|
Size: 17823
Comment:
|
Size: 29757
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 4: | Line 4: |
| The open-source software FreeSurfer can be used to extract the cortical envelope from a T1 MRI. It also registers the individual cortex surfaces to surface-based anatomical atlases (Desikan-Killiany, Destrieux, Brodmann). The process is fully automatic and the results can be imported in Brainstorm with just a few mouse clicks. '''Important note''': Whether you are using FreeSurfer for the T1 segmentation, the cortical atlases or the FSAverage subject default: please register on their website ([[https://surfer.nmr.mgh.harvard.edu/registration.html|registration page]]) and cite the appropriate references. |
The open-source software FreeSurfer can be used to extract the cortical envelope from a T1/T2 MRI and register it to an atlas. The process is fully automatic and the results can be imported in Brainstorm with just a few mouse clicks. If you are using FreeSurfer, please '''register''' on their website ([[https://surfer.nmr.mgh.harvard.edu/registration.html|registration page]]) and '''cite '''the appropriate references. Note that this software is available only for '''Linux and MacOS'''. |
| Line 10: | Line 8: |
| == Running FreeSurfer == 1. Downloading and installing FreeSurfer is very easy. It just takes some time because the distribution package is huge (get ready to download several Gb). You have two options: you have a Linux system and you want to install FreeSurfer, or you don't and you want to run FreeSurfer in a Linux virtual machine.<<BR>>Just follow the instructions: http://surfer.nmr.mgh.harvard.edu/fswiki/DownloadAndInstall |
== Run FreeSurfer manually == 1. Downloading and installing FreeSurfer is easy but long (several Gb). Two options: you have a Linux system and you want to install FreeSurfer, or you don't and you want to run FreeSurfer in a Linux virtual machine. Just follow the instructions:<<BR>> http://surfer.nmr.mgh.harvard.edu/fswiki/DownloadAndInstall |
| Line 19: | Line 17: |
| 1. Set up the FreeSurfer environment in a '''bash''' environment (add those lines at the end of your $HOME/.bashrc script): | 1. Set up the FreeSurfer environment in a '''bash''' environment (add these lines at the end of your $HOME/.bashrc script): |
| Line 27: | Line 25: |
| 1. Run the reconstruction: | 1. Run the reconstruction (6-12 hours): |
| Line 32: | Line 30: |
| 1. Go home, come back the next day. The process is fully automatic, but quite resource-consuming. 1. Done. Everything is ready to be imported in Brainstorm. The results are usually good, but depending on the quality of the structural MR, it may fail. Because of this unpredictable behavior, you always need to check visually the final surfaces. The FreeSurfer wiki suggests that you check all the steps with FreeSurfer. We suggest instead that you load it all in Brainstorm and go back to the manual checking/editing only if it looks bad. 1. More detailed instructions for setting up the environment and tuning the reconstruction here:<<BR>>http://surfer.nmr.mgh.harvard.edu/fswiki/RecommendedReconstruction == Importing the results in Brainstorm == 1. Switch to the anatomy side of the database explorer 1. Create a new subject, set the default anatomy option to "No, use individual anatomy" 1. Right-click on the subject > Import anatomy folder...<<BR>><<BR>> {{attachment:import1.gif}} <<BR>><<BR>> 1. Select the file format "FreeSurfer folder" and select the top folder of your subject <subject_id> (/.../data/freesurfer/subjects/subject_id)<<BR>>To import the cortical thickness maps at the same time, you can select the format "FreeSurfer folder + Thickness maps" instead. 1. Then you're prompted for the number of vertices you want in the final cortex surface. This will by extension define the number of dipoles to estimate during the source estimation process. By default we set this value to 15000 for the entire brain (it means 7500 for each hemisphere).<<BR>><<BR>> {{attachment:nVertices.gif}} <<BR>><<BR>> 1. The MRI Viewer appears, and a help window asks you to validate the orientation of the MRI and to define the 6 fiducial points. If something doesn't look right at this step, for instance if the MRI is not presented with a correct orientation, you should stop this automatic import process and follow the manual instructions in the basic tutorial pages. <<BR>><<BR>> {{attachment:mriviewer.gif}} <<BR>><<BR>> 1. Place the six fiducials. If you need help, refer to this page: CoordinateSystems 1. Click on Save to keep your modifications, and the automatic import will go on. 1. The files that are imported from the subject_id folder are the following: * /mri/'''T1.mgz''' (T1 MRI volume) * /mri/'''aseg.mgz''' (segmentation of subcortical structures) * /surf/'''?h.pial''' (grey/csf interface) * /surf/'''?h.white '''(grey/white matter interface) * /surf/'''?h.sphere.reg''' (registered parametrized sphere, for subject co-registration) * /label/'''?h.*.annot''' (cortical surface-based atlases) * /surf/'''?h.thickness''' (cortical thickness map) 1. The successive steps that are performed automatically by Brainstorm: * Import all the surfaces (left/right, white/pial) * Load all the atlases available for each surface (note that the .pial and .white surfaces are matching point-to-point, so the same annotation files are imported for both surface types) * Load the registered spheres for the the left and right hemispheres * Downsample each hemisphere to the number specified in the options (by default 7500, half of the total default number 15000) * Merge left and right hemispheres for the two surface types: white matter and cortex envelope * Delete all the unnecessary surfaces * Generate a head surface from the MRI * Read the sub-cortical atlas aseg.mgz as a set of labelled surfaces * Read the cortical thickness maps 1. The files you can see in the database explorer at the end: <<BR>><<BR>> {{attachment:checkDb.gif}} <<BR>><<BR>> * '''MRI''': The T1 MRI of the subject, imported from the MGH file format (.mgz) * '''head mask''' (10000,0,2): Scalp surface generated by Brainstorm. The numbers indicate the parameters that were used automatically for this head: vertices=10000, erode factor=0, fill holes=2 (those are detailed later) * '''cortex_300000V''': High-resolution cortex surface that was generated by FreeSurfer, that contains usually between 200,000 and 300,000 vertices. * '''cortex_15000V''': Low-resolution cortex surface, downsampled using the reducepatch function from Matlab (it keeps a meaningful subset of vertices from the original surface). It appears in green in the database explorer, ie. it is going to be used as the default by the processes that require a cortex surface. * '''white_300000V''': High-resolution white matter envelope from FreeSurfer * '''white_15000V''': Low-resolution white matter, processed with reducepatch * '''aseg atlas''': Atlas of subcortical regions 1. A figure is automatically shown at the end of the process, to check visually that the low-resolution cortex and head surfaces were properly generated and imported. If it doesn't look like the following picture, do not go any further in your source analysis, fix the anatomy first.<<BR>><<BR>> {{attachment:checkResult.gif}} <<BR>><<BR>> |
1. '''Contralateral registration''': Computes the registration of each hemisphere to the contralateral atlas, which later allows projecting scouts between hemispheres, but adds about one hour of computation. [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Contralateral_registration|More information]]<<BR>><<BR>> '''FreeSurfer 6.x''': Add the option [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Contralateral_registration|-contrasurfreg]] to the recon-all command line. {{{ recon-all -all -contrasurfreg -subjid <subject_id> }}} '''FreeSurfer 7.x''': The option -contrasurfreg was removed for [[https://www.mail-archive.com/freesurfer@nmr.mgh.harvard.edu/msg73877.html|unknown reasons]]. Execute manually the following commands after recon-all:<<BR>> {{{ mris_register -curv -reverse $SUBJECTS_DIR/<subject_id>/surf/lh.sphere $FREESURFER_HOME/average/rh.folding.atlas.acfb40.noaparc.i12.2016-08-02.tif $SUBJECTS_DIR/<subject_id>/surf/lh.rh.sphere.reg mris_register -curv -reverse $SUBJECTS_DIR/<subject_id>/surf/rh.sphere $FREESURFER_HOME/average/lh.folding.atlas.acfb40.noaparc.i12.2016-08-02.tif $SUBJECTS_DIR/<subject_id>/surf/rh.lh.sphere.reg }}} 1. If you have an additional T2 or FLAIR image, you can improve the cortex reconstruction: https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all#UsingT2orFLAIRdatatoimprovepialsurfaces 1. Additional volume or surface parcellations can be computed at this stage. For details see section [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Cortical_parcellations|Cortical parcellations]] below. 1. Done. Everything is ready to be imported in Brainstorm. Because the process may occasionally fail, always check visually the final surfaces. The FreeSurfer wiki suggests that you check all the steps with FreeSurfer. We suggest instead that you load it all in Brainstorm and go back to the manual checking/editing only if it looks bad. 1. More instructions for setting up the environment and tuning the reconstruction here: http://surfer.nmr.mgh.harvard.edu/fswiki/RecommendedReconstruction == Run FreeSurfer from Brainstorm == Alternatively, it is possible to call the FreeSurfer recon-all pipeline from Brainstorm. * Switch to the anatomy side of the database explorer. * Create a new subject, set the default anatomy option to "No, use individual anatomy". * Import the T1 MRI for this subject. * Optional: Import the T2 MRI for this subject, if available. * Set the [[https://neuroimage.usc.edu/brainstorm/Tutorials/ImportAnatomy#Fiducial_points|fiducial points]] manually (NAS/LPA/RPA) or [[https://neuroimage.usc.edu/brainstorm/Tutorials/ImportAnatomy#MNI_normalization|compute the MNI normalization]]. * Select the T1 MRI (or the T1+T2 MRIs simultaneously). * Right-click > '''MRI segmentation > FreeSurfer'''. Two options can be selected interactively: the number of final vertices in the cortex surface, and the extra [[https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all|command-line options]] for recon-all.<<BR>><<BR>> {{attachment:run_fs_menu.gif}} * Alternatively, use the process: '''Import anatomy > Segment MRI with FreeSurfer'''. <<BR>><<BR>> {{attachment:run_fs_process.gif}} * This process saves the output of the recon-all pipeline in the regular the regular FreeSurfer subjects folder (typically in /usr/local/freesurfer/version/subjects). If a sub-folder with the subject name already exists in that folder, it would return an error unless you select the option "'''Delete FreeSurfer subject'''". * The option "'''Run asynchronously'''" starts the recon-all pipeline on the system in a separate process, but does not track its execution. Some instructions are displayed in the Matlab command window to help you follow or cancel the FreeSurfer computation. At the end, you need to call manually the menu '''Import anatomy folder''' described below. * The option '''-contrasurfreg '''allows the computation of the [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Contralateral_registration|contralateral registration]], which allows projecting scouts between hemispheres. But takes an extra hour of computation and is only available with '''FreeSurfer 6.x'''. If you want to skip this step, remove this option from the command-line options. If you are using '''FreeSurfer 7.x''', refer to the [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Run_FreeSurfer_manually|instructions above]]. == Import the results in Brainstorm == 1. Switch to the anatomy view of the database explore. 1. Create a new subject, set the default anatomy option to "No, use individual anatomy". 1. Right-click on the subject > Import anatomy folder.<<BR>><<BR>> {{attachment:import1.gif}} * '''Import anatomy folder''': Interactive import: asks for the number of vertices expected in the final cortex surfaces and for the location of the fiducials NAS/LPA/RPA. Select this option in the case of an MEG study, when you know exactly where the fiducials were digitized during the MEG acquisition. See tutorial [[https://neuroimage.usc.edu/brainstorm/Tutorials/ChannelFile#Automatic_registration|MRI registration]]. * '''Import anatomy folder (auto)''': Automatic import: Computes the [[https://neuroimage.usc.edu/brainstorm/CoordinateSystems#MNI_coordinates|linear MNI normalization]], uses default positions from the MNI atlas for the NAS/LPA/RPA fiducials, and uses 15000 vertices for the cortex downsampled surfaces. 1. Select one of the FreeSurfer import options and select the top folder of your subject <subject_id> (/.../data/freesurfer/subjects/subject_id) * '''FreeSurfer''': Import the T1 MRI, pial and white cortex surfaces, surface parcellations, surface spherical registration, ASEG surface parcellation. Reconstruct the head surface. * '''FreeSurfer + Volume atlas''': Same as above, with the following additions: Import the volume parcellations available in the mri subfolder; reconstruct the '''mid''' surfaces (intermediate between the pial and white surfaces). * '''FreeSurfer + Volume atlas + Thickness''': Same as above, with the cortical thickness maps imported as source maps. 1. Manual import: You're prompted for the number of vertices you want in the final cortex surface. This will by extension define the number of dipoles to estimate during the source estimation process. By default we set this value to 15000 for the entire brain (it means 7500 for each hemisphere).<<BR>><<BR>> {{attachment:nVertices.gif}} <<BR>><<BR>> 1. Manual import: The MRI Viewer appears for you to define the anatomical fiducials. See tutorial [[https://neuroimage.usc.edu/brainstorm/Tutorials/ImportAnatomy#Fiducial_points|Import anatomy]]. Click on Save to keep your modifications, and the import will continue. <<BR>><<BR>> {{attachment:mriviewer.gif}} 1. A figure is automatically shown at the end of the process, to check visually that the low-resolution cortex and head surfaces were properly generated and imported. If it doesn't look like the following picture, do not go any further in your source analysis, fix the anatomy first.<<BR>><<BR>> {{attachment:checkResult.gif}} {{{#!wiki note From July 2024, Brainstorm also supports importing the output folder from <<BR>>'''`recon-all-clinical.sh`'''. ⚠️ The '''`recon-all-clinical.sh`''' in !FreeSurfer 7.4 has a small bug. More information in:<<BR>> https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all-clinical }}} == Files imported == The files that are imported from the subject_id folder are the following: * /mri/'''T1.mgz''' (T1 MRI volume) + optional /mri/'''T2.mgz''' * /mri/'''*aseg.mgz''' (volume parcellations) * /surf/'''?h.pial''' (grey/csf interface) * /surf/'''?h.white '''(grey/white matter interface) * /surf/'''?h.sphere.reg''' (registered parametrized sphere, for subject co-registration) * /label/'''?h.*.annot''' (cortical surface-based atlases) * /surf/'''?h.thickness''' (cortical thickness map) The successive steps that are performed by Brainstorm: * Import the T1/T2 MRI * For "auto" import: Compute the linear MNI registration, otherwise asks for placing the fiducials * Import all the surfaces (left/right, white/pial) * Load all the atlases available for each surface (note that the .pial and .white surfaces are matching point-to-point, so the same annotation files are imported for both surface types) * Load the registered spheres for the left and right hemispheres * Downsample each hemisphere to the number specified in the options (by default 7500, half of the total default number 15000) * Compute the mid-surfaces (average of the pial + white surfaces vertex by vertex) * Merge left and right hemispheres for the two surface types: white matter and cortex envelope * Generate a head surface from the MRI * Import the sub-cortical atlas *aseg.mgz as volume atlases and labelled surfaces * Import the cortical thickness maps The files you can see in the database explorer at the end (with Volume atlases + Thickness): * '''MRI''': The T1 MRI of the subject, imported from the MGH file format (T1.mgz) * '''Volume parcellations''': *aseg.mgz, see tutorial [[https://neuroimage.usc.edu/brainstorm/Tutorials/ExploreAnatomy#Subcortical_regions:_Volume|Explore the anatomy]]. * '''head mask''': Scalp surface generated by Brainstorm. The numbers in the filename indicate the parameters that were used for this head: vertices=10000, erode factor=0, fill holes=2, background threshold=detected (these are detailed later) * '''cortex_300000V''': High-resolution pial surface that was generated by FreeSurfer, that contains usually between 200,000 and 300,000 vertices. * '''cortex_15000V''': Low-resolution pial surface, downsampled using the reducepatch function from Matlab (it keeps a meaningful subset of vertices from the original surface). It appears in green in the database explorer, ie. it is going to be used as the default by the processes that require a cortex surface. * '''white_300000V''': High-resolution white matter envelope from FreeSurfer * '''white_15000V''': Low-resolution white matter, processed with reducepatch * '''mid_*V''': Mid-cortex surfaces (average of the pial + white surfaces vertex by vertex) * '''subcortical''': Atlas of subcortical regions imported as labelled surfaces {{attachment:checkDb.gif}} |
| Line 74: | Line 124: |
| ==== How to check the quality of the result ==== It's hard to estimate what would be a good cortical reconstruction. What you are trying to spot at this level is mostly the obvious errors, like when the early stages of the brain extraction didn't perform well, just with a visual inspection. Play with the ''Smooth'' slider in the ''Surface ''tab. If it looks like a brain (two separate hemispheres) in both smooth and original views, it is probably ok. |
=== How to check the quality of the result === It's hard to estimate what would be a good cortical reconstruction. Try to spot the most obvious errors, e.g. when the early stages of the brain extraction didn't perform well. Play with the ''Smooth'' slider in the ''Surface ''tab. If it looks like a brain (two separate hemispheres) in both smooth and original views, it is probably OK. |
| Line 81: | Line 131: |
| ==== The cortex looks bad ==== | === The cortex looks bad === |
| Line 84: | Line 134: |
| If after following those instructions you still don't manage to get good surfaces, you can try to run the automatic MRI segmentation from [[Tutorials/SegBrainVisa|BrainVISA]] or [[Tutorials/SegBrainSuite|BrainSuite]]. ==== The head surface looks bad ==== It is not mandatory to have a perfect head surface to use any of the Brainstorm features: you don't necessarily have to recognize the face (for the anonymity of the figures, it can be even better if you don't). The head surface is important mostly for the alignment of the MEG sensors and the MRI. If you digitized the head shape with a Polhemus device, you can align automatically the head surface (hence the MRI) with the MEG sensors (in the same referential as the Polhemus points). The quality of this automatic registrations depends on the quality of both surfaces: the Polhemus head shape (green points) and the head surface from the MRI (grey surface). If you placed lots of points on the nose but your head surface doesn't have a nose, those points are not going to help. Except for that, a nice head shape is mainly useful for producing nicer figures. {{attachment:checkAlignMeg.gif||height="243",width="298"}} If the default head surface looks bad, you can try generating another one: right-click on the subject folder > Generate head surface. The options are: |
If after following these instructions you still don't manage to get good surfaces, you can try to run the automatic MRI segmentation from [[https://neuroimage.usc.edu/brainstorm/Tutorials/SegCAT12|CAT12]], [[Tutorials/SegBrainSuite|BrainSuite]] or [[Tutorials/SegBrainVisa|BrainVISA]]. === The head surface looks bad === It is not mandatory to have a perfect head surface to use many Brainstorm features. (For anonymity, it may actually be necessary to distort or obscure the face in figures.) The head surface is important mainly for two reasons. First, for the alignment of the MEG sensors and the MRI, and second, for generating head models that incorporate the scalp surface (e.g. multi-layer BEM or FEM). If you digitized the head shape with a Polhemus device, you can automatically align the head surface (hence the MRI) with the MEG sensors (in the same referential as the Polhemus points). The quality of this automatic registration depends on the quality of both surfaces: the Polhemus head shape (green points) and the head surface from the MRI (grey surface). If you digitized lots of points on the nose but your head surface doesn't have a nose, these points are not going to help. . {{attachment:checkAlignMeg.gif||width="298",height="243"}} If the default head surface looks bad, you can try generating another one: right-click on the subject folder > '''Generate head surface'''. The options are: |
| Line 97: | Line 147: |
| * '''Fill holes factor''': Number of dimensions in which the holes should be identified and closed. Increasing this number removes more of the cavities of the head surface (0=no correction, 1=removes holes inside the surface, 3=closes all the features that make the surface non-convex)<<BR>><<BR>> {{attachment:generateHead.gif}} | * '''Fill holes factor''': Number of dimensions in which the holes should be identified and closed. Increasing this number removes more of the cavities of the head surface (0=no correction, 1=removes holes inside the surface, 3=closes all the features that make the surface non-convex) * '''Background threshold''': Intensity below which most MRI voxels will be considered as background. This threshold is set automatically after analyzing the volume histogram, but this detection sometime fails. Redefining manually the background threshold may help obtaining a better head surface for noisy MRI scans. If the background was already removed and the background voxels set to zero, you should set this threshold to 1.<<BR>>To help you define this threshold, you can get the intensity value of the selected voxel in the top-right corner of the MRI Viewer. <<BR>><<BR>> {{attachment:generateHead.gif||width="165",height="280"}} {{attachment:mriValue.gif||width="425",height="214"}} === Error in mne_read_surface.m === If you get an error message mentioning mne_read_surface.m, and the files pial.lh and pial.rh are only 2Kb, you probably had an issue with the transfer of symbolic links. See these forum posts: * https://neuroimage.usc.edu/forums/t/error-in-mne-read-surface-m/26418 * https://neuroimage.usc.edu/forums/t/failure-to-load-freesurfer-mri-data/39590/2 |
| Line 100: | Line 157: |
| The default analysis pipeline in FreeSurfer implements an automatic parcellation of the cortical surface in anatomical regions. The description of this feature is available here:<<BR>>http://freesurfer.net/fswiki/CorticalParcellation With FreeSurfer 5.3, 4 atlases are available on all the individual brains: * Destrieux atlas (?h.aparc.a2009s.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/CorticalParcellation|more information]] * Desikan-Killiany atlas (?h.aparc.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/CorticalParcellation|more information]] * Mindboggle (?h.aparc.DKTatlas40.annot): [[http://mindboggle.info/data.html|more information]] * Brodman areas (?h.BA.annot and ?h.BA.thresh.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/BrodmannAreaMaps|more information]] Those atlases are imported in Brainstorm as scouts (cortical regions of interest), and saved directly in the surface files. To check where they are saved: right-click on the low-resolution cortex file > File > View .mat file. You can see that 4 structures "Atlas" are available, the first one that has Name='User scouts', and the second one Name='Destrieux'.<<BR>><<BR>> {{attachment:viewMat.gif}} <<BR>><<BR>> To access them from the interface: Double-click on the cortex and go to the ''Scout'' tab, and click on the drop-down list to select another ''Atlas ''(ie group of scouts): {{attachment:scoutTab.gif}} |
The default analysis pipeline in FreeSurfer computes several parcellations of the cortical surface in anatomical regions. The description of this feature is available here:<<BR>>http://freesurfer.net/fswiki/CorticalParcellation With FreeSurfer 7, the following parcellations are always available: * '''Brodmann''' areas (?h.BA*.annot and ?h.BA*.thresh.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/BrodmannAreaMaps|more information]] * '''Destrieux''' atlas (?h.aparc.a2009s.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/CorticalParcellation|more information]] * '''Desikan-Killiany''' atlas (?h.aparc.annot): [[http://ftp.nmr.mgh.harvard.edu/fswiki/CorticalParcellation|more information]] * '''DKT / Mindboggle6''' (?h.aparc.DKTatlas.annot): [[http://mindboggle.info/data.html|more information]] * '''VcAtlas''' (?h.mpm.vpnl.annot): [[http://vpnl.stanford.edu/vcAtlas/|more information]] Additional parcellations can be computed from FreeSurfer with [[https://freesurfer.net/fswiki/mri_surf2surf|mri_surf2surf]] ([[https://github.com/ftadel/IntrAnat/issues/7#issuecomment-768996270|example]]): * '''Brainnetome''' (?h.BN_Atlas.annot): [[https://www.nitrc.org/frs/shownotes.php?release_id=3459|more information]] | [[https://neuroimage.usc.edu/forums/t/brainnetome-atlas-in-default-anatomy/16849/6|forum]] * '''HCP-MMP1''' (?h.HCP-MMP1.annot): [[https://figshare.com/articles/dataset/HCP-MMP1_0_projected_on_fsaverage/3498446?file=5528837|more information]] * '''CHubs/OASIS''' (?h.oasis.chubs.annot): [[https://surfer.nmr.mgh.harvard.edu/fswiki/Chubs|more information]] * '''PALS-B12''' (?h.PALS_B12_*.annot): [[https://surfer.nmr.mgh.harvard.edu/fswiki/PALS_B12|more information]] * '''Schaefer2018''' (?h.Schaefer2018_*Parcels_*Networks_order.annot): [[https://github.com/ThomasYeoLab/CBIG/tree/master/stable_projects/brain_parcellation/Schaefer2018_LocalGlobal/Parcellations/project_to_individual|more information]] * '''Yeo2011''' (?h.Yeo2011_7Networks_N1000, ?h.Yeo2011_17Networks_N1000): [[https://surfer.nmr.mgh.harvard.edu/fswiki/CorticalParcellation_Yeo2011|more information]] * '''Lausanne''' '''2008''' (?h.myaparc_*.annot): [[https://github.com/jguillon/easy_lausanne|more information]] These parcellations are imported in Brainstorm as [[https://neuroimage.usc.edu/brainstorm/Tutorials/Scouts|scouts]] (cortical regions of interest). To access them from the interface: double-click on the cortex, go to the ''Scout'' tab, and click on the drop-down list to select another ''Atlas ''(ie group of scouts): {{attachment:scoutTab.png}} |
| Line 127: | Line 192: |
| The scouts are saved directly in the surface files. To check where they are saved: right-click on the low-resolution cortex file > File > View file contents. You can see that multiple "Atlas" structures are available, the first one that has Name='User scouts', and the second one Name='Brodmann'. {{attachment:viewMat.gif}} == Volume parcellations == All the volume parcellations available in the mri subfolder are imported when the option "FreeSurfer + Volumes atlases" is selected. This includes: * '''ASEG''': mri/aseg.mgz, subcortical structures only * '''Destrieux''': aparc.a2009s+aseg.mgz * '''Desikan-Killiany''': aparc+aseg.mgz * '''DKT''': aparc.DKTatlas+aseg.mgz {{attachment:volatlas.gif}} |
|
| Line 128: | Line 207: |
| The file aseg.mgz contains a volume atlas of 40 subcortical regions. Brainstorm reads those volume labels and tesselates some of those regions, groups all the meshes in a large surface file where the regions are identified in an atlas called "Structures". It identifies: 8 bialateral structures (accumbens, amygdala, caudate, hippocampus, pallidum, putamen, thalamus, cerebellum) and 1 central structure (brainstem). You can easily extract one structure (for example the brainstem or the cerebellum) by selecting the corresponding entries in the scouts list and selecting the menu Scout > Edit surface > Keep only selected scouts. It creates a new surface with only the selected regions. If you want to remove one or several structures, use the menu "Remove selected scouts" instead. |
The file aseg.mgz contains a volume atlas of 40 subcortical regions. Brainstorm reads these volume labels and tesselates some of these regions, groups all the meshes in a large surface file where the regions are identified in an atlas called "Structures". It identifies: 8 bialateral structures (accumbens, amygdala, caudate, hippocampus, pallidum, putamen, thalamus, cerebellum) and 1 central structure (brainstem). You can easily extract one structure (for example the brainstem or the cerebellum) by selecting the corresponding entries in the scouts list and selecting the menu Scout > Edit surface > Keep only selected scouts. It creates a new surface with only the selected regions. If you want to remove one or several structures, use the menu "Remove selected scouts" instead. [[https://neuroimage.usc.edu/brainstorm/Tutorials/DeepAtlas#Select_deep_structures|More information]]. |
| Line 136: | Line 215: |
| == Registered sphere == The registered sphere is saved in each surface file in the field Reg.Sphere.Vertices. There is nothing that can be done with this information at this point, but it will become helpful when projecting the source results from the individual brains to the default anatomy of the protocol, for a group analysis of the results: [[Tutorials/CoregisterSubjects|Subject coregistration]]. |
== Registered spheres == The registered spheres are saved in each surface file in the field Reg.Sphere.Vertices. There is nothing that can be done with this information at this point, but it will become helpful when projecting the source results from the individual brains to the default anatomy of the protocol, for a group analysis of the results: [[Tutorials/CoregisterSubjects|Subject coregistration]]. |
| Line 140: | Line 219: |
== Contralateral registration == When computing the contralateral registration (see instructions [[https://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Run_FreeSurfer_manually|above]]), two additional files are generated in the FreeSurfer subject folder: surf/lh.rh.sphere.reg and surf/rh.lh.sphere.reg. These files define [[https://neuroimage.usc.edu/brainstorm/Tutorials/CoregisterSubjects#Using_FreeSurfer_registered_spheres|FreeSurfer spheres]] that register each hemisphere to the contralateral atlas, and therefore allow interpolating information from one hemisphere to the other. This is needed for projecting scouts between hemispheres, as done by the Scout tab menu [[https://neuroimage.usc.edu/brainstorm/Tutorials/Scouts#Menu:_Scout|Project to: Contralateral hemisphere]]. When these extra files are available in the FreeSurfer subject folder, they are imported automatically in the cortex surface (field Reg.SphereLR). But note that this option is not part of the standard recon-all pipeline and the contralateral spheres must be computed [[http://neuroimage.usc.edu/brainstorm/Tutorials/LabelFreeSurfer#Run_FreeSurfer_manually|manually]]. Read more about the FreeSurfer 6.x option -contrasurfreg on the software wiki:<<BR>>https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all#ContralateralSurfaceRegistation.28SphericalMorph.29.28-.3Cno.3Econtasurfreg.29 When the contralateral registration is not available for a subject, it is still possible to project scouts between hemispheres using the anatomy template. If the template available in the ''(Default anatomy)'' folder of the protocol includes the contralateral registration (e.g. ICBM152_2022). The scouts are projected to the template, then from one hemisphere to the other of the template, and finally back to the subject space. This procedure is less accurate than the direct projection using the subject-level contralateral projection. {{attachment:project_contra.gif}} == Mollweide projection == The FreeSurfer registered spheres can be used to compute a flat 2D projection of the cortex surface using the Mollweide projection, as described in this article: [[https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3445499/|(Kang et al. 2019) Hemispherically-Unified Surface Maps of Human Cerebral Cortex: Reliability and Hemispheric Asymmetries]]. * Right-click on a cortex surface > '''MRI registration > 2D projection (Mollweide)''': <<BR>><<BR>> {{attachment:mollweide_scouts.gif||width="627",height="226"}} * Right-click on a source file > '''Cortical activations > 2D projection (Mollweide)''': <<BR>><<BR>> {{attachment:mollweide_sources.gif||width="626",height="291"}} |
|
| Line 147: | Line 243: |
| Instead of the MNI Colin27 brain, you can use the FreeSurfer average subject "FSAverage" as your default anatomy in Brainstorm. This template is an average of 40 subjects using a spherical averaging described in [[http://nmr.mgh.harvard.edu/~fischl/reprints/morphing_human_brain_mapping_reprint.pdf|(Fischl et al. 1999)]]. To change the default, right-click on "(Default anatomy)" > Use template > FSAverage. If it is not available on your computer yet, it will be downloaded automatically from the server to your user folder: $HOME/.brainstorm/templates/anatomy. |
Instead of the MNI ICBM152 brain, you can use the FreeSurfer average subject "FSAverage" as your default anatomy in Brainstorm. This template is an average of 40 subjects using a spherical averaging described in [[http://nmr.mgh.harvard.edu/~fischl/reprints/morphing_human_brain_mapping_reprint.pdf|(Fischl et al. 1999)]]. To change the default, right-click on "(Default anatomy)" > Use template > FSAverage. If it is not available on your computer yet, it will be automatically downloaded from the server to your user folder: $HOME/.brainstorm/templates/anatomy. See tutorial: [[https://neuroimage.usc.edu/brainstorm/Tutorials/DefaultAnatomy#Subject_configuration|Using the anatomy templates]]. |
| Line 156: | Line 252: |
| You can import the FreeSurfer folders from scripts, but you have to provide manually the position for all the fiducial points: process Import anatomy > Import anatomy folder. | You can import the FreeSurfer folders from scripts, but you have to provide manually the position for all the fiducial points: process Import > Import anatomy > Import anatomy folder. |
| Line 166: | Line 262: |
| 1. Right-click on the subject folder > Import MRI > Select "mri/T1.mgz" 1. Set the 6 fiducial points, save 1. Right-click on the subject folder > Import surfaces > Select the FreeSurfer file format > Select simultaneously from the "surf" folder: lh.pial, lh.white, rh.pial, rh.white 1. Double-click on lh.pial toi display it. In the scout tab: Atlas > Load atlas > select all the lh.*.annot files available in the label folder. Close the figure. |
1. Right-click on the subject folder > Import MRI > Select "mri/T1.mgz". 1. Set the 6 fiducial points, or click on "Compute MNI normalization" (option maff8). Save. 1. Right-click on the subject folder > Import surfaces > Select the FreeSurfer file format > Select simultaneously from the "surf" folder: lh.pial, lh.white, rh.pial, rh.white. Question "Apply transformation?": NO. 1. Double-click on lh.pial to display it. In the scout tab: Atlas > Load atlas > select all the lh.*.annot files available in the label folder. Close the figure. |
| Line 174: | Line 270: |
| 1. Select lh.pial, rh.pial, right-click > Merge surfaces: Generates a surface cortex_250000V 1. Select lh.white, rh.white, right-click > Merge surfaces: Generates a surface white_250000V |
1. Select lh.pial + rh.pial, right-click > Merge surfaces: Generates a surface cortex_300000V 1. Select lh.white + rh.white, right-click > Merge surfaces: Generates a surface white_300000V |
| Line 180: | Line 276: |
| 1. Right-click on the subject folder > Import surfaces > Select the file format "Volume mask of atlas" > Select the file mri/aseg.mgz 1. Go to the functional view of the protocol, create a condition "FreeSurfer". Leave your mouse for a second over the new folder, an note the study index (iStudy). |
1. Right-click on the subject folder > Import surfaces > Select the file format "Volume mask or atlas (subject space)" > Select the file mri/aseg.mgz. Question "Apply transformation?": NO. 1. Right-click on the subject folder > Import MRI > Select the file format "Volume atlas (subject space)" > Select all the files mri/*aseg.mgz. * Question "Apply standard transformation?": NO * Question "Apply transformation?": YES * Question "How to register?": Ignore * Question "Reslice volume?": No 1. Go to the functional view of the protocol, create a folder "FreeSurfer". Leave your mouse for a second over the new folder, an note the study index (iStudy). |
| Line 188: | Line 289: |
| == Additional documentation == * Infant FreeSurfer: https://neuroimage.usc.edu/forums/t/incorrect-t1-orientation-when-loading-infant-freesurfer-output/32266 * https://neuroimage.usc.edu/forums/t/error-in-mne-read-surface-m/26418 * https://neuroimage.usc.edu/forums/t/failure-to-load-freesurfer-mri-data/39590/2 |
Using FreeSurfer
Authors: Francois Tadel
The open-source software FreeSurfer can be used to extract the cortical envelope from a T1/T2 MRI and register it to an atlas. The process is fully automatic and the results can be imported in Brainstorm with just a few mouse clicks. If you are using FreeSurfer, please register on their website (registration page) and cite the appropriate references. Note that this software is available only for Linux and MacOS.
Contents
- Run FreeSurfer manually
- Run FreeSurfer from Brainstorm
- Import the results in Brainstorm
- Files imported
- Handling errors
- Cortical parcellations
- Volume parcellations
- Subcortical structures: aseg atlas
- Registered spheres
- Contralateral registration
- Mollweide projection
- Cortical thickness
- FSAverage template
- Running the folder import as a process
- Manual import of the anatomy
- Additional documentation
Run FreeSurfer manually
Downloading and installing FreeSurfer is easy but long (several Gb). Two options: you have a Linux system and you want to install FreeSurfer, or you don't and you want to run FreeSurfer in a Linux virtual machine. Just follow the instructions:
http://surfer.nmr.mgh.harvard.edu/fswiki/DownloadAndInstallSet up the FreeSurfer in a csh/tcsh environment: run the following lines, or add them at the end of your $HOME/.cshrc script for permanent change. If you're not sure what csh is: type "echo $SHELL" to know what is the name of the shell that you use. If it says "/bin/tcsh" or "/bin/csh", this is for you.
setenv FREESURFER_HOME /.../local/freesurfer setenv SUBJECTS_DIR /.../data/freesurfer/subjects setenv FUNCTIONALS_DIR /.../data/freesurfer/sessions source /.../local/freesurfer/FreeSurferEnv.csh
Set up the FreeSurfer environment in a bash environment (add these lines at the end of your $HOME/.bashrc script):
export FREESURFER_HOME=/.../local/freesurfer export SUBJECTS_DIR=/.../data/ftadel/freesurfer export FUNCTIONALS_DIR=/.../data/ftadel/freesurfer source $FREESURFER_HOME/FreeSurferEnv.sh
- Run the reconstruction (6-12 hours):
recon-all -i <mri_file> -subjid <subject_id> recon-all -all -subjid <subject_id>
Contralateral registration: Computes the registration of each hemisphere to the contralateral atlas, which later allows projecting scouts between hemispheres, but adds about one hour of computation. More information
FreeSurfer 6.x: Add the option -contrasurfreg to the recon-all command line.recon-all -all -contrasurfreg -subjid <subject_id>
FreeSurfer 7.x: The option -contrasurfreg was removed for unknown reasons. Execute manually the following commands after recon-all:
mris_register -curv -reverse $SUBJECTS_DIR/<subject_id>/surf/lh.sphere $FREESURFER_HOME/average/rh.folding.atlas.acfb40.noaparc.i12.2016-08-02.tif $SUBJECTS_DIR/<subject_id>/surf/lh.rh.sphere.reg mris_register -curv -reverse $SUBJECTS_DIR/<subject_id>/surf/rh.sphere $FREESURFER_HOME/average/lh.folding.atlas.acfb40.noaparc.i12.2016-08-02.tif $SUBJECTS_DIR/<subject_id>/surf/rh.lh.sphere.reg
If you have an additional T2 or FLAIR image, you can improve the cortex reconstruction: https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all#UsingT2orFLAIRdatatoimprovepialsurfaces
Additional volume or surface parcellations can be computed at this stage. For details see section Cortical parcellations below.
Done. Everything is ready to be imported in Brainstorm. Because the process may occasionally fail, always check visually the final surfaces. The FreeSurfer wiki suggests that you check all the steps with FreeSurfer. We suggest instead that you load it all in Brainstorm and go back to the manual checking/editing only if it looks bad.
More instructions for setting up the environment and tuning the reconstruction here: http://surfer.nmr.mgh.harvard.edu/fswiki/RecommendedReconstruction
Run FreeSurfer from Brainstorm
Alternatively, it is possible to call the FreeSurfer recon-all pipeline from Brainstorm.
- Switch to the anatomy side of the database explorer.
- Create a new subject, set the default anatomy option to "No, use individual anatomy".
- Import the T1 MRI for this subject.
- Optional: Import the T2 MRI for this subject, if available.
Set the fiducial points manually (NAS/LPA/RPA) or compute the MNI normalization.
- Select the T1 MRI (or the T1+T2 MRIs simultaneously).
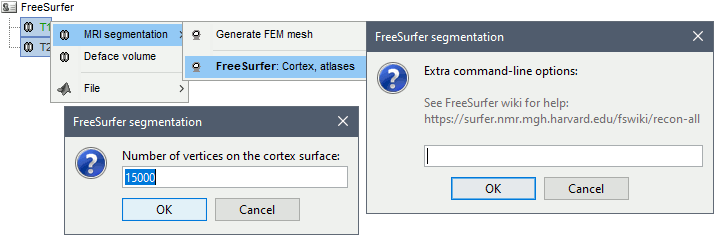
Right-click > MRI segmentation > FreeSurfer. Two options can be selected interactively: the number of final vertices in the cortex surface, and the extra command-line options for recon-all.
Alternatively, use the process: Import anatomy > Segment MRI with FreeSurfer.
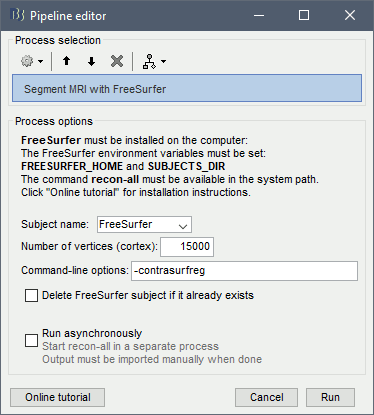
This process saves the output of the recon-all pipeline in the regular the regular FreeSurfer subjects folder (typically in /usr/local/freesurfer/version/subjects). If a sub-folder with the subject name already exists in that folder, it would return an error unless you select the option "Delete FreeSurfer subject".
The option "Run asynchronously" starts the recon-all pipeline on the system in a separate process, but does not track its execution. Some instructions are displayed in the Matlab command window to help you follow or cancel the FreeSurfer computation. At the end, you need to call manually the menu Import anatomy folder described below.
The option -contrasurfreg allows the computation of the contralateral registration, which allows projecting scouts between hemispheres. But takes an extra hour of computation and is only available with FreeSurfer 6.x. If you want to skip this step, remove this option from the command-line options. If you are using FreeSurfer 7.x, refer to the instructions above.
Import the results in Brainstorm
- Switch to the anatomy view of the database explore.
- Create a new subject, set the default anatomy option to "No, use individual anatomy".
Right-click on the subject > Import anatomy folder.
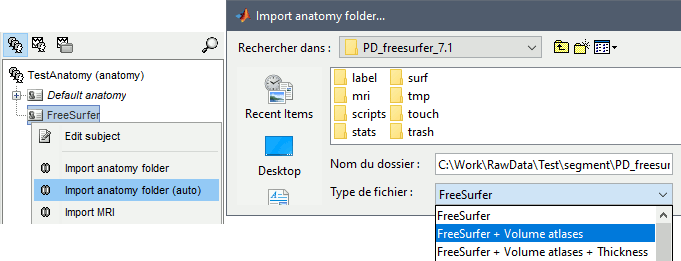
Import anatomy folder: Interactive import: asks for the number of vertices expected in the final cortex surfaces and for the location of the fiducials NAS/LPA/RPA. Select this option in the case of an MEG study, when you know exactly where the fiducials were digitized during the MEG acquisition. See tutorial MRI registration.
Import anatomy folder (auto): Automatic import: Computes the linear MNI normalization, uses default positions from the MNI atlas for the NAS/LPA/RPA fiducials, and uses 15000 vertices for the cortex downsampled surfaces.
Select one of the FreeSurfer import options and select the top folder of your subject <subject_id> (/.../data/freesurfer/subjects/subject_id)
FreeSurfer: Import the T1 MRI, pial and white cortex surfaces, surface parcellations, surface spherical registration, ASEG surface parcellation. Reconstruct the head surface.
FreeSurfer + Volume atlas: Same as above, with the following additions: Import the volume parcellations available in the mri subfolder; reconstruct the mid surfaces (intermediate between the pial and white surfaces).
FreeSurfer + Volume atlas + Thickness: Same as above, with the cortical thickness maps imported as source maps.
Manual import: You're prompted for the number of vertices you want in the final cortex surface. This will by extension define the number of dipoles to estimate during the source estimation process. By default we set this value to 15000 for the entire brain (it means 7500 for each hemisphere).
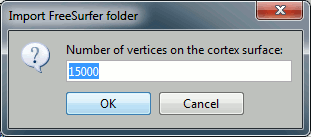
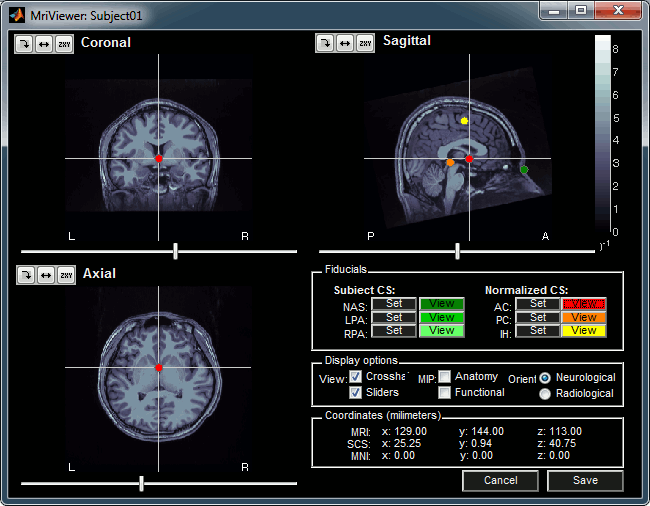
Manual import: The MRI Viewer appears for you to define the anatomical fiducials. See tutorial Import anatomy. Click on Save to keep your modifications, and the import will continue.
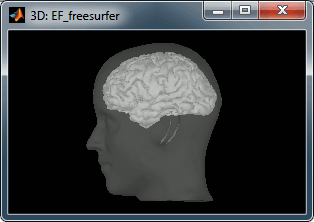
A figure is automatically shown at the end of the process, to check visually that the low-resolution cortex and head surfaces were properly generated and imported. If it doesn't look like the following picture, do not go any further in your source analysis, fix the anatomy first.
From July 2024, Brainstorm also supports importing the output folder from
recon-all-clinical.sh.
⚠️ The recon-all-clinical.sh in FreeSurfer 7.4 has a small bug. More information in:
https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all-clinical
Files imported
The files that are imported from the subject_id folder are the following:
/mri/T1.mgz (T1 MRI volume) + optional /mri/T2.mgz
/mri/*aseg.mgz (volume parcellations)
/surf/?h.pial (grey/csf interface)
/surf/?h.white (grey/white matter interface)
/surf/?h.sphere.reg (registered parametrized sphere, for subject co-registration)
/label/?h.*.annot (cortical surface-based atlases)
/surf/?h.thickness (cortical thickness map)
The successive steps that are performed by Brainstorm:
- Import the T1/T2 MRI
- For "auto" import: Compute the linear MNI registration, otherwise asks for placing the fiducials
- Import all the surfaces (left/right, white/pial)
- Load all the atlases available for each surface (note that the .pial and .white surfaces are matching point-to-point, so the same annotation files are imported for both surface types)
- Load the registered spheres for the left and right hemispheres
- Downsample each hemisphere to the number specified in the options (by default 7500, half of the total default number 15000)
- Compute the mid-surfaces (average of the pial + white surfaces vertex by vertex)
- Merge left and right hemispheres for the two surface types: white matter and cortex envelope
- Generate a head surface from the MRI
- Import the sub-cortical atlas *aseg.mgz as volume atlases and labelled surfaces
- Import the cortical thickness maps
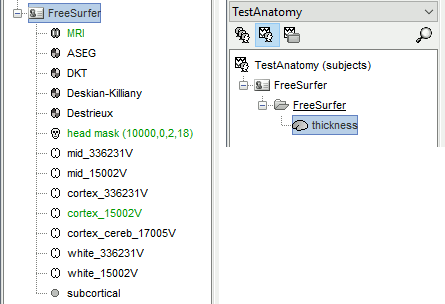
The files you can see in the database explorer at the end (with Volume atlases + Thickness):
MRI: The T1 MRI of the subject, imported from the MGH file format (T1.mgz)
Volume parcellations: *aseg.mgz, see tutorial Explore the anatomy.
head mask: Scalp surface generated by Brainstorm. The numbers in the filename indicate the parameters that were used for this head: vertices=10000, erode factor=0, fill holes=2, background threshold=detected (these are detailed later)
cortex_300000V: High-resolution pial surface that was generated by FreeSurfer, that contains usually between 200,000 and 300,000 vertices.
cortex_15000V: Low-resolution pial surface, downsampled using the reducepatch function from Matlab (it keeps a meaningful subset of vertices from the original surface). It appears in green in the database explorer, ie. it is going to be used as the default by the processes that require a cortex surface.
white_300000V: High-resolution white matter envelope from FreeSurfer
white_15000V: Low-resolution white matter, processed with reducepatch
mid_*V: Mid-cortex surfaces (average of the pial + white surfaces vertex by vertex)
subcortical: Atlas of subcortical regions imported as labelled surfaces
Handling errors
How to check the quality of the result
It's hard to estimate what would be a good cortical reconstruction. Try to spot the most obvious errors, e.g. when the early stages of the brain extraction didn't perform well. Play with the Smooth slider in the Surface tab. If it looks like a brain (two separate hemispheres) in both smooth and original views, it is probably OK.
Display the cortex surface on top of the MRI slices, to make sure that they are well aligned, that the surface follows well the folds, and that left and right were not flipped: right-click on the low-resolution cortex > MRI registration > Check MRI/surface registration...
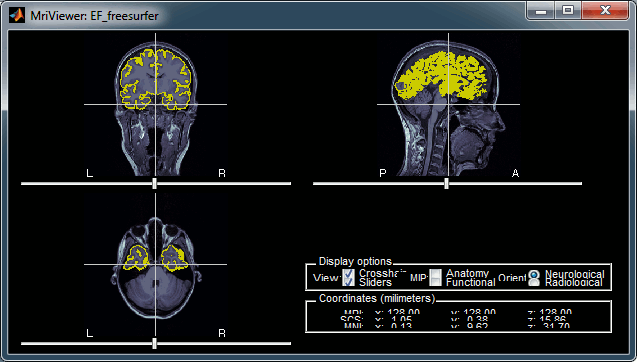
The cortex looks bad
It is critical to get a good cortex surface for source estimation. If the final cortex surface looks bad, it means that something didn't work well somewhere along the FreeSurfer pipeline. You can refer to the following page to fix the problems manually:
http://surfer.nmr.mgh.harvard.edu/fswiki/RecommendedReconstruction
If after following these instructions you still don't manage to get good surfaces, you can try to run the automatic MRI segmentation from CAT12, BrainSuite or BrainVISA.
The head surface looks bad
It is not mandatory to have a perfect head surface to use many Brainstorm features. (For anonymity, it may actually be necessary to distort or obscure the face in figures.)
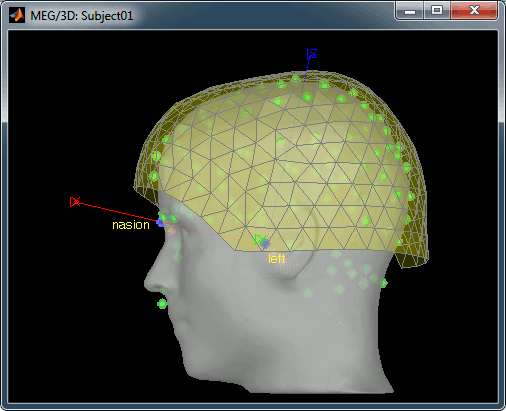
The head surface is important mainly for two reasons. First, for the alignment of the MEG sensors and the MRI, and second, for generating head models that incorporate the scalp surface (e.g. multi-layer BEM or FEM). If you digitized the head shape with a Polhemus device, you can automatically align the head surface (hence the MRI) with the MEG sensors (in the same referential as the Polhemus points). The quality of this automatic registration depends on the quality of both surfaces: the Polhemus head shape (green points) and the head surface from the MRI (grey surface). If you digitized lots of points on the nose but your head surface doesn't have a nose, these points are not going to help.
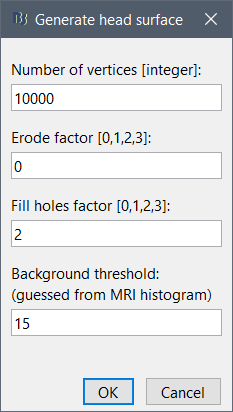
If the default head surface looks bad, you can try generating another one: right-click on the subject folder > Generate head surface. The options are:
Number of vertices: Number of points that are kept from the initial isosurface computed from the MRI. Increasing this number may increase the quality of the final surface.
Erode factor: Number of pixels to erode after the first binary threshold of the MRI. Increasing this number removes small components that are connected to the head.
Fill holes factor: Number of dimensions in which the holes should be identified and closed. Increasing this number removes more of the cavities of the head surface (0=no correction, 1=removes holes inside the surface, 3=closes all the features that make the surface non-convex)
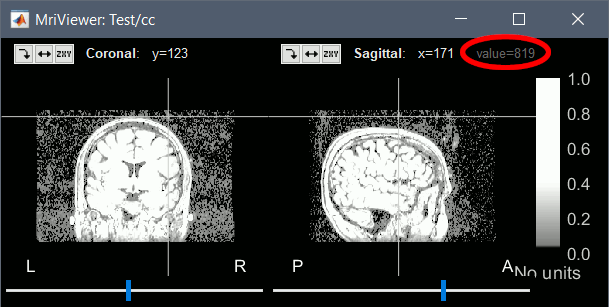
Background threshold: Intensity below which most MRI voxels will be considered as background. This threshold is set automatically after analyzing the volume histogram, but this detection sometime fails. Redefining manually the background threshold may help obtaining a better head surface for noisy MRI scans. If the background was already removed and the background voxels set to zero, you should set this threshold to 1.
To help you define this threshold, you can get the intensity value of the selected voxel in the top-right corner of the MRI Viewer.
Error in mne_read_surface.m
If you get an error message mentioning mne_read_surface.m, and the files pial.lh and pial.rh are only 2Kb, you probably had an issue with the transfer of symbolic links. See these forum posts:
https://neuroimage.usc.edu/forums/t/error-in-mne-read-surface-m/26418
https://neuroimage.usc.edu/forums/t/failure-to-load-freesurfer-mri-data/39590/2
Cortical parcellations
The default analysis pipeline in FreeSurfer computes several parcellations of the cortical surface in anatomical regions. The description of this feature is available here:
http://freesurfer.net/fswiki/CorticalParcellation
With FreeSurfer 7, the following parcellations are always available:
Brodmann areas (?h.BA*.annot and ?h.BA*.thresh.annot): more information
Destrieux atlas (?h.aparc.a2009s.annot): more information
Desikan-Killiany atlas (?h.aparc.annot): more information
DKT / Mindboggle6 (?h.aparc.DKTatlas.annot): more information
VcAtlas (?h.mpm.vpnl.annot): more information
Additional parcellations can be computed from FreeSurfer with mri_surf2surf (example):
Brainnetome (?h.BN_Atlas.annot): more information | forum
HCP-MMP1 (?h.HCP-MMP1.annot): more information
CHubs/OASIS (?h.oasis.chubs.annot): more information
PALS-B12 (?h.PALS_B12_*.annot): more information
Schaefer2018 (?h.Schaefer2018_*Parcels_*Networks_order.annot): more information
Yeo2011 (?h.Yeo2011_7Networks_N1000, ?h.Yeo2011_17Networks_N1000): more information
Lausanne 2008 (?h.myaparc_*.annot): more information
These parcellations are imported in Brainstorm as scouts (cortical regions of interest). To access them from the interface: double-click on the cortex, go to the Scout tab, and click on the drop-down list to select another Atlas (ie group of scouts):
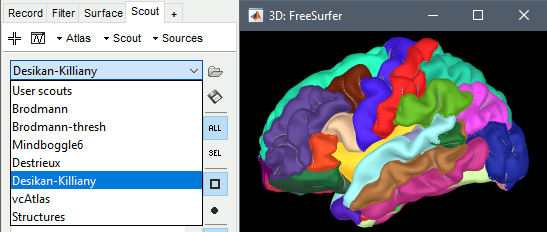
Desikan-Killiany atlas
Displayed respectively in: FreeSurfer, Brainstorm (high-resolution) and Brainstorm (15000 vertices)
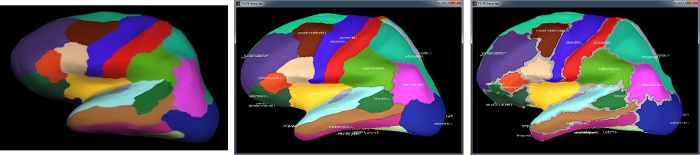
Destrieux atlas
Displayed in Brainstorm with the original scouts colors (left) or classified in 6 regions (right): pre-frontal, frontal, central, parietal, temporal, occipital, occipital. You can switch between the two views with the button "Identify regions with colors" in the toolbar on the right of the scouts list.
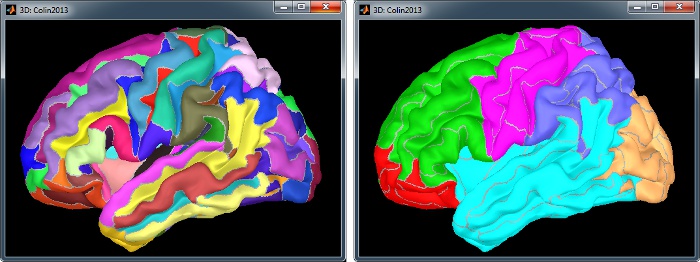
The scouts are saved directly in the surface files. To check where they are saved: right-click on the low-resolution cortex file > File > View file contents. You can see that multiple "Atlas" structures are available, the first one that has Name='User scouts', and the second one Name='Brodmann'.
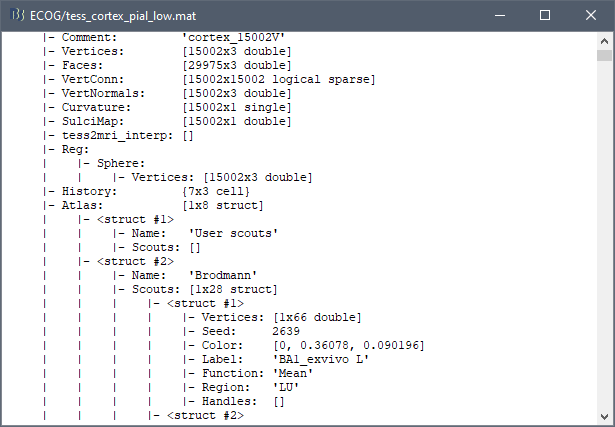
Volume parcellations
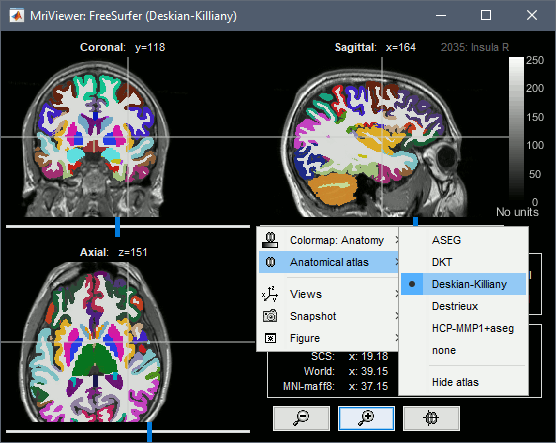
All the volume parcellations available in the mri subfolder are imported when the option "FreeSurfer + Volumes atlases" is selected. This includes:
ASEG: mri/aseg.mgz, subcortical structures only
Destrieux: aparc.a2009s+aseg.mgz
Desikan-Killiany: aparc+aseg.mgz
DKT: aparc.DKTatlas+aseg.mgz
Subcortical structures: aseg atlas
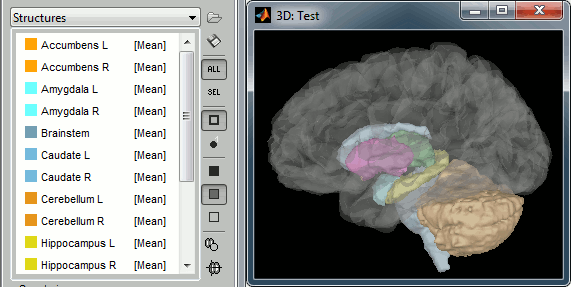
The file aseg.mgz contains a volume atlas of 40 subcortical regions. Brainstorm reads these volume labels and tesselates some of these regions, groups all the meshes in a large surface file where the regions are identified in an atlas called "Structures". It identifies: 8 bialateral structures (accumbens, amygdala, caudate, hippocampus, pallidum, putamen, thalamus, cerebellum) and 1 central structure (brainstem).
You can easily extract one structure (for example the brainstem or the cerebellum) by selecting the corresponding entries in the scouts list and selecting the menu Scout > Edit surface > Keep only selected scouts. It creates a new surface with only the selected regions. If you want to remove one or several structures, use the menu "Remove selected scouts" instead. More information.
Read more about the FreeSurfer subcortical atlas on the software wiki:
http://ftp.nmr.mgh.harvard.edu/fswiki/SubcorticalSegmentation
Registered spheres
The registered spheres are saved in each surface file in the field Reg.Sphere.Vertices. There is nothing that can be done with this information at this point, but it will become helpful when projecting the source results from the individual brains to the default anatomy of the protocol, for a group analysis of the results: Subject coregistration.
Read more about the FreeSurfer registration process on the software wiki:
https://surfer.nmr.mgh.harvard.edu/fswiki/SurfaceRegAndTemplates
Contralateral registration
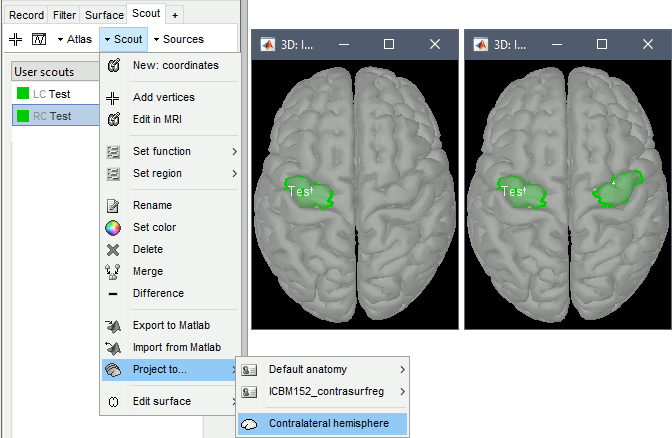
When computing the contralateral registration (see instructions above), two additional files are generated in the FreeSurfer subject folder: surf/lh.rh.sphere.reg and surf/rh.lh.sphere.reg. These files define FreeSurfer spheres that register each hemisphere to the contralateral atlas, and therefore allow interpolating information from one hemisphere to the other. This is needed for projecting scouts between hemispheres, as done by the Scout tab menu Project to: Contralateral hemisphere.
When these extra files are available in the FreeSurfer subject folder, they are imported automatically in the cortex surface (field Reg.SphereLR). But note that this option is not part of the standard recon-all pipeline and the contralateral spheres must be computed manually.
Read more about the FreeSurfer 6.x option -contrasurfreg on the software wiki:
https://surfer.nmr.mgh.harvard.edu/fswiki/recon-all#ContralateralSurfaceRegistation.28SphericalMorph.29.28-.3Cno.3Econtasurfreg.29
When the contralateral registration is not available for a subject, it is still possible to project scouts between hemispheres using the anatomy template. If the template available in the (Default anatomy) folder of the protocol includes the contralateral registration (e.g. ICBM152_2022). The scouts are projected to the template, then from one hemisphere to the other of the template, and finally back to the subject space. This procedure is less accurate than the direct projection using the subject-level contralateral projection.
Mollweide projection
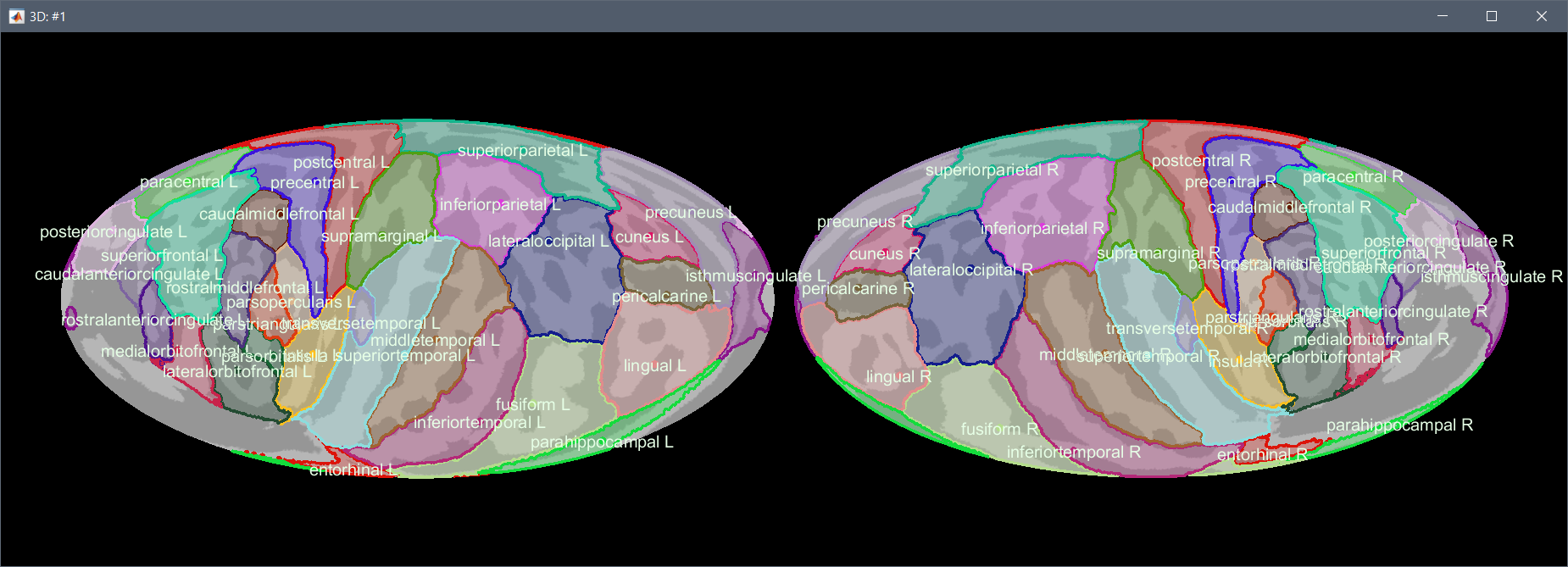
The FreeSurfer registered spheres can be used to compute a flat 2D projection of the cortex surface using the Mollweide projection, as described in this article: (Kang et al. 2019) Hemispherically-Unified Surface Maps of Human Cerebral Cortex: Reliability and Hemispheric Asymmetries.
Right-click on a cortex surface > MRI registration > 2D projection (Mollweide):
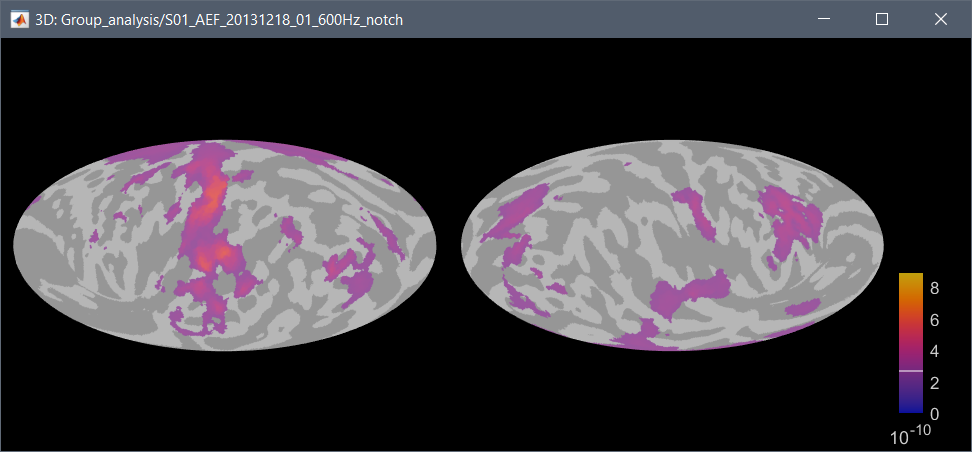
Right-click on a source file > Cortical activations > 2D projection (Mollweide):
Cortical thickness
The cortical thickness can be saved as a cortical map in the database (a "results" file). This result is generated when using the file format "FreeSurfer folder + Thickness maps" in the Import anatomy folder selection.
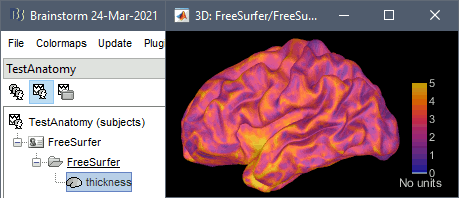
FSAverage template
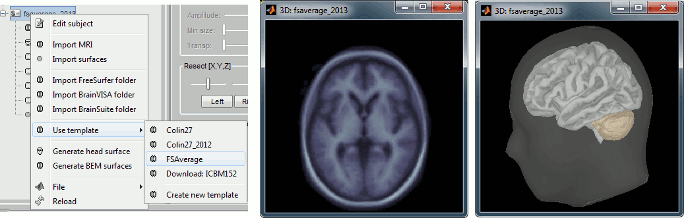
Instead of the MNI ICBM152 brain, you can use the FreeSurfer average subject "FSAverage" as your default anatomy in Brainstorm. This template is an average of 40 subjects using a spherical averaging described in (Fischl et al. 1999).
To change the default, right-click on "(Default anatomy)" > Use template > FSAverage. If it is not available on your computer yet, it will be automatically downloaded from the server to your user folder: $HOME/.brainstorm/templates/anatomy. See tutorial: Using the anatomy templates.
If you are using the FSAverage template but not a regular user of FreeSurfer, please register on their website: registration page.
Running the folder import as a process
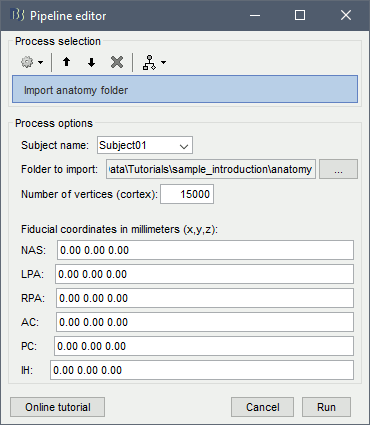
You can import the FreeSurfer folders from scripts, but you have to provide manually the position for all the fiducial points: process Import > Import anatomy > Import anatomy folder.
The corresponding process function is: brainstorm3/toolbox/process/functions/process_import_anatomy.m
Manual import of the anatomy
In case you need to import the MRI, surfaces and atlases separately instead of using the menu "Import anatomy folder", here is the sequence of operations to perform to get to the same result:
From the Anatomy side of the database explorer: create a subject.
Right-click on the subject folder > Import MRI > Select "mri/T1.mgz".
- Set the 6 fiducial points, or click on "Compute MNI normalization" (option maff8). Save.
Right-click on the subject folder > Import surfaces > Select the FreeSurfer file format > Select simultaneously from the "surf" folder: lh.pial, lh.white, rh.pial, rh.white. Question "Apply transformation?": NO.
Double-click on lh.pial to display it. In the scout tab: Atlas > Load atlas > select all the lh.*.annot files available in the label folder. Close the figure.
- Repeat for the other surfaces: lh.white, rh.pial, rh.white
Right-click on lh.pial > MRI registration > Load FreeSurfer sphere > Select "surf/lh.sphere.reg"
- Repeat with the other surfaces (use rh.sphere.reg for the right hemisphere, white and pial surfaces)
Select all the surfaces, right-click > Less vertices > 7500 vertices > Select the first option "Matlab reducepatch"
Select lh.pial + rh.pial, right-click > Merge surfaces: Generates a surface cortex_300000V
Select lh.white + rh.white, right-click > Merge surfaces: Generates a surface white_300000V
Select lh.pial_7500V, rh.pial_7500V, right-click > Merge surfaces: Generates a surface cortex_15000V
Select lh.white_7500V, rh.white_7500V, right-click > Merge surfaces: Generates a surface white_15000V
- Delete all the separate hemispheres: ?h.pial, ?h.white
- Double-click on cortex_15000V to set it as the default cortex
Right-click on the subject folder > Import surfaces > Select the file format "Volume mask or atlas (subject space)" > Select the file mri/aseg.mgz. Question "Apply transformation?": NO.
Right-click on the subject folder > Import MRI > Select the file format "Volume atlas (subject space)" > Select all the files mri/*aseg.mgz.
- Question "Apply standard transformation?": NO
- Question "Apply transformation?": YES
- Question "How to register?": Ignore
- Question "Reslice volume?": No
Go to the functional view of the protocol, create a folder "FreeSurfer". Leave your mouse for a second over the new folder, an note the study index (iStudy).
- From the Matlab command window, you can import the thickness maps with the following call:
ThickFile = import_sources(iStudy, CortexHiFile, ThickLhFile, ThickRhFile, 'FS');
CortexHiFile = full path to the high-res cortex file (right-click > File > Copy file path to clipboard)
ThickLhFile = full path to the surf/lh.thickness file
ThickRhFile = full path to the surf/rh.thickness file
Additional documentation
Infant FreeSurfer: https://neuroimage.usc.edu/forums/t/incorrect-t1-orientation-when-loading-infant-freesurfer-output/32266
https://neuroimage.usc.edu/forums/t/error-in-mne-read-surface-m/26418
https://neuroimage.usc.edu/forums/t/failure-to-load-freesurfer-mri-data/39590/2