the final tensors are constrained to fits the volume of the equivalent isotropic tensor volume. |
| Line 121: | Line 111: |
| ---- /!\ '''Edit conflict - other version:''' ---- In the case where the DWI is not available, or the users desire to evaluate the effect of the conductivity change on the model, the artificial conductivity can be used. |
==== Visulation of FEM mesh tensors ==== Once the FEM tensors are successfully computed, they are stored in the FEM head node. By right-clicking on the FEM head, new menu items are added that gives the possibilities to display the FEM tensors either as ellipsoids or as vectors in the direction of the main eigenvector. |
| Line 124: | Line 114: |
| ---- /!\ '''Edit conflict - your version:''' ---- In the case where the DWI is not available, or the users desire to evaluate the effect of the conductivity change on the model, the artificial conductivity can be used. |
{{attachment:menuDisplayTensors.jpg||width="250",height="300"}} |
| Line 127: | Line 116: |
| ---- /!\ '''End of edit conflict''' ---- | The tensors can be displayed either on the FEM mesh or overlaid on the MRI. The following figures show an example of the obtained tensors displayed on the white matter. |
| Line 129: | Line 118: |
| Two approaches are integrated within Brainstorm. Either the | {{attachment:meshViewTensorsLines.JPG||width="350",height="300"}} {{attachment:meshViewTensorsTensorsTops.JPG||width="350",height="300"}} On the left, the tensors as a line on the direction of the main eigenvector. On the right, the tensors displayed as ellipsoids. The orientation of the tensor is color-coded as follows: red for right-left, green for anterior-posterior, and blue for superior-inferior. Note that the quality of the tensors depends on the DWI data and the number of acquisition direction. Users can also display the tensors on specific tissues, for example on the white matter (left figure) or overlay on the MRI (right figure). {{attachment:meshViewTensorsLinesWM2.JPG||width="350",height="300"}} {{attachment:tensorsOnMri.JPG||width="300",height="300"}} ==== Recommendation ==== In the case where the user wants to use generate isotropic tensors, then the DTI is not required. For that case, keep all the options to 'isotropic', the recommended display is the 'Ellipsoids', and the final shape will be a sphere (isotropic direction). In the case where more than one FEM head model is in the database, the highlighted one in the green color will be used. <<TAG(Advanced)>> == Simulated conductivity tensor == In the case where the DWI is not available, or in the case where the users desire to evaluate the effect of the conductivity change on the head model, the artificial conductivity can be used. Users can reach this option by following this tutorial and select the third method in this panel. {{attachment:artificialTensors.JPG||width="280",height="450"}} Two approaches are integrated within Brainstorm. Either Wang's constraint or the volume's constraint (Wolters). The common feature between these methods is the ratio between the transversal and longitudinal conductivity ratio. A common example is the skull anisotropy simulation, where the longitudinal conductivity can be higher than the transversal conductivity, the ratio can vary from 2 to 10 [ref]. In this tutorial, we keep all the tissue as isotropic, except the skull, we use a ratio of 0.1 and select the volume constraint. The following figures show the results of this example. "eigenvalues parallel (longitudinal) and perpendicular (transverse) to the fiber directions" for 1:10 anisotropy (transverse:longitudinal) {{attachment:skullAniso.JPG||width="300",height="250"}} {{attachment:skullAniso2.JPG||width="300",height="250"}} == Troubleshooting == To be completed soon and linked to BrainSuite website |
| Line 132: | Line 150: |
| ===TODO=== Check the error in the simnibs mesh in X direction and overlay on mri check the error with the brain2mesh Correct the ratio from integer to float check the meaning of transversal/longitidunal in the code add an interactive way yo change the size of the tensor.. important correct the name of the simulated method, correct the EMC and remove the VC and change the coefficcient |
[TUTORIAL UNDER CONSTRUCTION: NOT READY FOR PUBLIC USE]
FEM tensors estimation with BrainSuite
Authors: Takfarinas Medani, Francois Tadel, Anand Joshi and Richard Leahy
In this tutorial, we describe the estimation of realistic conductivity tensors of living brain tissues using the BrainSuite software. These results are used in FEM forward modeling, as described in the tutorials: FEM with DUNEuro and FEM median nerve example.
The realistic tensors are estimated from the Diffusion-Weighted Images (DWI): Brainstorm calls the BrainSuite software to compute the diffusion tensors on each brain MRI voxel (DTI), then Effective Medium Approach (EMA) is applied to estimate the conductivity tensors for each element of a tetrahedral FEM mesh. This is particularly interesting for the modeling the anisotropy of the white matter.
BrainSuite is also used for other purposes in Brainstorm, particularly the T1 MRI segmentation, as documented in this tutorial: MRI segmentation: BrainSuite.
Contents
Download and installation
Requirements
- You have already followed all the introduction tutorials.
- You have a working copy of Brainstorm installed on your computer.
- For the DWI data, only the NIfTI files (.nii) are supported.
Install Brainsuite
Download the latest version of BrainSuite from http://forums.brainsuite.org/download/.
Install it on your computer by following the instructions in BrainSuite's quick start installation guide.
You will be using BrainSuite Diffusion Pipeline (BDP), so you need to install a compatible MATLAB Runtime (2019b for BrainSuite 21a).
In Brainstorm, menu File > Edit preferences > Enter the BrainSuite installation folder:
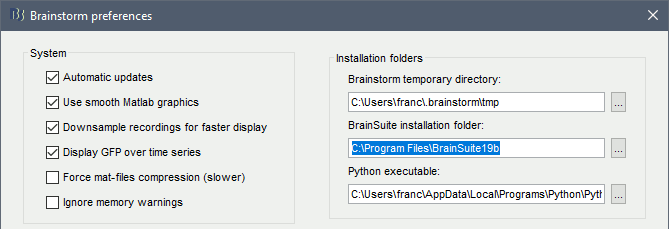
Download the dataset
Download the files: MRI T1w and MRI DWI (from the BrainSuite diffusion tutorial).
- Unzip it outside of any of the Brainstorm folders (program folder or database folder).
- Start Brainstorm (Matlab scripts or stand-alone version)
Select the menu File > Create new protocol. Name it "TutorialTensors" and select:
- No, use individual anatomy
- No, use one channel file per condition
Import the anatomy
T1 MRI
- Switch to the "anatomical data" view, the left button in the toolbar above the database explorer.
Right-click on the TutorialFem folder > New subject > Subject01
- Keep the default options you set for the protocol.
Right-click on the subject node > Import MRI:
Set the file format: All MRI files (subject space)
Select the T1 file: BrainSuiteTutorialSVReg/2523412.nii.gz
Click on the link "Click here to compute MNI normalization": option "maff8". This estimates an affine transformation to the MNI space and sets default positions for the anatomical fiducials. The NAS/LPA/RPA fiducials are needed for defining the Brainstorm subject coordinate system, in which the surfaces and FEM meshes are stored.
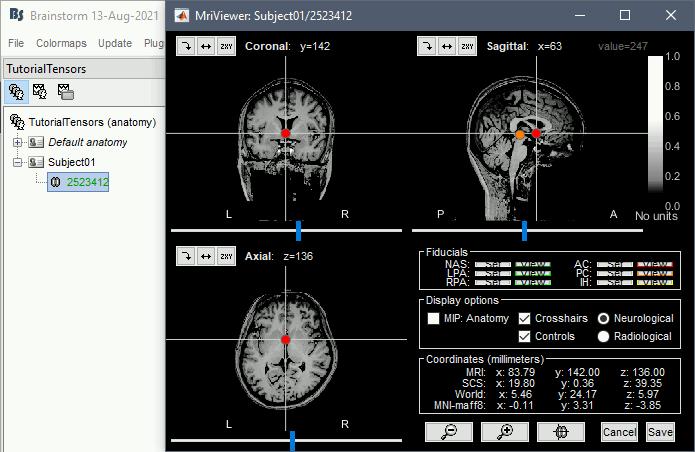
Diffusion imaging
This computes the This requires BrainSuite to be installed on your computer, with the bdp program available in the system path.
Right-click on Subject01 > Convert DWI to DTI
Select the DWI file: DWI/2523412.dwi.nii.gz
The associated text files *.bvec (orientation of the gradient) and *.bval (value of the gradient) must be in the same folder, with the same file name. Theses files are created from for the DWI acquisition. If you don't have them, ask the person who programmed your DWI sequence and get the files that are specific to your use case.
The process can take up to 30min. At the end, a new file DTI-EIG appears in the database (DTI=diffusion tensors images, EIG=eigenvalue). This file contains 12 volumes, ie. 12 values for each voxel. From 1 to 9: components of the three eigenvectors; from 10 to 12: the values of their norm to the eigenvalue.
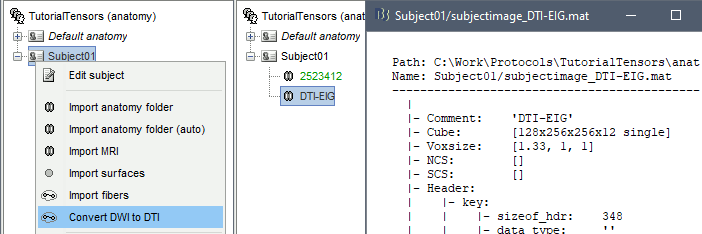
FEM mesh
The FEM approach requires a segmentation of the head volume in different tissues, represented as hexahedral or tetrahedral 3D meshes. The methods available within Brainstorm are listed in the tutorial FEM mesh generation. Here we illustrate only the use of Brain2mesh: this is not the most accurate solution for MRI segmentation but it is probably the fastest solution to obtain a tetrahedral mesh of the head with 5 tissues (gray matter, white matter, CSF, skull, skin). For more accurate results, we recommend using SimNIBS, as illustrated in the tutorial FEM median nerve example.
Right-click on the T1 MRI > MRI segmentation > Generate FEM mesh > Brain2mesh.
After less than 15 minutes, you will obtain a new FEM mesh in the database.
![[ATTACH] [ATTACH]](/moin_static198/brainstorm1/img/attach.png)
Conductivity tensor generation from DTI
The Effective Medium Approach is applied to convert the diffusion tensors to the conductivity tensors.
www.pnas.org/content/98/20/11697
FEM mesh head model
Note that this mesh is obtained only from the T1, the use of the T2 is highly recommended if it's available, as recommended in the FEM mesh tutorial.
Computation of FEM mesh tensors
Once the FEM mesh and the DTI tensors are available in the Brainstorm database, the next step for the FEM tensors can be performed by the following:
- Right-click on the FEM mesh - Compute FEM tensors
Brainstorm checks the available tissues in the FEM head model and displays the following panel
This panel lists the tissues available in the FEM head model and assigns a default value of the conductivity for each compartment. Users can change these values to their own if needed.
DTI values can be used to generate conductivity tensors for the white matter (and in some cases for the grey matter). Please, note that the DWI can be used only for the brain tissues and not for the outers compartments (skull and skin)
In this tutorial (and in most cases) we select the white matter. Select the WM anisotropy and keep all the other tissues as isotropic, then these additional options appear asking for the method to use.
The available methods are:
- Effective Medium approach (EMA)
- Effective Medium approach with volume constraints (EMA + VC)
- Simulated or the artificial anisotropy
Only the two first methods require the DTI. More information about these methods can be found on these references [ref1][ref2] and in our main paper [link]
In this tutorial, we use the method "EMA + VC", where the final tensors are constrained to fits the volume of the equivalent isotropic tensor volume.
Visulation of FEM mesh tensors
Once the FEM tensors are successfully computed, they are stored in the FEM head node. By right-clicking on the FEM head, new menu items are added that gives the possibilities to display the FEM tensors either as ellipsoids or as vectors in the direction of the main eigenvector.
The tensors can be displayed either on the FEM mesh or overlaid on the MRI. The following figures show an example of the obtained tensors displayed on the white matter.
On the left, the tensors as a line on the direction of the main eigenvector. On the right, the tensors displayed as ellipsoids. The orientation of the tensor is color-coded as follows: red for right-left, green for anterior-posterior, and blue for superior-inferior.
Note that the quality of the tensors depends on the DWI data and the number of acquisition direction.
Users can also display the tensors on specific tissues, for example on the white matter (left figure) or overlay on the MRI (right figure).
Recommendation
In the case where the user wants to use generate isotropic tensors, then the DTI is not required. For that case, keep all the options to 'isotropic', the recommended display is the 'Ellipsoids', and the final shape will be a sphere (isotropic direction). In the case where more than one FEM head model is in the database, the highlighted one in the green color will be used.
Simulated conductivity tensor
In the case where the DWI is not available, or in the case where the users desire to evaluate the effect of the conductivity change on the head model, the artificial conductivity can be used.
Users can reach this option by following this tutorial and select the third method in this panel. ![]()
Two approaches are integrated within Brainstorm. Either Wang's constraint or the volume's constraint (Wolters). The common feature between these methods is the ratio between the transversal and longitudinal conductivity ratio.
A common example is the skull anisotropy simulation, where the longitudinal conductivity can be higher than the transversal conductivity, the ratio can vary from 2 to 10 [ref]. In this tutorial, we keep all the tissue as isotropic, except the skull, we use a ratio of 0.1 and select the volume constraint. The following figures show the results of this example.
"eigenvalues parallel (longitudinal) and perpendicular (transverse) to the fiber directions" for 1:10 anisotropy (transverse:longitudinal)
Troubleshooting
To be completed soon and linked to BrainSuite website
References
===TODO===
Check the error in the simnibs mesh in X direction and overlay on mri check the error with the brain2mesh Correct the ratio from integer to float check the meaning of transversal/longitidunal in the code add an interactive way yo change the size of the tensor.. important correct the name of the simulated method, correct the EMC and remove the VC and change the coefficcient
