What's new
Brainstorm is in a very active development state: small or major bug fixes and improvements are issued almost everyday. To update your version of the software easily: Install and update.
See also the full list of updates: brainstorm3/doc/updates.txt | All GitHub commits
April 2026
Anatomy
Bugfix: Install SPM tissue probability maps (TPM.nii) if not present in the system 🔗 🔗
Improve primitive cylinder mesh generation 🔗
Input / output
Nihon Koden files, improve reading start of acquisition timestamp 🔗
Update reading montages from FieldTrip data files for versions > ft-20250829 🔗
IO: Add support for polygon file format .ply mesh files 🔗
Bugfix: allow importing raw recordings for subject without MRI volume but with CT volume 🔗 🔗
Containers
Visualization
Timeseries, improve display of tick values on relative time 🔗
NIRS, allow display of volume head model 🔗

March 2026
Digitization
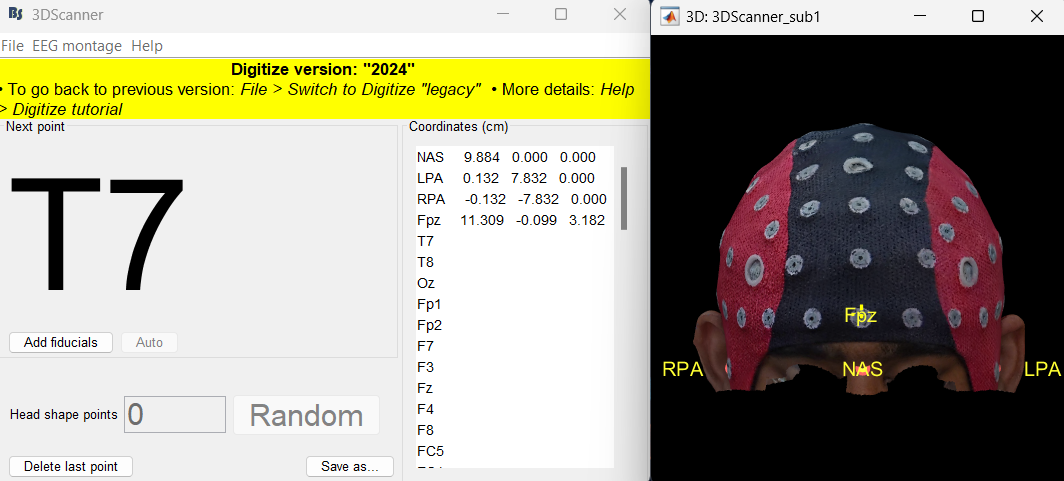
Auto-detect EEG locations for all EEG caps and custom montages 🔗

Events
Add GUI entry and process to merge overlapping extended events 🔗
Input / Output
Update and set timestamp for 0s in raw and imported recording files with GUI (right-click > "Set datetime for time = 0s") or with the process File > "Set datetime for time = 0s" 🔗
Bugfix: Apply montage on raw files with prop.times(1) ~= 0 🔗
Bugfix: Exporting recordings as EDF for raw files with prop.times(1) ~= 0 🔗
Bugfix: Rounding error on computing SamplesBounds for InRaw/OutRaw 🔗
Plugins
Simulations
Allow to simulate recordings from sources obtained with MEM 🔗
February 2026
Anatomy
Improve visualization of multiple structures and surfaces 🔗

Add non-interactive generation of primitive surfaces 🔗
Update parcellation labels for different BrainSuite atlases ("BrainSuiteAtlas1", "BCI-DNI_brain_atlas", and "USCBrain") 🔗 🔗
Head model
Add various directions for leadfield sensitivity visualization 🔗

Bst exlusion zone options interactive 🔗

IEEG
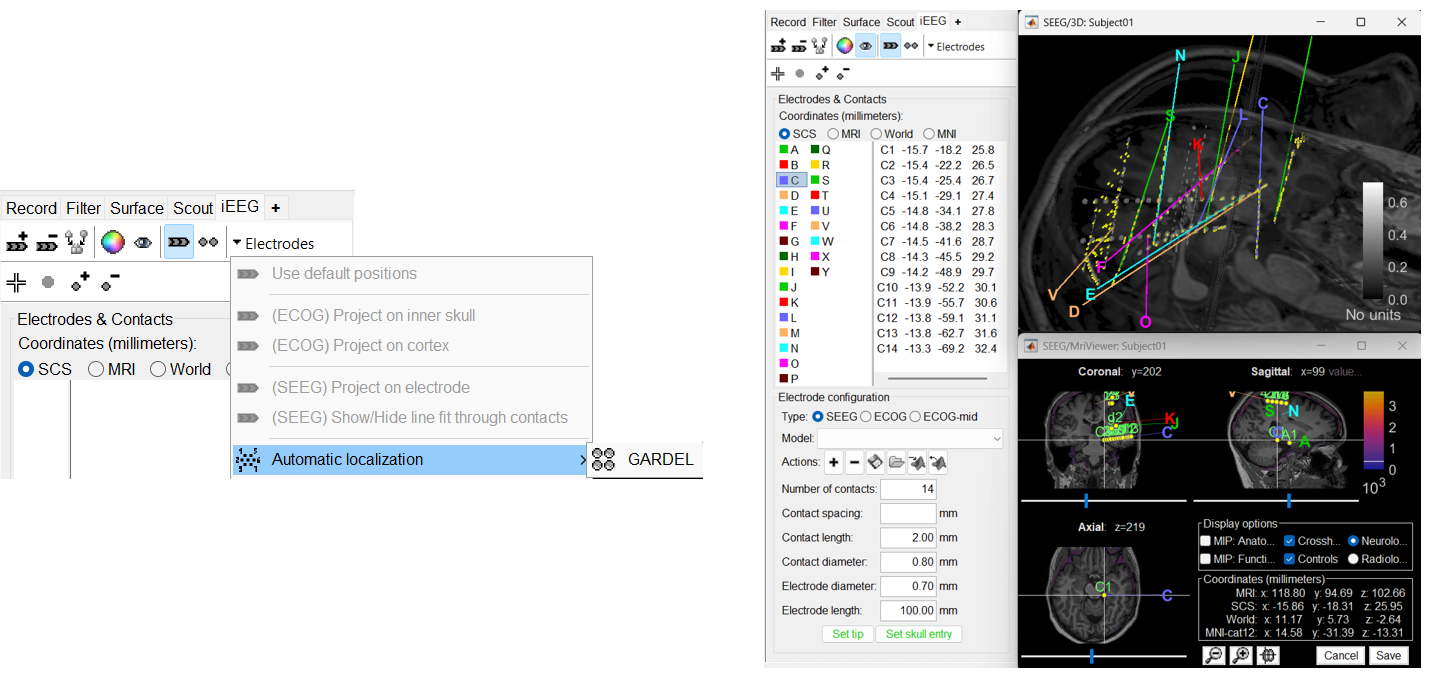
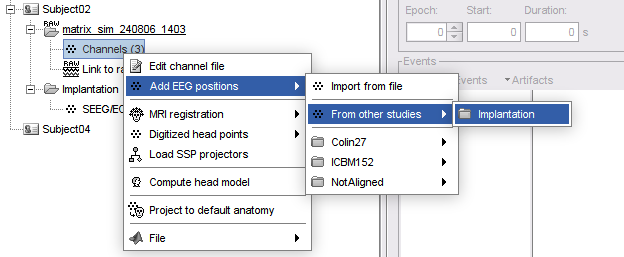
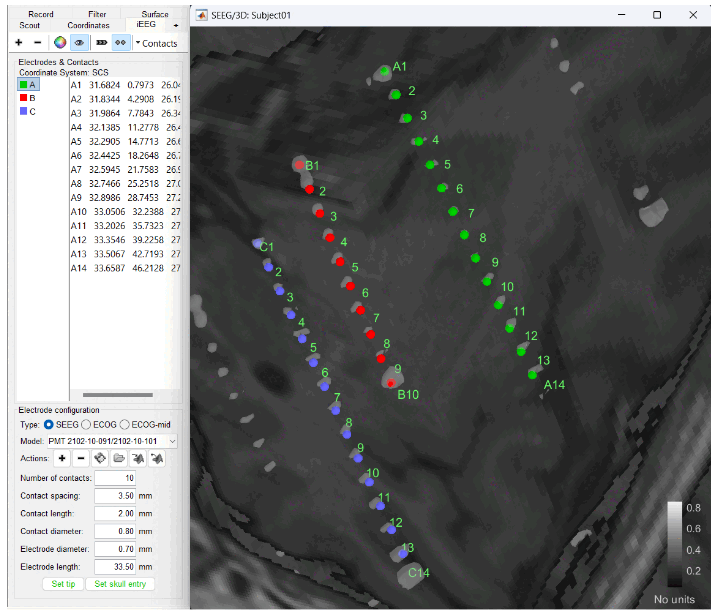
Improved detection of type and group of contacts for SEEG/ECOG channels 🔗
Connectivity
Link TF function matrix and graph figures 🔗
Same colormap behaviour in matrix and graph visualizations 🔗
Input / output
Read and store timestamp for 0s in recording files if available. This time can be used to display actual time in raw and imported recordings. 🔗

January 2026
Anatomy
Add generation of primitive surfaces (sphere, ellipsoid, cube, cylinder, cone) 🔗

Boolean operation between two surfaces (intersection, union and difference) 🔗

BIDS
Coregistration
Manual sensor coregistration, prompts selection of target folders to apply the same coregistreation 🔗
Events
Head model
FEM: Update extraction of FEM layers as surfaces for the case of non fully nested surfaces 🔗
FEM: Extract FEM layers to new FEM file 🔗
Interface
Bugfix: Context menus remained open for Matlab releases using New Deskptop (>=2023b). Context menu is now closed by clicking in the figure or when closing the figure. 🔗 🔗
Recordings
Montage: Bugfix on editing montages with channel separators (:) 🔗
Monatge: Do not allow user montage names to contain [tmp] tag 🔗
December 2025
Database
- Allow adding time to the Acquisition date of a Study

Input / output
Events
Plugins
Add, edit and remove HED tags within Brainstorm GUI with CTagger 🔗 🔗

Automatic EEG artifact removal with GEDAI 🔗 🔗

Software distribution
Add Linux/macOS and Windows wrappers for Brainstorm compiled with >= R2020a 🔗
Update 'spmtrip' functions 🔗
Compilation with Matlab 2026a 🔗
November 2025
BIDS
NIRS, export optode locations as _channel.tsv 🔗
Events
On importing epochs from raw data using events, keep events that were used on importing data 🔗
Head model
Allow refinement of tetrahedral FEM meshes by: (i) FEM layer, (2) ROI.
Left figure: original FEM mesh. Center figure: brain layer (gray) was refined. Right figure: all FEM layers inside the 25mm-radius sphere (transparent green) were refined. 🔗 🔗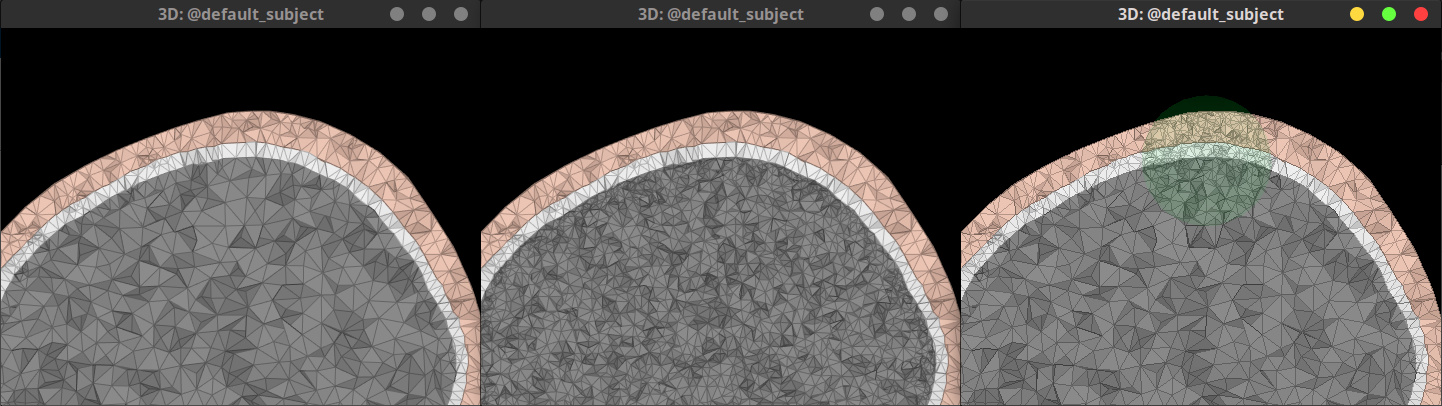
Plugins
Set iso2mesh to follow GitHub master 🔗
Bugfix, interaction between SPM and FieldTrip, setting SPM defaults 🔗
New Processes
New process: File > Select files by event info to select / ignore only files that have events with one of more of the indicated event name(s). 🔗
Software distribution
Distrib: Bugfix on compiling from different OS 🔗
October 2025
Anatomy
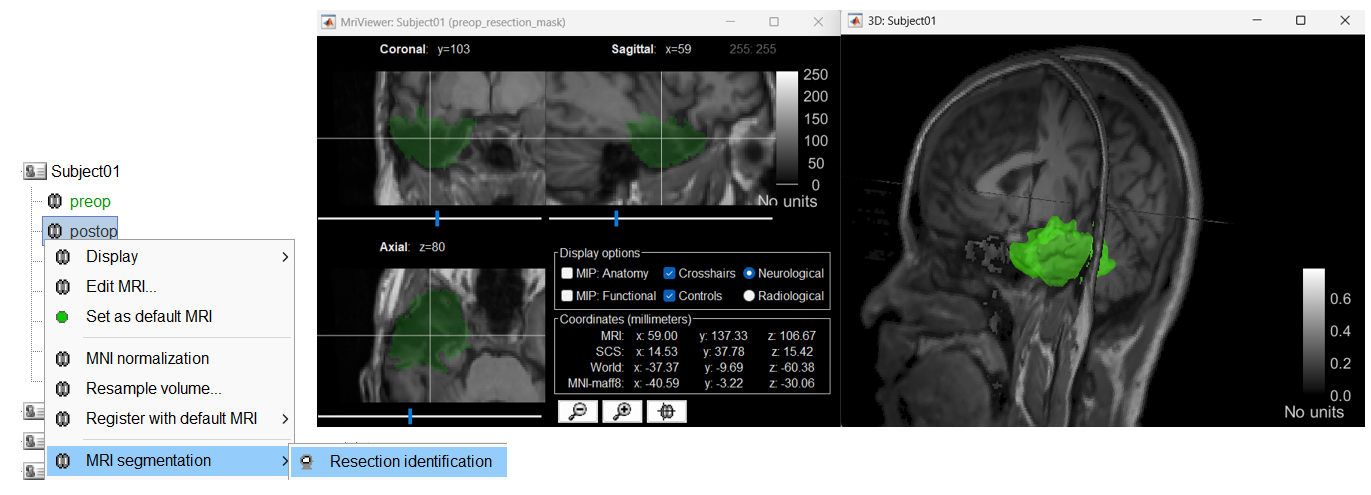
Add plugin "resection-identification" to: (1) coregister pre- and post-op MRIs of subjects that underwent surgical resection, and (2) identify resection and import it in Brainstorm as a volumetric mask 🔗 🔗
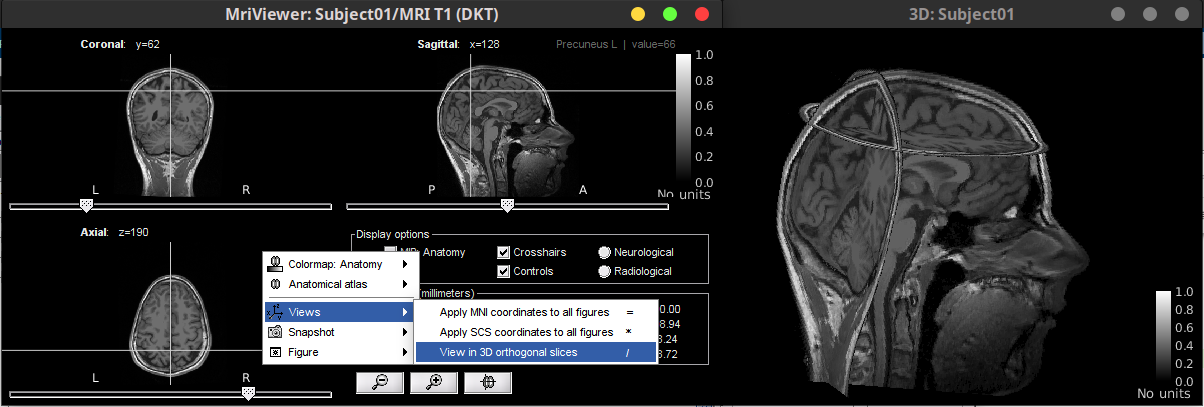
Allow overlaying Anatomy volumes on volume of choice 🔗
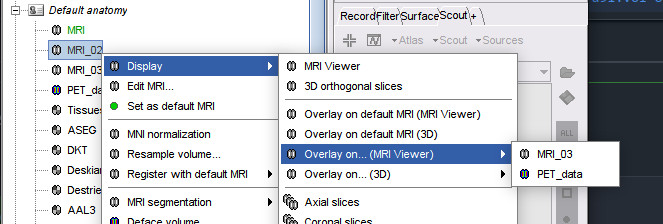
- New option in MRI viewer context menu: "View in 3D orthogonal slices" to set the current slices in the MRI viewer in 3D figure.
EEG
IEEG
Input / output
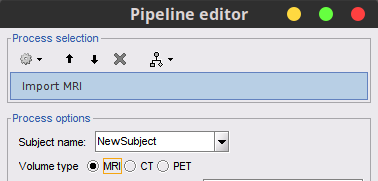
Extend "Import MRI" process to support CT and PET volumes 🔗
September 2025
Anatomy
Allow exporting fiducial locations as BIDS-compatible JSON file alongside MRI file 🔗
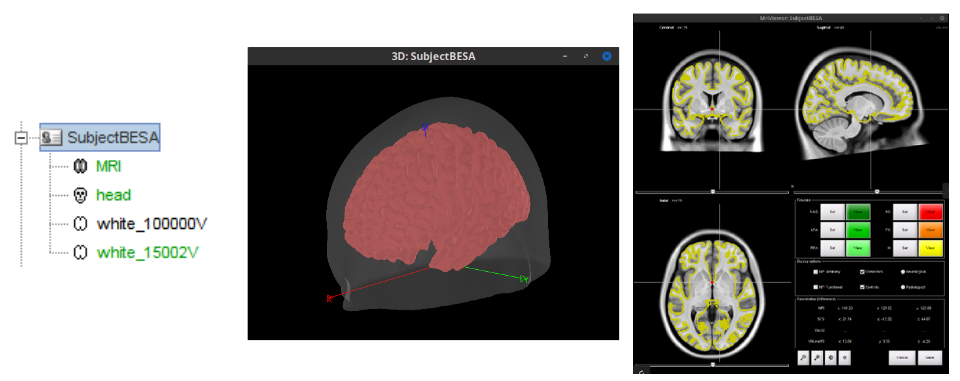
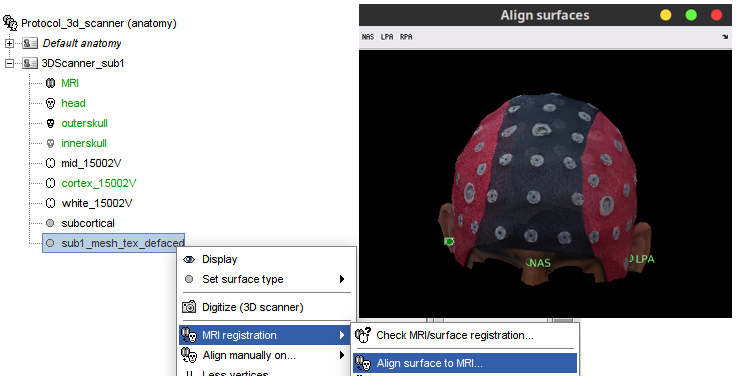
Alignment of (scalp and 3Dscanner) surfaces to MRI, by setting fiducials in surface 🔗
EEG
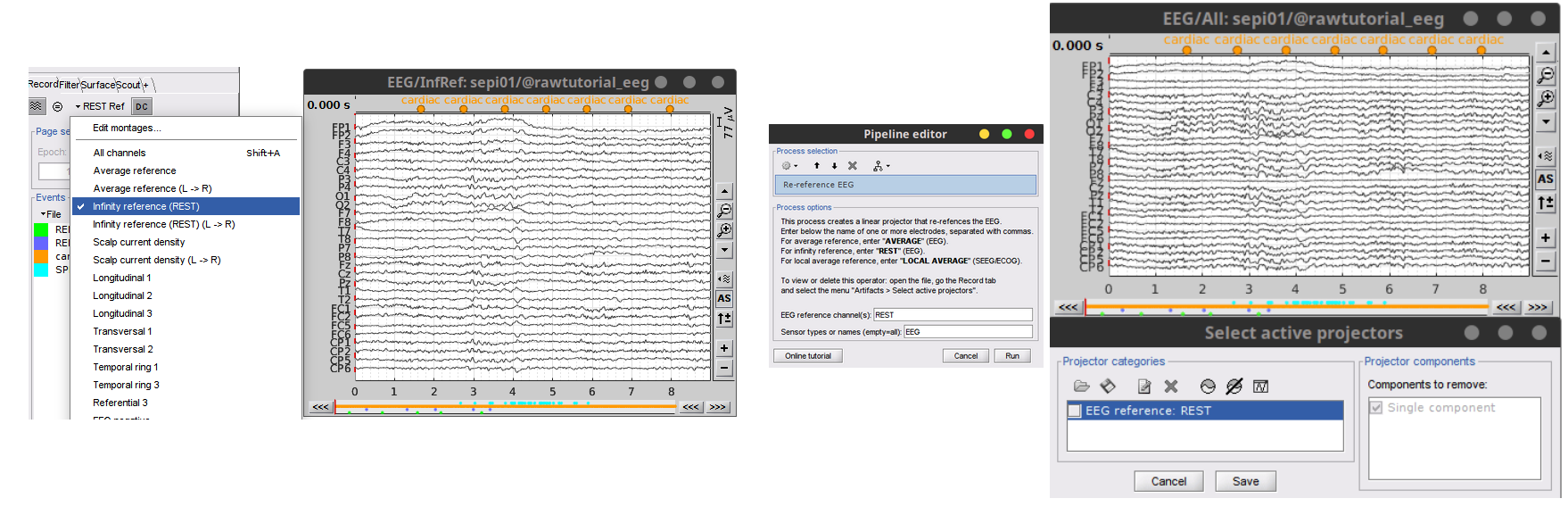
- Reference Electrode Standardization Technique (REST) aka Infinity reference is now available as montage or method for re-referencing
IEEG
Update isosurface from CT thresholding within Surface panel 🔗
Input / output
Add I/O support for text (.jnii) and binary (.bnii) JNIfTI files 🔗
Bugfix: Allow exporting files in any of their valid extensions 🔗
NIRS
Display sensitivity map for NIRS 🔗

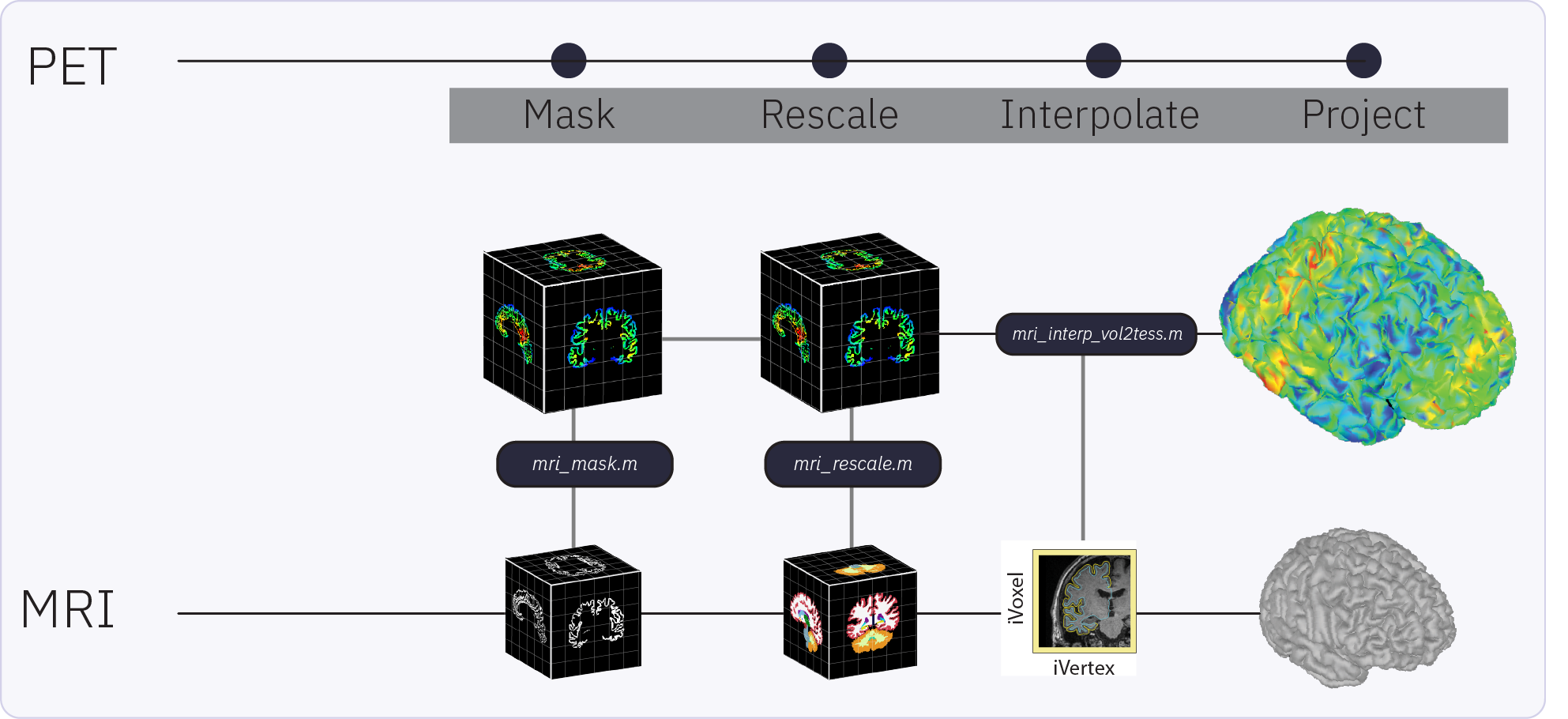
PET
Visualization
Bugfix: Update FEM tensor viz to follow moved slice in 3D figures 🔗
Use Lambert azimultal equal-area projection for 2D projection of 3D sensor locations 🔗
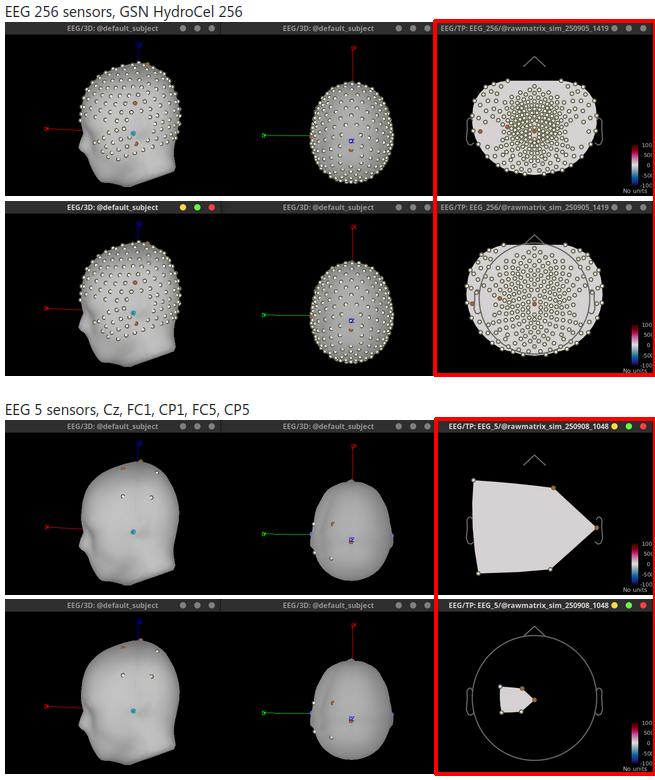
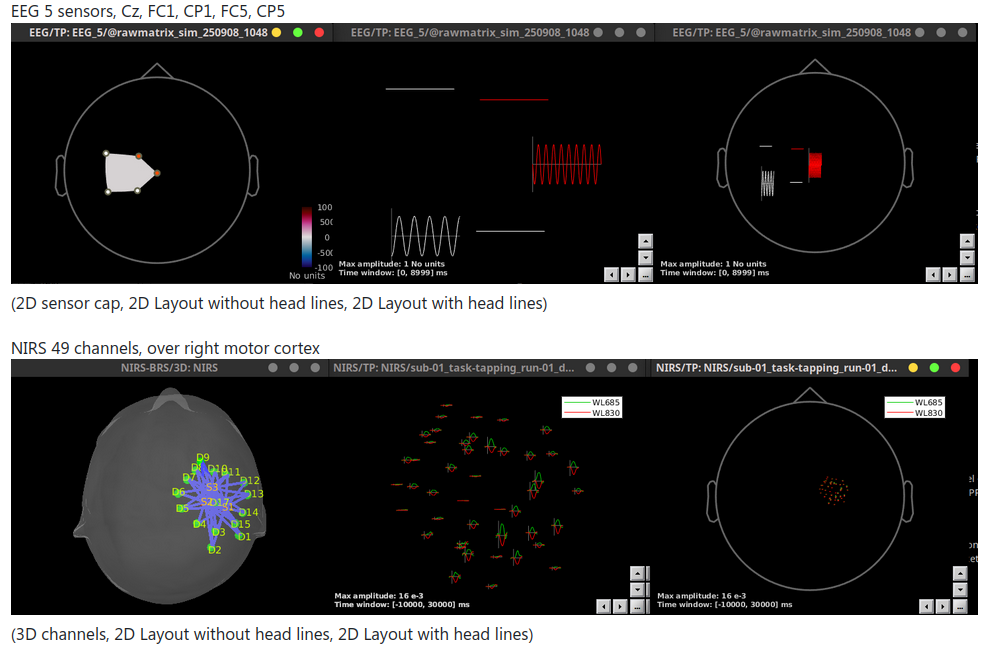
Add options of head lines in 2D Layout figures 🔗
Software distribution
Update 'spmtrip' functions 🔗
Compilation with Matlab 2025b 🔗
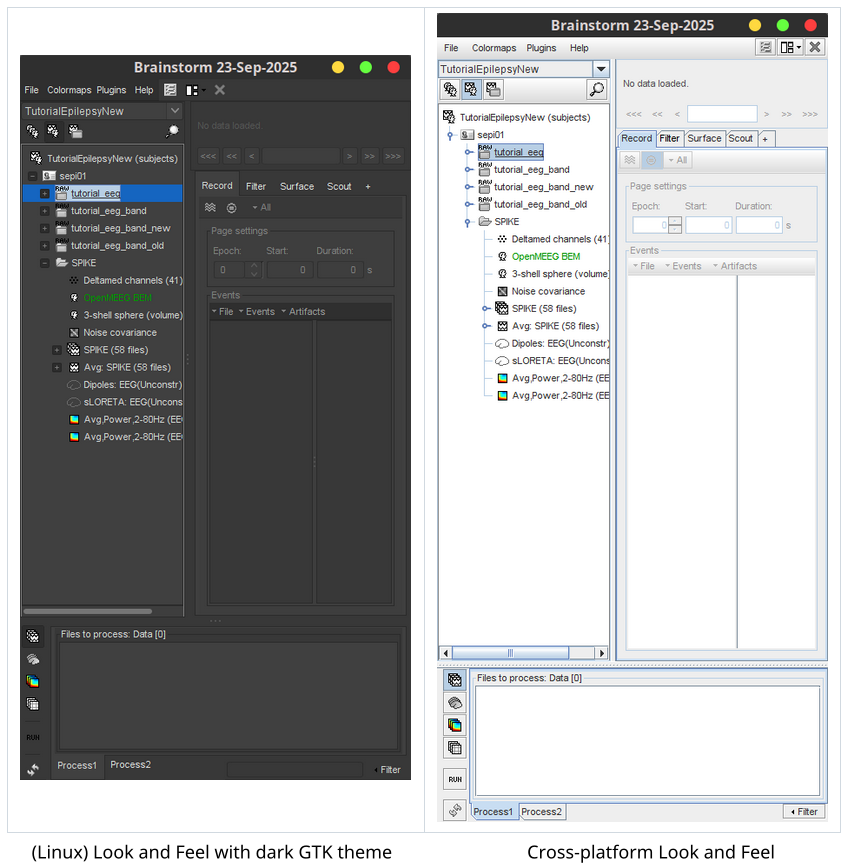
Add option for cross-platform Java Look and Feel in compiled 🔗
August 2025
Anatomy
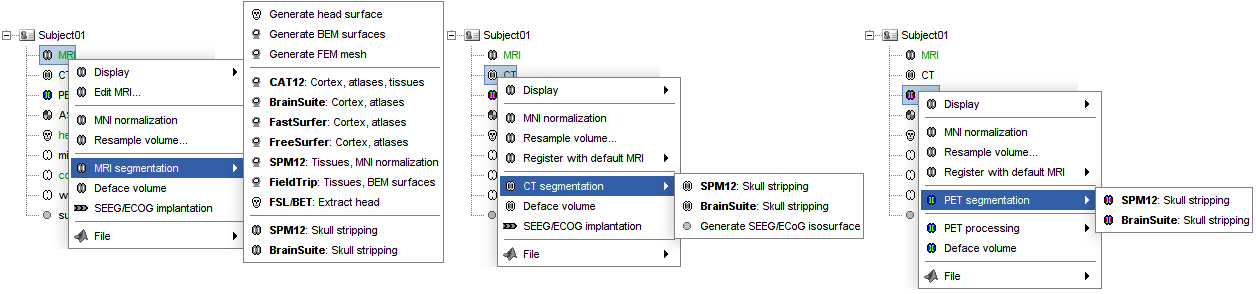
Skull stripping is now available for all volumes (MRI, CT and PET), and it does not require same volume size for coregistered CT and PET 🔗 🔗
BIDS
Connectivity
Handle flat signals for Granger causality 🔗
Update in time- and frequency-domain Granger causality. Recommended GC implementation uses the MVGC Toolbox 🔗
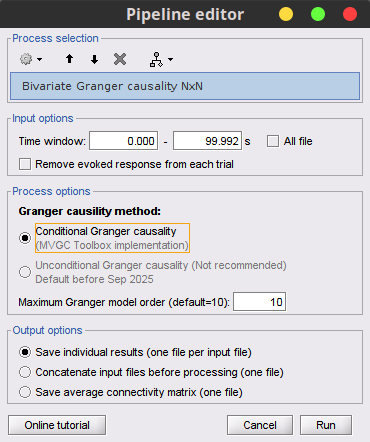
Viz, only hide self-connectivity if same channel from same file 🔗
Database
Bugfix: keep default anatomies for moved or copied Subject 🔗
Update: Head model, include NIRSmethod field in Study.HeadModel struct 🔗
Head model
Better integration of NIRS in GUI and process to compute head model 🔗
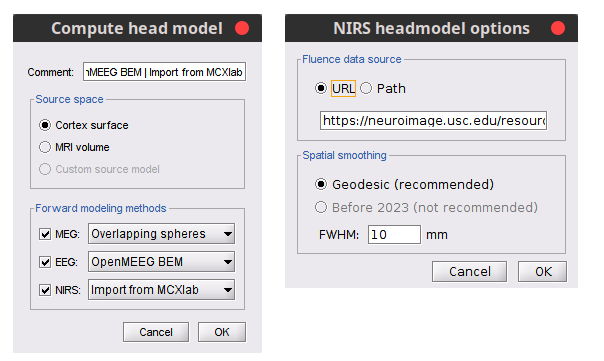
IEEG
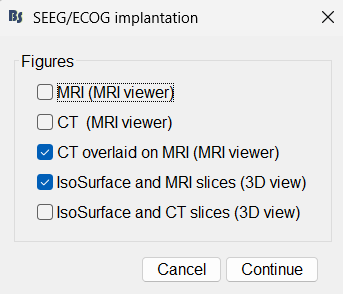
Simultaneous MRI view figures (e.g., to display MRI and CT in different figure) 🔗
Non-interactive call for anatomy labeling 🔗
- Improve ON/OFF indicator for tool to find centroid of surface "blobs"
Bugfix: Use correct surface when finding centroid of surface "blobs" 🔗
Bugfix: Show correct units for ECOG+SEEG plots 🔗
Input / output
Visualization
Improved 2D-Layout plot positioning for MEG, EEG, NIRS, ECOG and SEEG 🔗
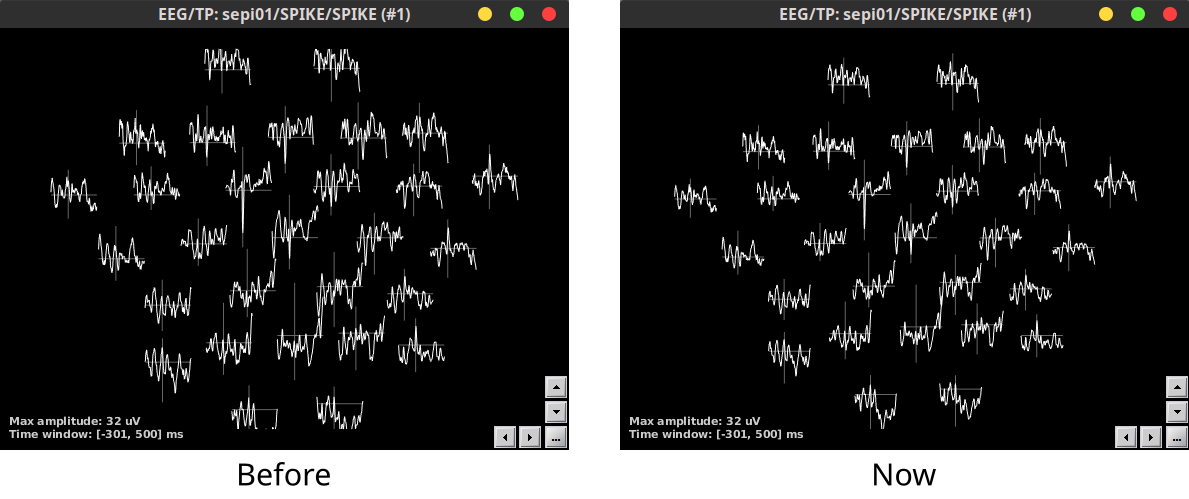
Software distribution
July 2025
Anatomy
On importing NIfTI files, vox2ras transformation from sform is preferred over qform if both are present 🔗
Allow non-interactive call for tess_force_envelop.m 🔗
Add various methods to smooth a surface mesh 🔗
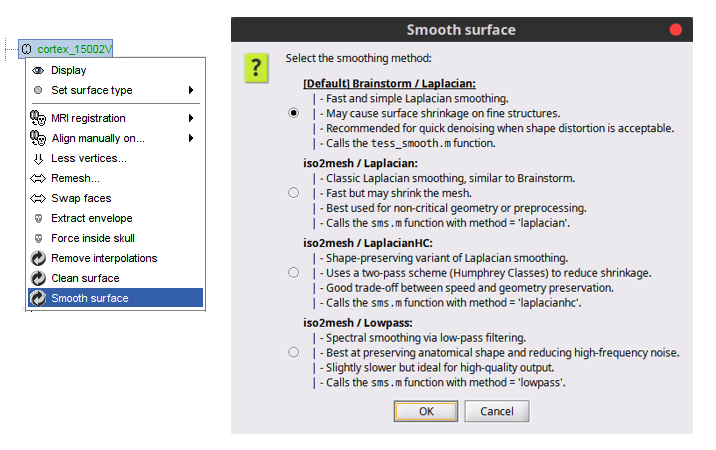
Compute mesh statistics for surface and volume meshes 🔗

BIDS
On importing BIDS-formatted datasets, the preferred names and locations for EEG electrodes are obtained from the _channels.tsv and the _electrodes.tsv files respectively over the names and locations in the EEG recording file 🔗
Events
Import/export events as BIDS (_events.tsv), the column onset is relative to beginning of recording 🔗
Export events as BIDS (_events.tsv), add channel information in column channel 🔗
Bugfix: Merge events, channels and notes are now properly handled 🔗
IEEG
Software distribution
- SPM12 is available as method to coregister, realign and skullstrip in compiled version of Brainstorm
June 2025
Coregistration
Bugfix: Correct conversion for locations of EEG points from .pos file on reviewing CTF data 🔗
Digitize: Improved support for Polhemus-Patriot device 🔗
Connectivity
Allow NaN as value for connectivity metrics. These values are not shown on figures 🔗
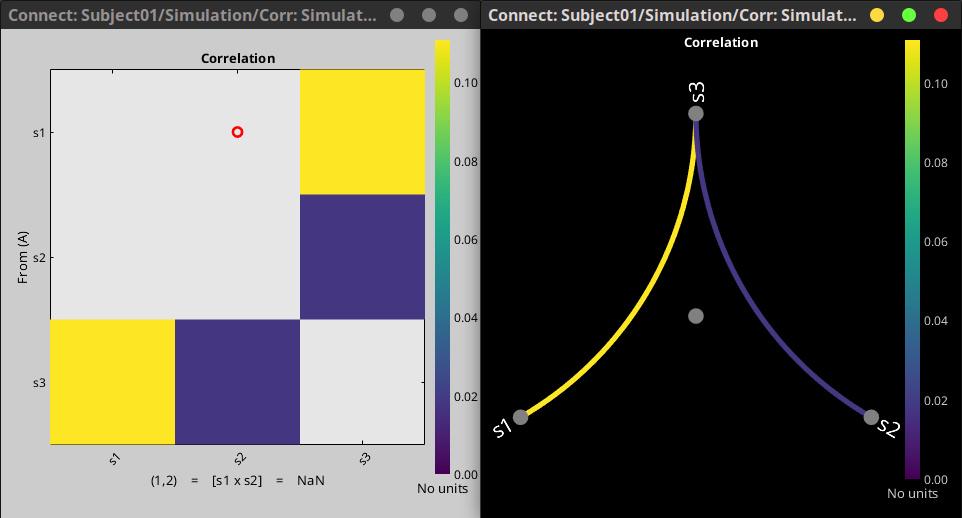
Visualization
For source PSD analysis, improved specparam (FOOOF) model figures (sources level, and vertex level) 🔗
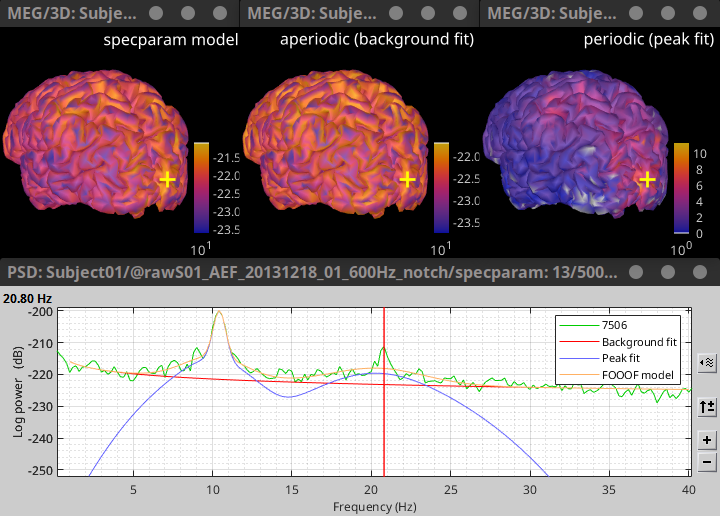
- Remember last display option [power, magnitude, log(power)] for TF representations
Scripting
May 2025
BIDS
Importing BIDS-formatted datasets, allow EEG recordings stored as ANT Neuro EEProbe
(despite not being part of the BIDS specification) 🔗 🔗
NIRS
April 2025
Anatomy
PET: Basic support to pre-process PET volumes (realign, smooth, aggregate frames) 🔗
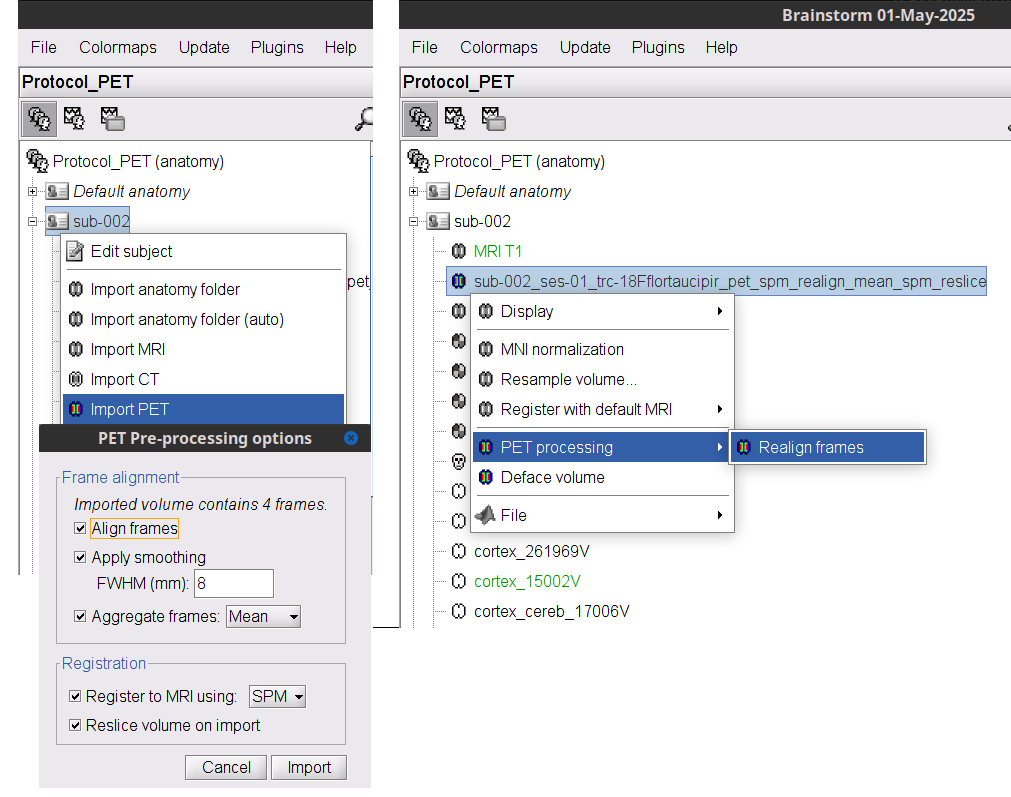
IEEG
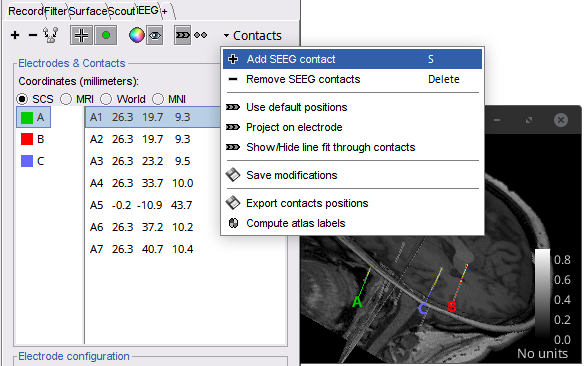
Add switch to select (or not) the centroid in a IsoSurface 🔗
Add/remove individual contacts in SEEG electrodes 🔗
Events
Software distribution
Server: Update URLS for neuroimage.usc.edu from HTTP to HTTPS 🔗
March 2025
Anatomy
Scouts: Export surface scouts as MRI mask 🔗
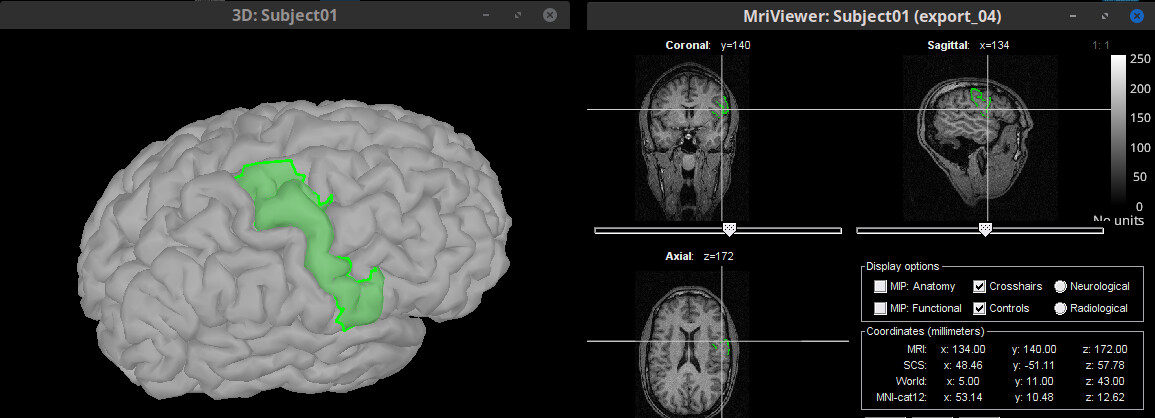
Add mark on colorbar to indicate threshold on overlayed anatomies (MRIview, 3D) 🔗
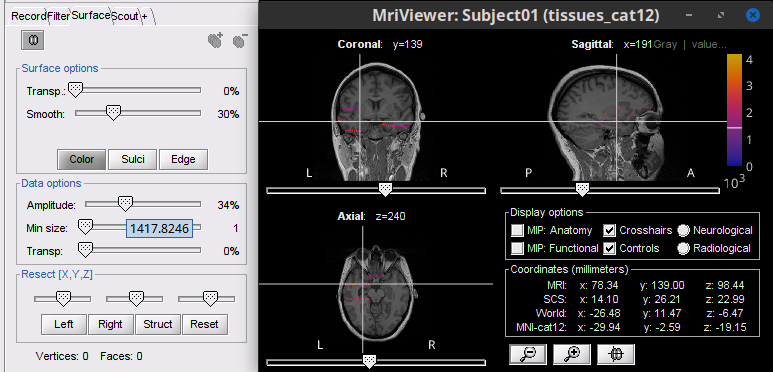
Add "isosurface" as available surface to add. Improve slider behaviour 🔗 🔗

Timefreq
Correct sign for imaginary part of the analytical signal using hilbert() 🔗
This change does not affect any previous result using hilbert() as the sign flip was consistent.
Software distribution
Bugfix: Improved detection of write permissions in Windows links and OneDrive folders 🔗
February 2025
Anatomy
Processes
Add support for frequency bands on computation of PSD features 🔗
Add option to have z-scores as output on comparing to normative distribution 🔗
GUI: Handle nested controller-classes in process options 🔗
Software distribution
Compilation with Matlab 2025a 🔗
December 2024
Anatomy
Scouts:, Options Copy and Paste to copy-pasting scouts (same atlas or between atlases).
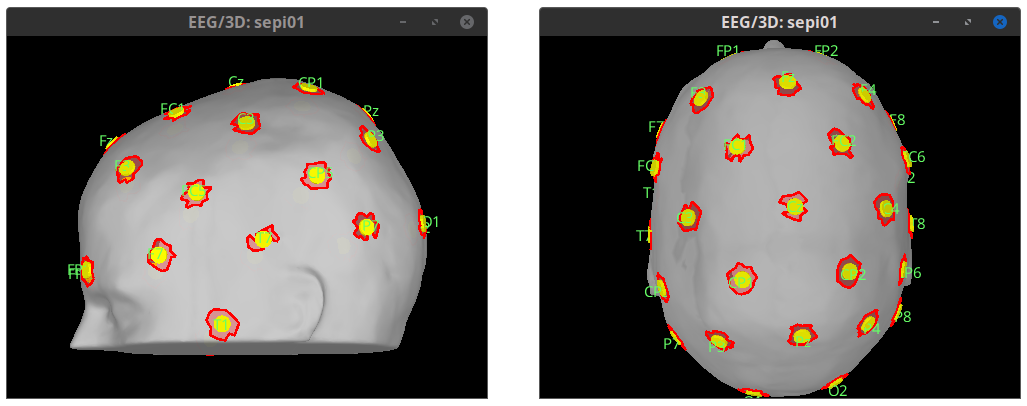
Scouts, create scalp scouts from sensor positions. Bellow, scalp scouts (red) were generated foor a radius of 10 mm around the EEG sensors (yellow) 🔗
Machine learning / Decoding
SVM decoder, Bugfix, allow ignore bad channels 🔗
Digitization
Software distribution
November 2024
Anatomy
Add NIRS template: 'Colin27_4NIRS_2024' 🔗
Integration of skull stripping (using SPM or BrainSuite) within Brainstorm 🔗
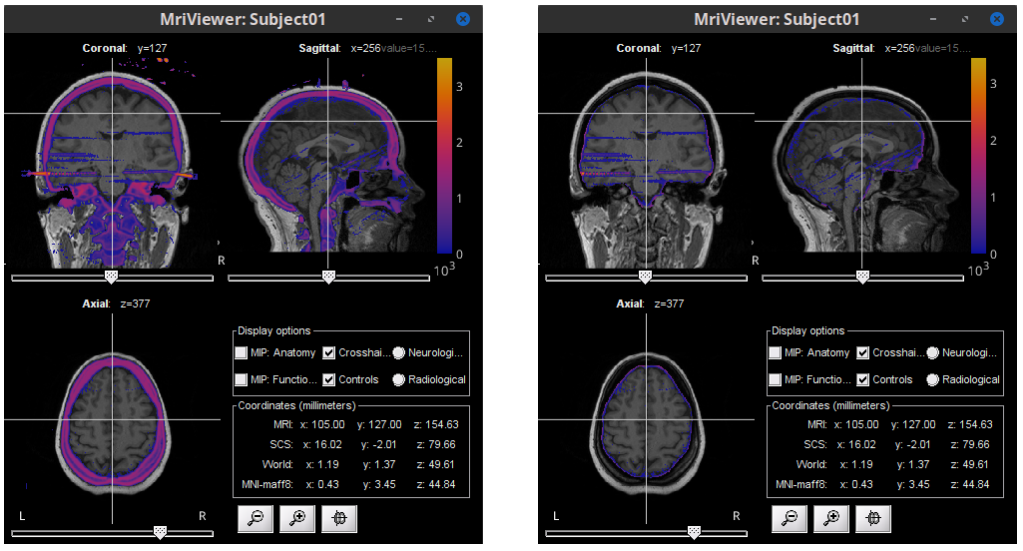
SEEG/ECOG
Allow iEEG implantation using MRI and/or CT and/or IsoSurface 🔗
Default EEG caps
Assess the registration between default anatomies and default EEG caps 🔗
Add caps 'ANT Wavegard 65', 'BrainProducts ActiCap 68', and 'WearableSensing DSI 24' for default anatomies 🔗
Coregistration
Plugins
Update iso2mesh, brain2mesh and xdf plugins to support Apple Silicon 🔗
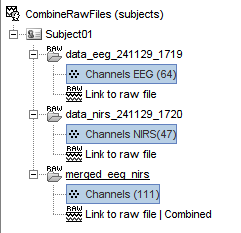
Input / output
Allow to merge (channel-wise) raw continue recordings with the same duration 🔗
Scripting
Update script for epilepsy tutorial, add source estimation with MEM 🔗 🔗
Script to automate the BEst (Brain Entropy in space and time) tutorial 🔗 🔗
October 2024
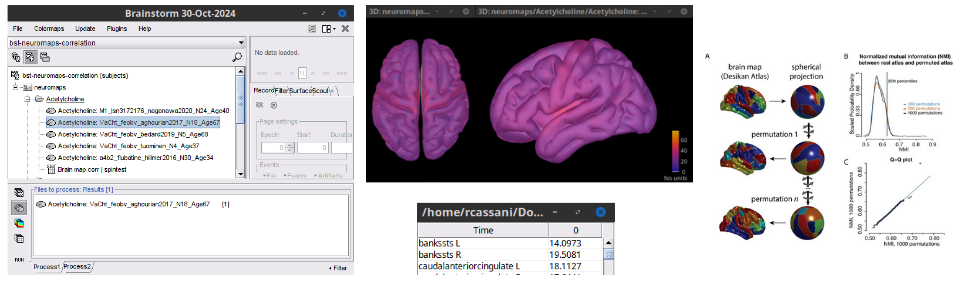
Anatomy
Add bst-neuromaps plugin and its tutorial: 🔗 🔗
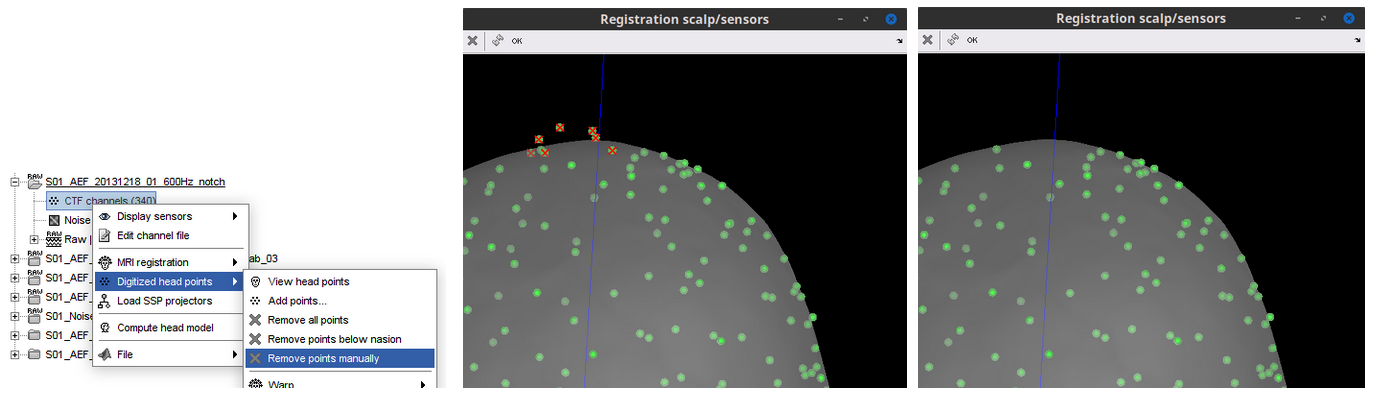
Allow selecting and deleting head points from 3D figure 🔗
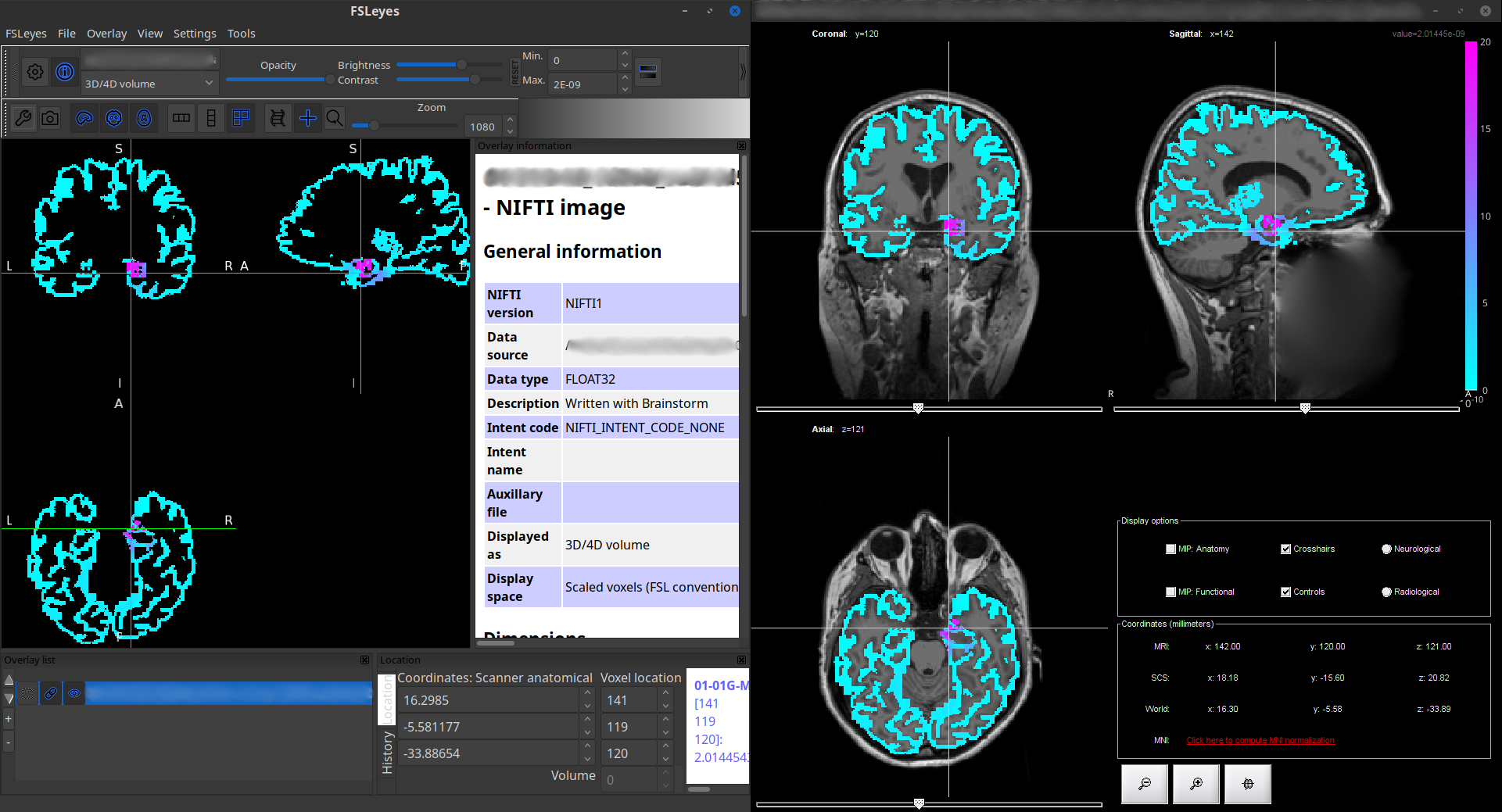
Export mixed source files as NIfTI files 🔗
Expose manual setting of background level ('bgLevel') in tess_isohead() 🔗
Time frequency analysis
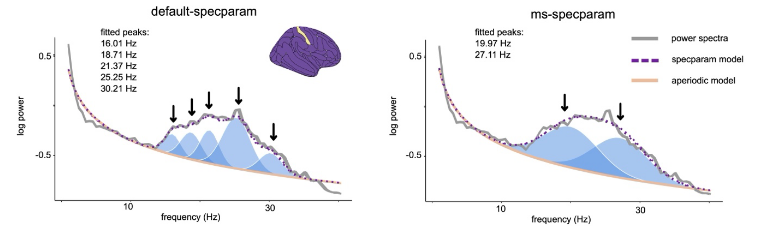
Model selection (ms) for FOOOF (aka specparam) and SPRiNT 🔗 🔗
Update macOS (Intel and Apple Silicon) .mex files for PAC 🔗
Input / output
Support multiple-rate EDF files, through EDF reader from FieldTrip 🔗
Bugfix: correct scaling of MEG sensor positions for data from FieldTrip 🔗
NIRS
Bugfix: ensure that wavelengths are integers 🔗
September 2024
Artifacts
ICA: Explained variance per IC is now shown 🔗
Scripting
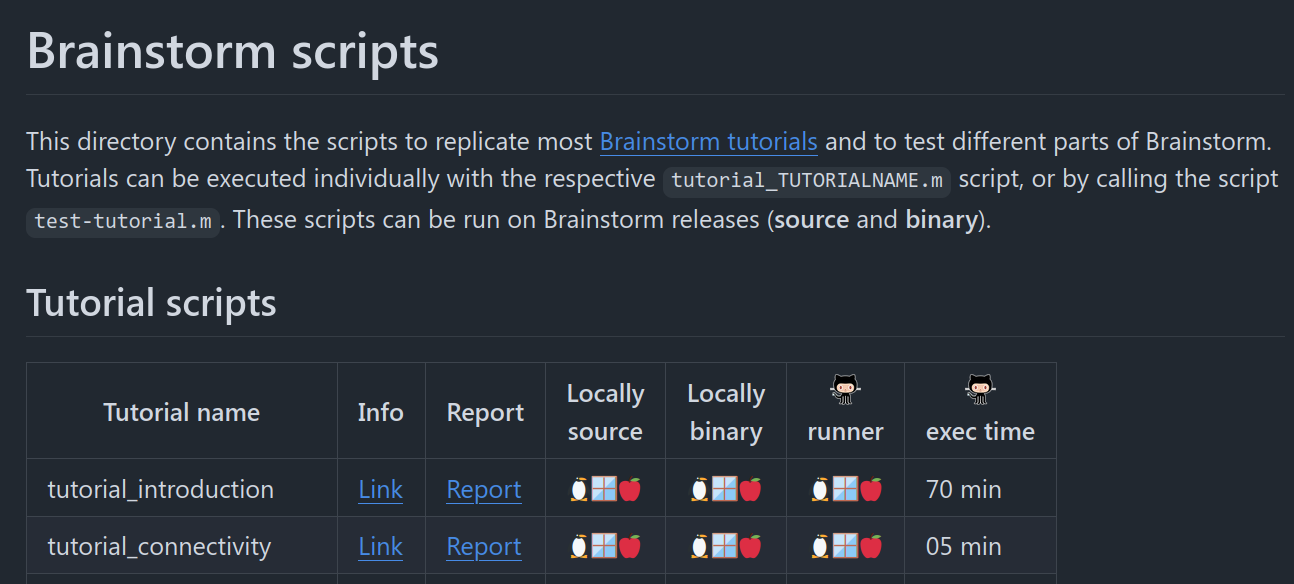
Script to run different Brainstorm tutorials test_tutorial.m: 🔗
Reproducibility
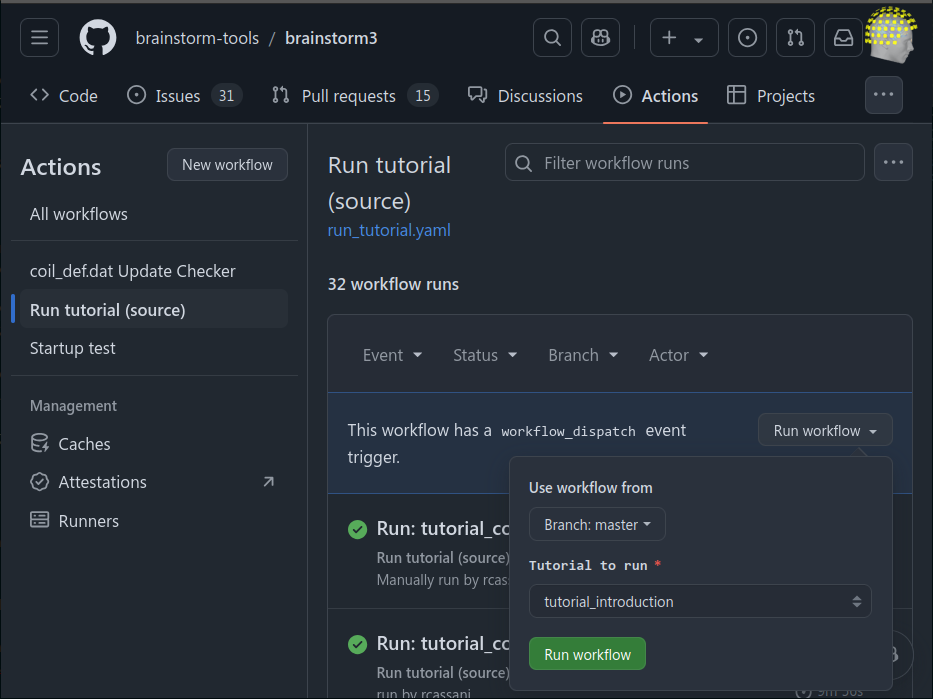
Allow running test_tutorial.m for a subset of tutorials using GitHub workflows 🔗
Process: Add input of types list_vertical and list_horizontal for processes 🔗
Allow cells and matrices as script argument for Brainstorm server 🔗
Plugins
Bugfix: Wrong interactions between SPM and FieldTrip plugins on source code 🔗 and standalone 🔗
Bugfix: Wrong interaction between MNE functions in external and from SPM or FieldTrip plugins 🔗
Bugfix: Clean persistent variables on Loading FieldTrip plugin 🔗
Interface
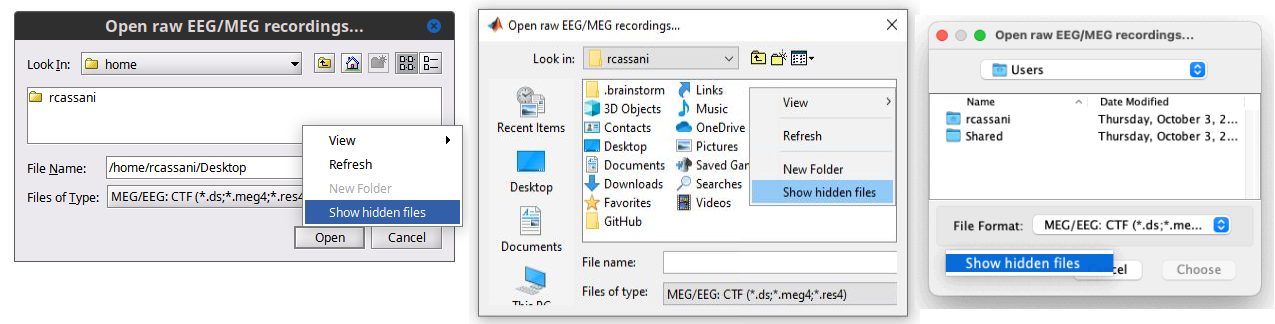
Show/hide hidden files in the File Selector window 🔗 🔗
Report: Add 'Elapsed time' and errors/warnings count to text-only report 🔗
Optimization: Faster generation of Plugin menu 🔗
Bugfix: Setting current Protocol after renaming protocol 🔗
- Text based report has time initial time stamp and error/warning counters
August 2024
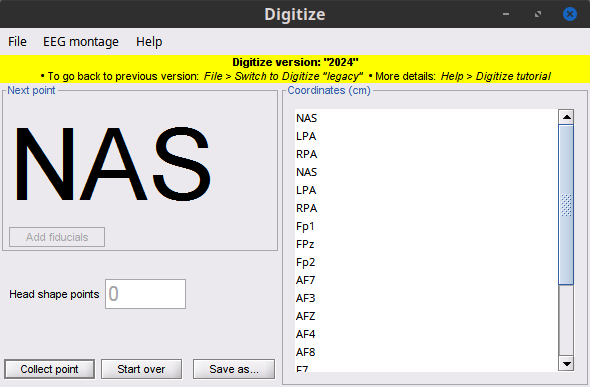
Coregistration
Digitize: New panel, more features 🔗
IEEG
Processes
Add computation of deviation maps from PSD features. Tutorial 🔗
Allow setting files in Process tabs with processes File > Select files: ... 🔗
Scripting
Script to automate the DBA (Deep Brain Activity) tutorial 🔗
Process: Add class-controller support combobox_label 🔗
Anatomy
Scouts, improve volume calculation 🔗
Input / output
EDF+ support for negative gains 🔗
Software distribution
Linux and macOS, standalone version can automatically find Matlab runtime engine in full Matlab installation 🔗
Update ICBM template on compilation 🔗
July 2024
Anatomy
Bugfix: importing non-Atlas MRI volumes given in MNI space 🔗
Add category Others for anatomy templates that do not correspond to any of the above
Artifacts
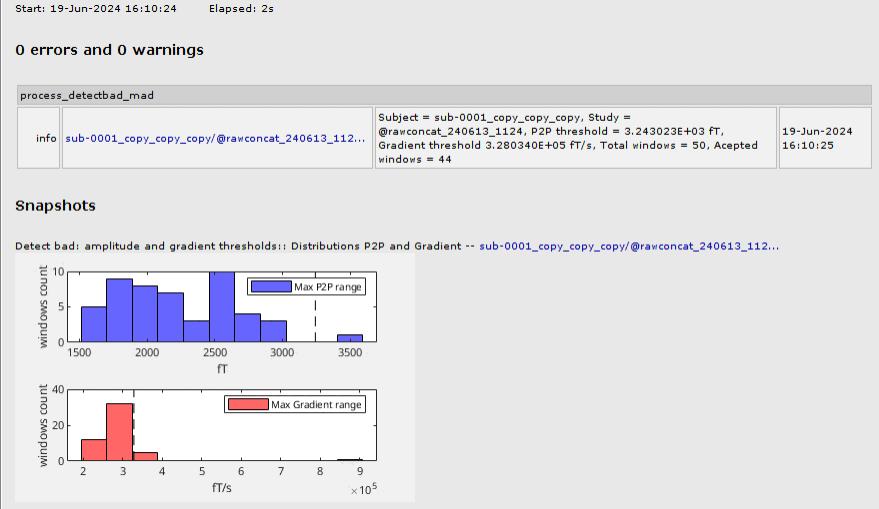
Automatic method to detect artifacts, based on thresholding the peak-to-peak amplitude (p2p) and the gradient of the signal using N times the median absolute deviations (MAD) 🔗 🔗
Head model
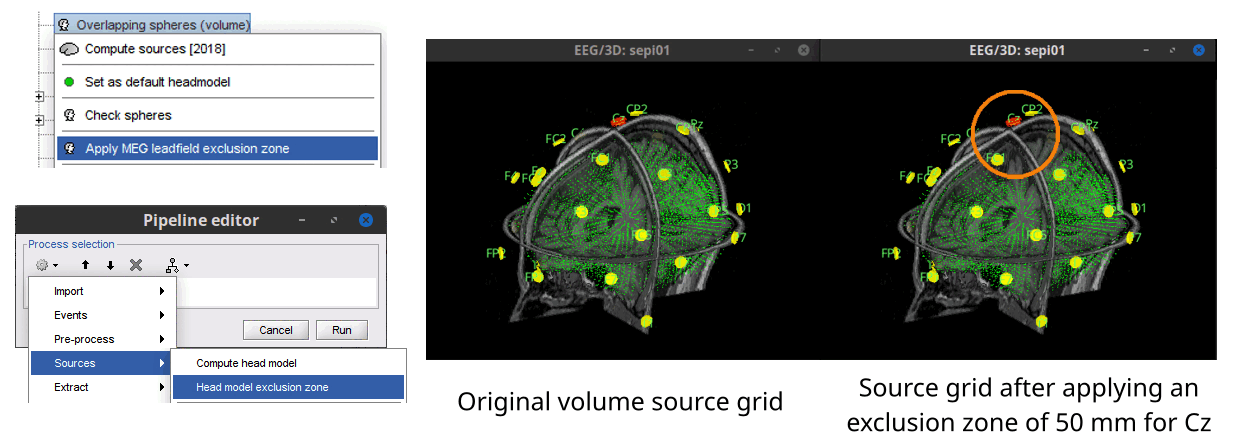
Process to create an exclusion zone around sensors in volume source grids 🔗
Plugins
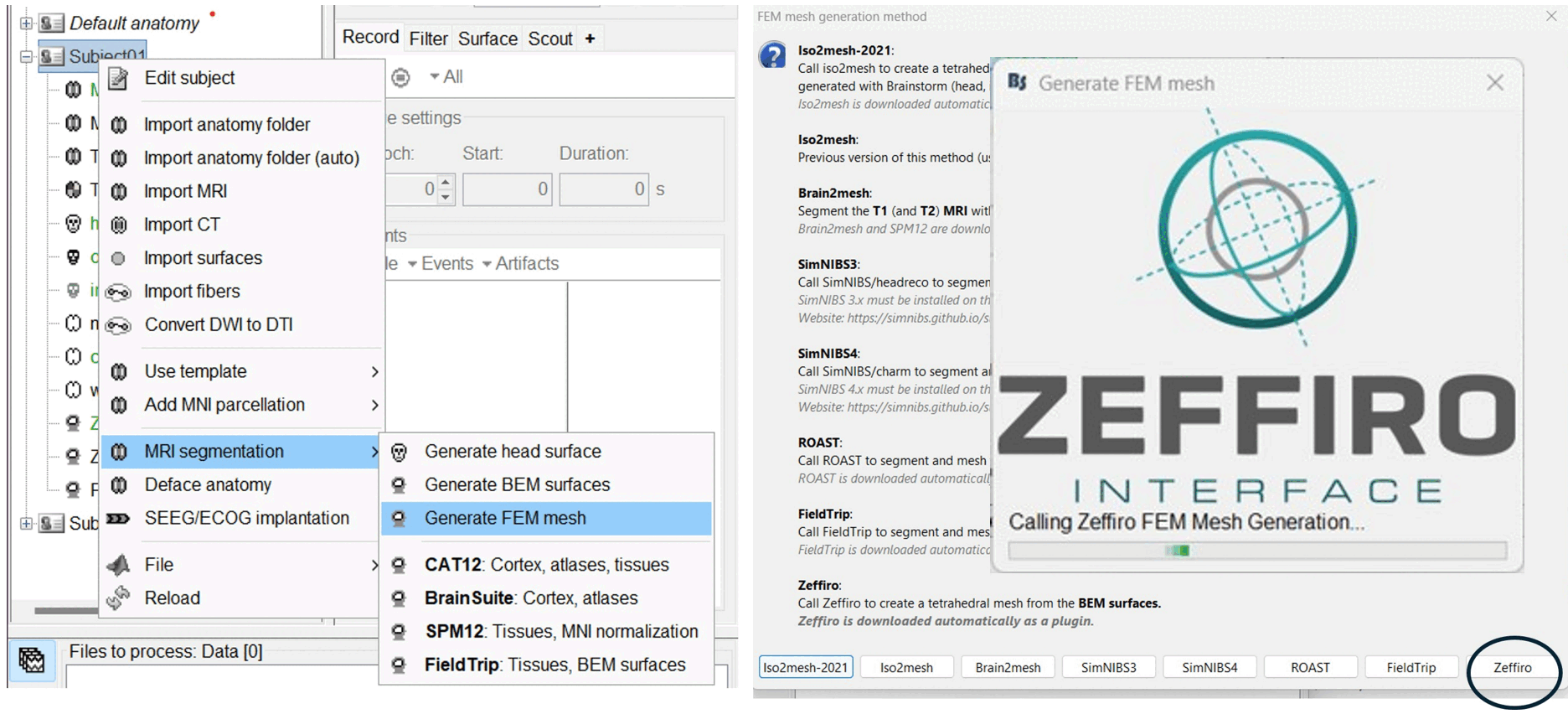
Add Zeffiro plugin for FEM mesh generation 🔗
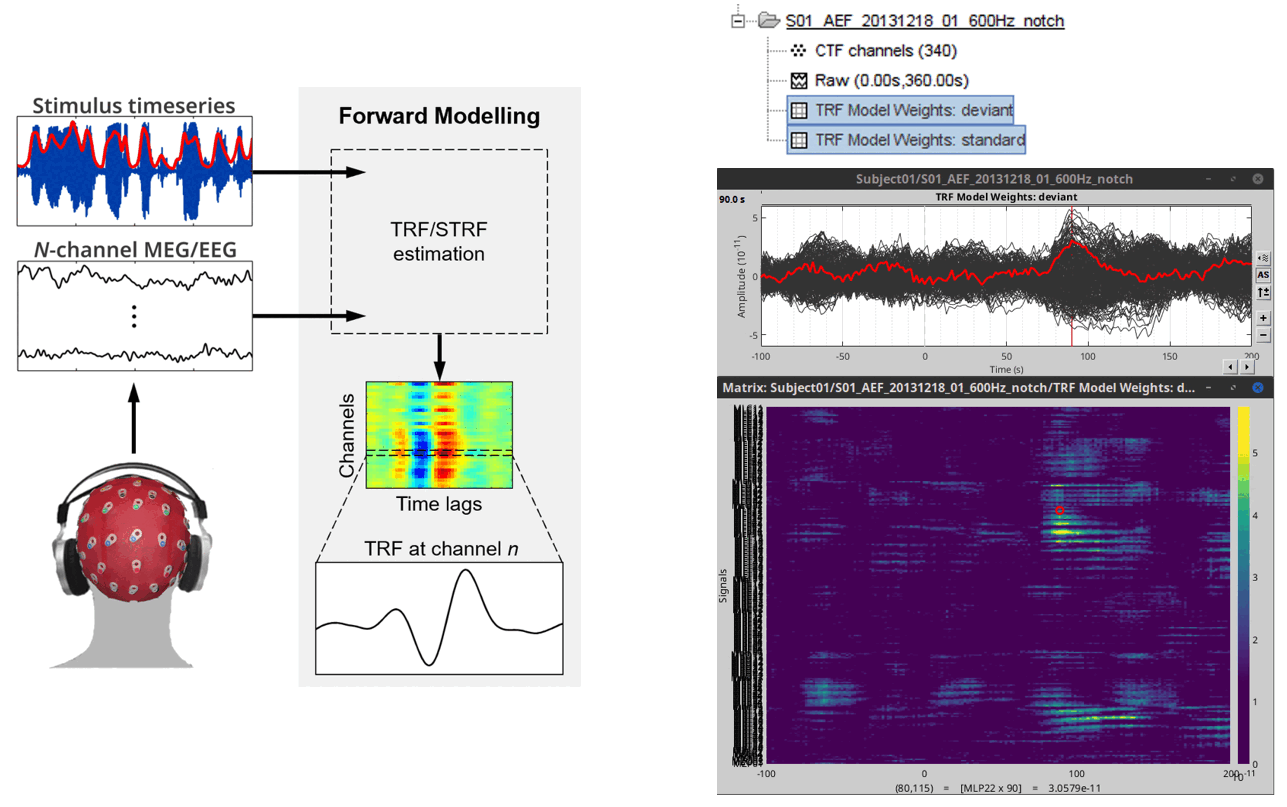
Add the mTRF-Toolbox plugin for multivariate temporal response function analysis process and tutorial 🔗 🔗
Update OpenMEEG plugin to support Apple Silicon (as v2.5.8) 🔗
Update mxclab plugin version, also add Apple Silicon support 🔗
Scripting
Allow to export/import stored pipelines as Matlab variables with:
bst_get('Pipelines') and bst_set('Pipelines') 🔗
Software distribution
Add System info menu to show information about Brainstorm, Matlab and Computer
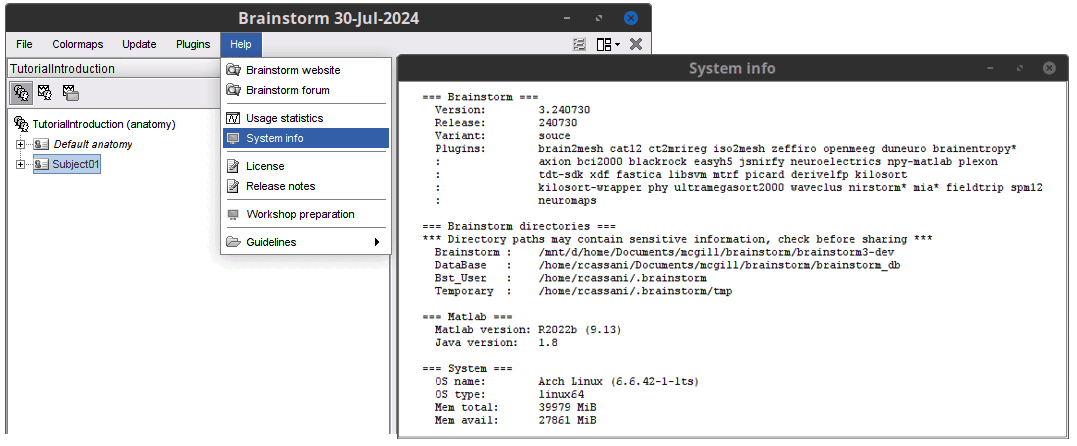
June 2024
Anatomy
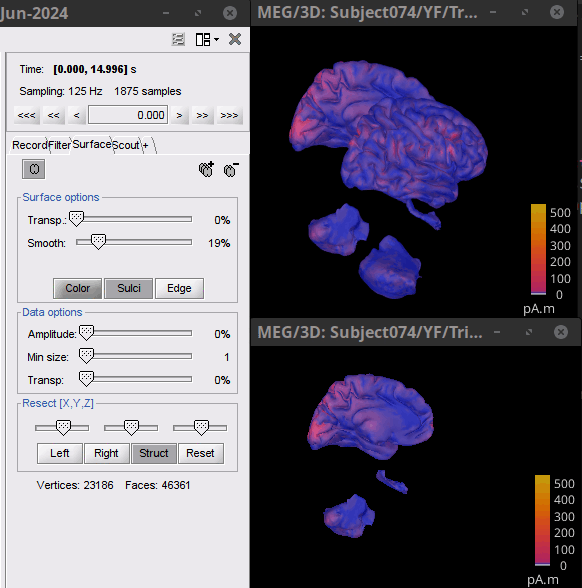
Resection: Allow combining multiple resection approaches. E.g., Struct and y-axis (SCS).
Contactsheets: Contactsheets from MRIviewer support overlayed Atlases and Volumes. E.g., Sagittal contactsheets of AAL3 atlas on Default anatomy.
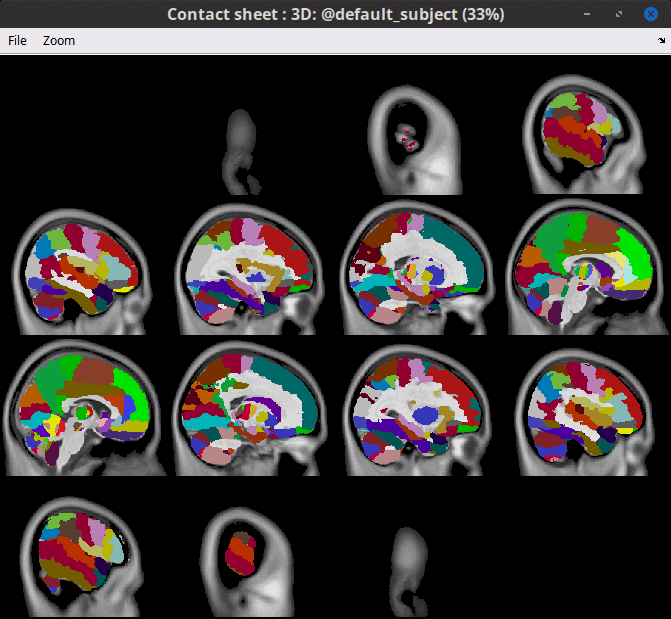
Input / output
Intan RHS: Bugfix, incorrect block selection 🔗
Neuralynx: Bugfix reading NSE files 🔗, added support for NTT files 🔗
Software distribution
Brainstorm is now compiled with Matlab 2023a: Installation instructions
May 2024
Anatomy
Bugfix at selecting proper number type (int16 or int32) when exporting MRI as NIfTI. 🔗
Events
Add option Every sample to convert extended events to a set of consecutive simple events. 🔗
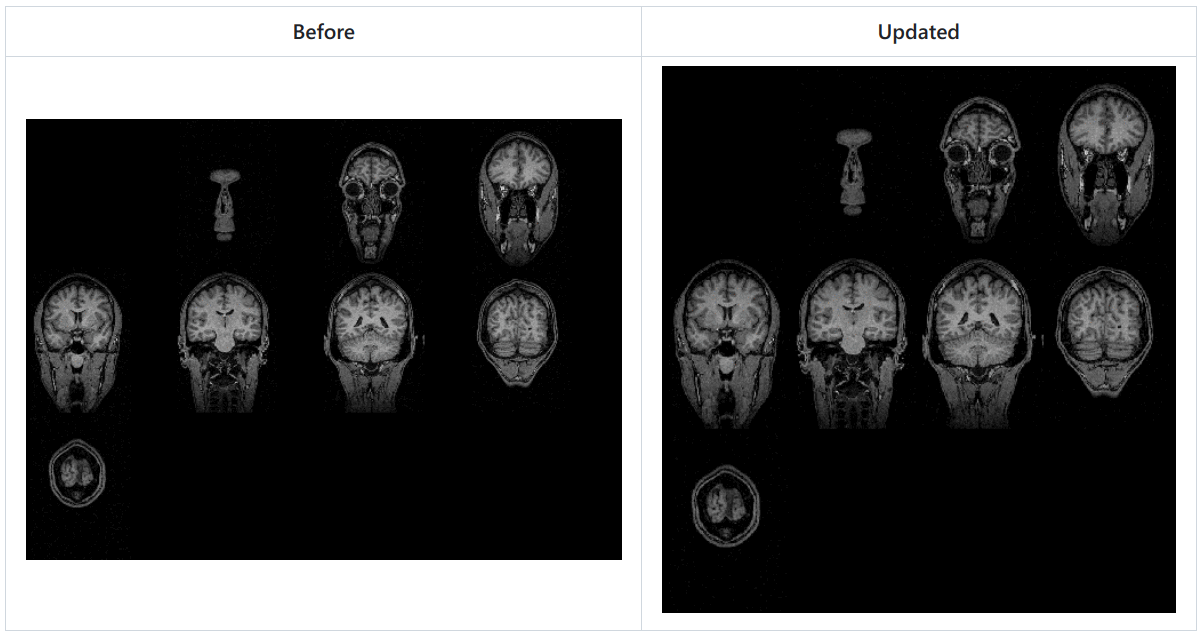
Update contactsheets
Add colormap and timestamp for contactsheets from MRI viewer and 3D slices. 🔗
- Cleaner useless background, each contact image in sheet has the same size.
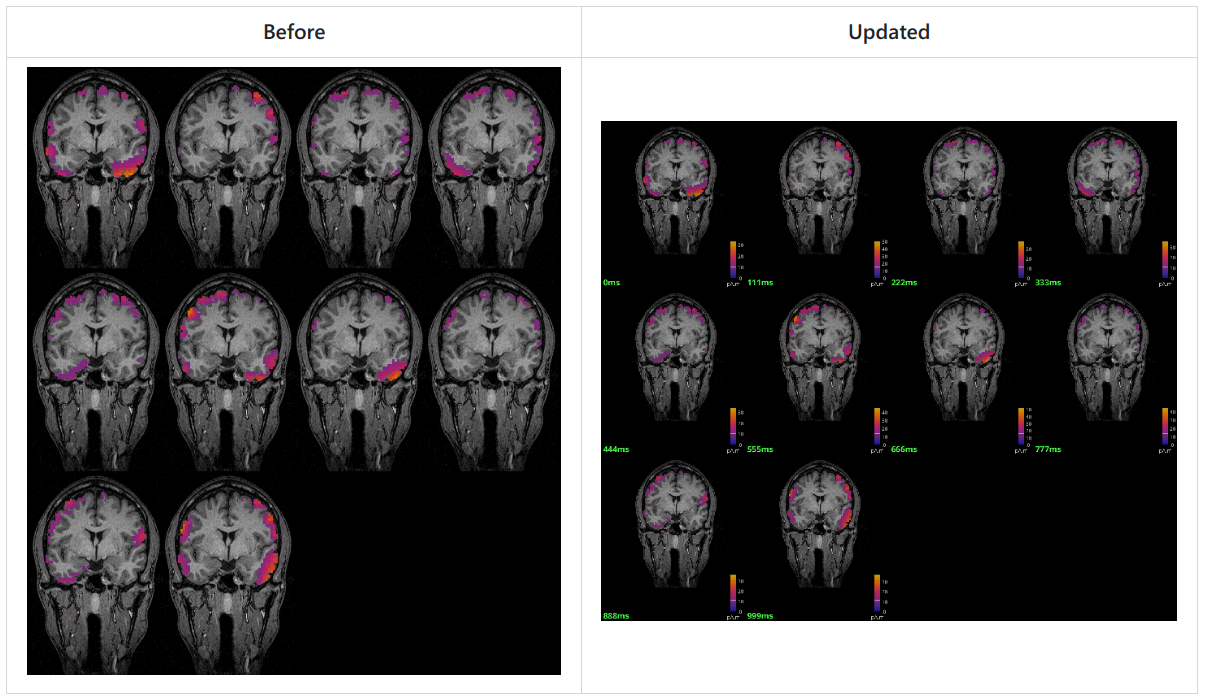
Input / output
Open Ephys: Support for recordings performed OE-GUI >=0.6.0 🔗
.fif: Add support for double, complex-float and complex-double data 🔗
Plugins
npy-matlab: Moved from /external to plugin. 🔗
User-defined plugins User can register plugins to list of supported plugins. 🔗 and 🔗
April 2024
Anatomy
AAL1: This parcellation now available in the Add MNI parcellation menu. 🔗
2D Layout spectral data
Plot frequency-resolved data from sensors in a 2DLayout. Different spectral files: PSD, FFT, and FOOOF results, and available for all sensor modalities (EEG, MEG, SEEG, etc). Compare easily averages between experimental conditions: Select multiple PSD files in the database explorer, right-click on any of them > 2DLayout. 🔗
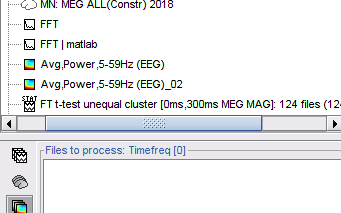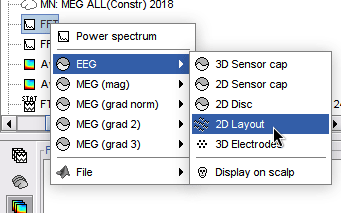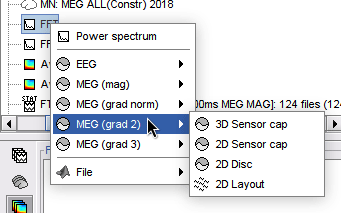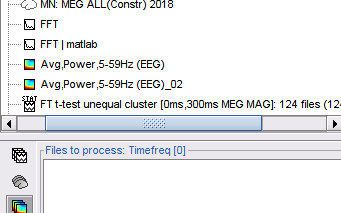
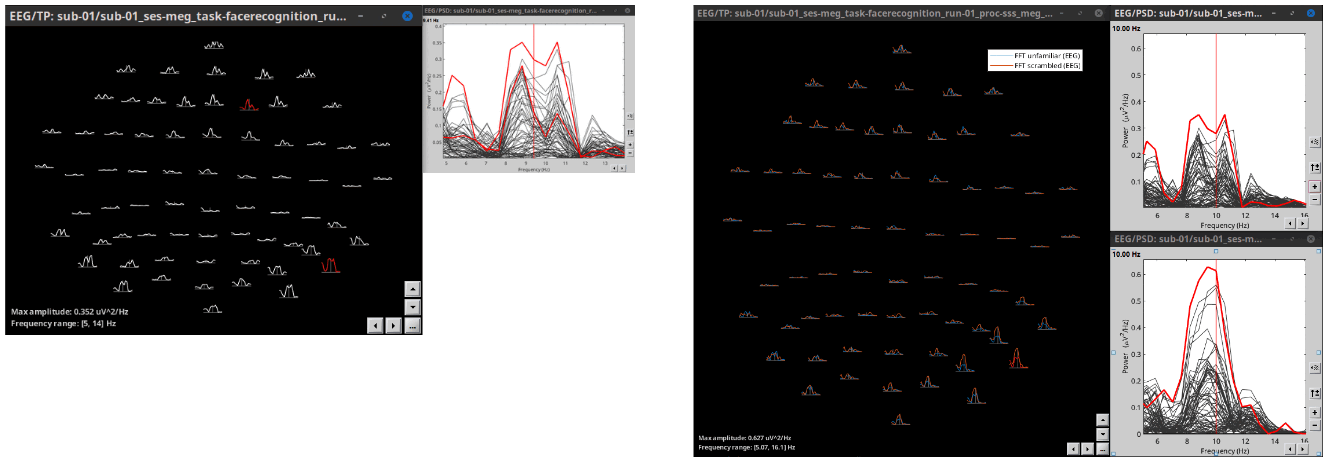
Processes
Continuous 'raw' data support to process Artifacts > Detect bad channels: peak-to-peak'. 🔗
Software distribution
Update automatic detection of Matlab RunTime path in Linux and macOS. 🔗
March 2024
SEEG/ECOG
Improved localization: Update of iEEG tab. Localization of SEEG/ECOG contacts can be performed in 3D figures assisted by iso-surfaces from thresholded CT volumes. This facilitates the localization of iEEG contacts. 🔗
Events
Anatomy
- Prevent modification of template scout atlases
Scouts: Add 'Duplicate' operation for scouts. RMS is available as scout function (and cluster function) 🔗
Inverse
Prevent errors by explicitly checking that bad channels in noise covariance are also bad channels in recordings 🔗
Input / output
Update: support for ITAB 🔗
Update: support for Localite .csv files 🔗
Bugfix: coordinate system on for Curry .pom channel files 🔗
February 2024
Database
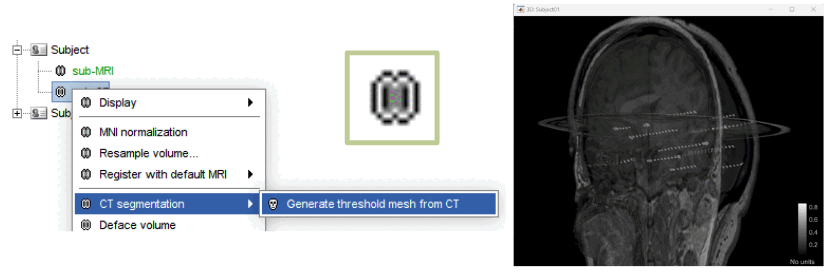
CT volumes: Integration of CT volumes in Brainstorm database. CT volumes are their own anatomy type now, this allows popup menus and options different from the MRI ones. E.g. CT segmentation to generate an iso-surface from thresholded CT volumes. 🔗
Portability of raw files Links to raw files are automatically updated, when mounting point or disk letter change, e.g. database in HDD open in different computers. 🔗
Anatomy
Scouts Add option to compute the intersection among a set of scouts 🔗
Warp Bugfix: Default surfaces after warping default anatomy 🔗
Input / output
Synchronize raw signals: Allow to synchronize raw signals using common events 🔗
January 2024
Input / output
Import channel-wise events from BrainVision BrainAmp
December 2023
Software distribution
Basic support Apple silicon (OsType 'mac64arm')
Input / output
Add process Export to file to export or or multple data, sources, timefreq and matrix files.
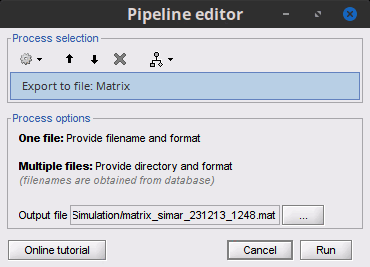
November 2023
CT to MRI co-registration
The method CT2MRI was added as an option to co-register CT and MRI volumes. CT2MRI plugin and BrainSuite are required.
- CT2MRI co-registration offers the option of skull stripping for the registered CT.
Plugins
- Remove EASYH5 and JSNIRF code from Brainstorm, add them as plugins
Input / output
- Always Export EDF+ with UTF-8 encoding (bugfix)
Support export to .xlsx files for Matlab >= R2019a
- Export EEG data as Brainsight format
October 2023
iEEG electrode models
- Save, Load, Export and Import electrode models
Software distribution
- Compilation with Matlab 2023a/2023b
September 2023
Anatomy
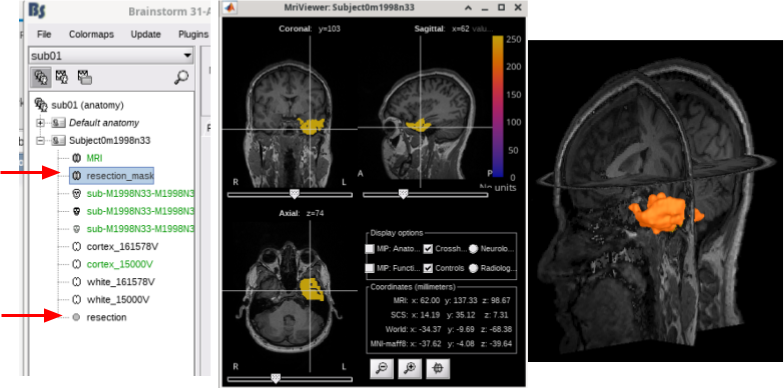
Import resection mask from BrainSuite SVReg. See tutorial
Input / output
Export raw data as FieldTrip structure
August 2023
Brainstorm ChatBot
Users can now interact with the ChatBrainstorm3 a chatbot based on the ChatGPT 3.5. ChatBrainstorm3 is fine tuned using ~11M characters, from all the Brainstorm website pages, from all the forum's discussions and from the Brainstorm GitHub repo.
This ChatBot is available both on the Brainstorm webpages and on the forum. You can find it on the bottom right on your screen. Click on the 🗨️ icon and start your discussion with it ![]()
Interface
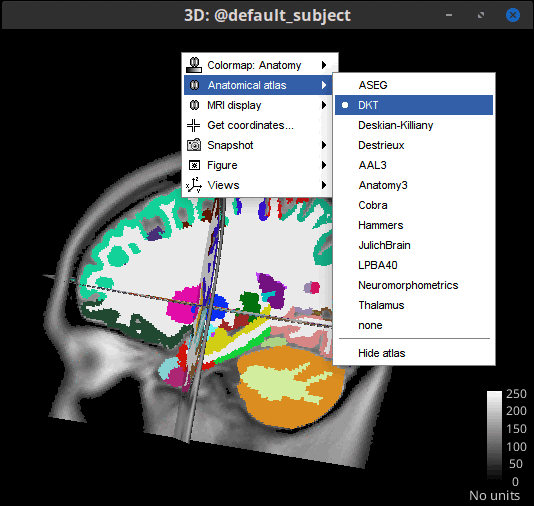
- Display anatomical atlases on 3D orthogonal slices view.
Connectivity
- Improve across-trials average of phase metrics
- Reorganize connectivity metrics
July 2023
Anatomy
Scouts: Export scouts as FreeSurfer annotations
Electrophysiology
Tunning curves: Bugfix at their computation
June 2023
PCA
New PCA tutorial page.
- Major fixes and improvements for Scouts and Flattening unconstrained sources
- Fix 1st PC sign inconsistency across epochs and conditions
- PCA for scouts rescaled. "% power kept" fixed and saved in history
Connectivity
- Connectivity: New option to flatten unconstrained sources with PCA
- Connectivity: Fix scouts on volume and mixed models
May 2023
Anatomy
Support importing FreeSurfer wfiles as textures
Plugins
Blackrock: NPMK library update to be fetched from its GitHub HEAD
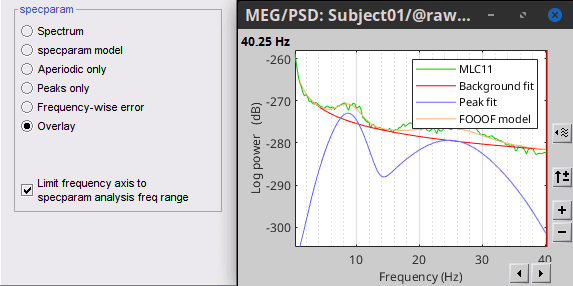
FOOOF
Display Allow adjusting the frequency axis to the frequency range defined by specparam model.
April 2023
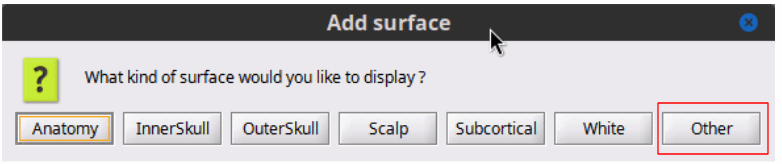
Anatomy
3D Figure Allow to add 'Other' surfaces
NIRS
- Project channel files between subjects
March 2023
Temporary files
The temporary files in Brainstorm are saved by default in the user home folder ($HOME/.brainstorm/tmp). Previsouly, all the files were always with the same name, directly in the tmp folder. Now, each process saving temporary files creates a sub-folder with a timestamp. This avoids conflicts between multiple Brainstorm sessions and enables parallel computing for some processes (more information).
For example, when epoching MEG/EEG recordings, Brainstorm would first create a temporary folder import_yymmdd_hhmmss, store all the epochs in it, then move them to the database when the epoching process is completed. The name of the temporary folder indicates its creation time (year/month/day_hour_minutes_seconds). At the end of the process, all the temporary files should be deleted automatically. If the process crashes or is killed before it returns, the files remain in the tmp folder and need to be explicitly deleted, either with a function call or from the Brainstorm preferences.
When starting the Brainstorm interface, users are now prompted to delete files in the temporary folder. They can be safely deleted if they result from a previous unfinished process, but must be kept if in use from a different instance of Matlab/Brainstorm.
Database
Events: Update storage of events structures: The fields "channels" and "notes" are now allowed to be empty. This can save Gb of storage and long minutes of loading when working with recordings including hundreds of thousands of events (e.g. single neuron spiking activity).
Channel clusters: The clusters of channels are now saved in the channel files, while there were previously available only temporarily in the graphic interface. The Cluster tab is now opened automatically when loading recordings for which clusters have been defined. Associated function db_set_clusters.m and process Import clusters of channels.
February 2023
FEM anatomy
FEM mesh generation with SimNIBS4/CHARM
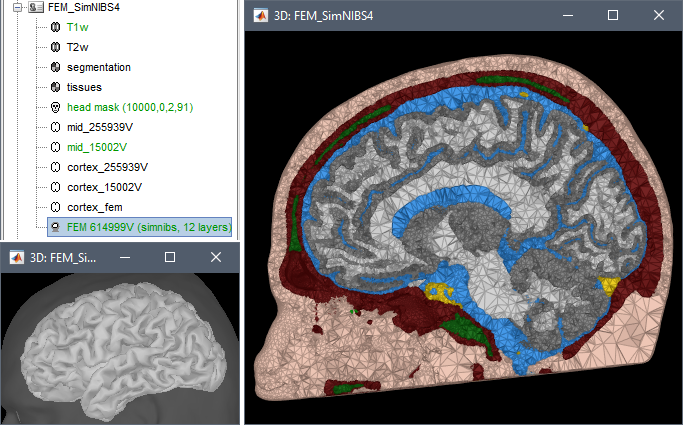
Merge/rename/delete FEM layers: new popup menu
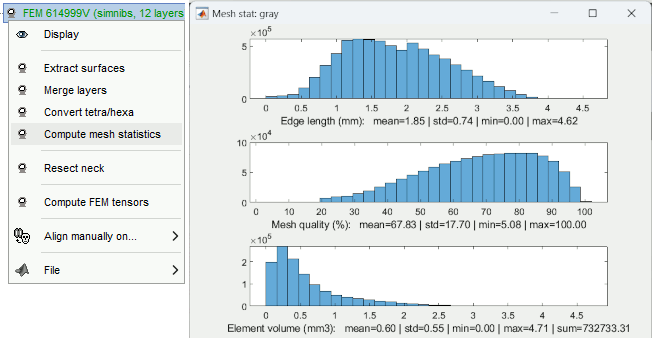
Compute FEM mesh statistics
Statistics
Non-parametric tests: Added FieldTrip threshold-free cluster enhancement (TFCE)
Fixed error in data covariance computation: The baseline was not removed when the baseline time window was not included in data time window. See issue #602.
Input / output
- Export matrix files as EDF+
January 2023
Anatomy
New MNI template: ICBM152 2023
- Project scouts between hemispheres using the anatomy template
- Removed selection of fiducials when setting anatomy template
Software distribution
Brainstorm is now compiled with Matlab 2022b: Installation instructions
The compilation is now supported on Linux and MacOS: How to compile
Interface
- Display scouts on source-level PSD as PSD figures
November 2022
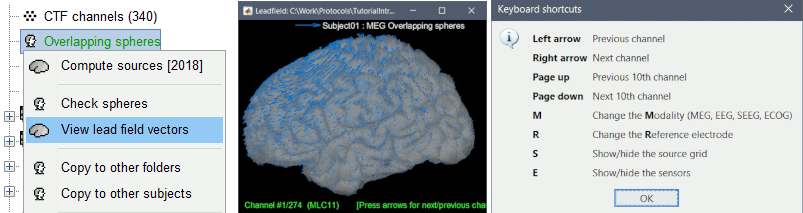
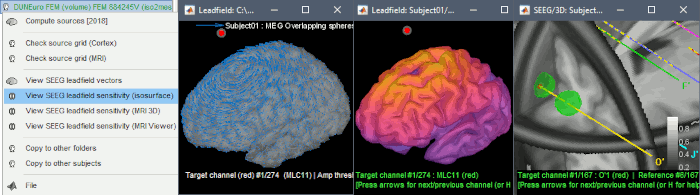
Leadfield sensitivity
The leadfield for each MEG sensor, or each pair of EEG/SEEG electrode/reference, can now be displayed as vectors, or as sensitivity maps: on the cortex surface, in the the MRI volume, or as an isosurface at a specific sensitivity value.
BIDS import
Support for ACPC and CapTrak coordinate systems
- Support for NIRS extension
Support for AssociatedEmptyRoom + automatic computation of noise covariance
MNI normalization
New MNI template: ICBM152 2022
- EEG: Project channel files between subjects
- EEG: When using menu menu "Add EEG positions", convert from MNI space to subject space
- Project dipoles files between subjects
September 2022
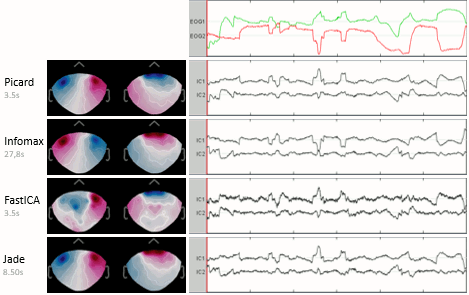
ICA analysis
The interface of the ICA process has been improved and two new popular and efficient methods have been added: PICARD and FastICA. Picard is now the default option for ICA analysis in Brainstorm. More information.
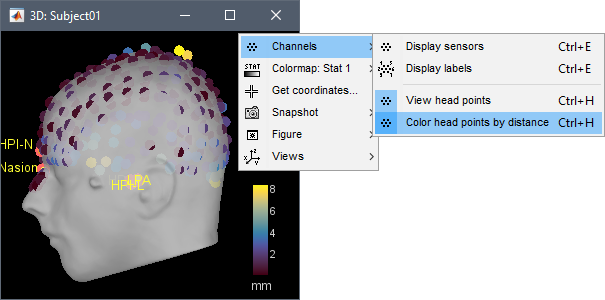
Coregistration
Color-coded distance: When checking visually the registration between the sensors and the anatomy, a new option allows to color the digitized head points based on the distance to the scalp surface. This can help identify and fix manually coregistration issues. More information.
Fiducials on head surface: It is now possible to position the anatomical fiducials (NAS, LPA, RPA) on the surface instead of the slices in the MRI Viewer. More information.
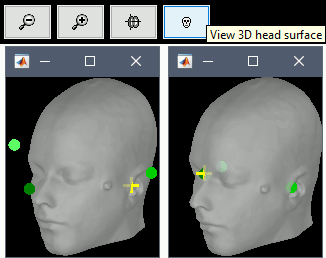
Input / output
- Support for BCI2000 .dat recordings
Support for BIOPAC AcqKnowledge .acq recordings
- Support for XDF EEG recordings
- Updated Nicolet EEG reader
June 2022
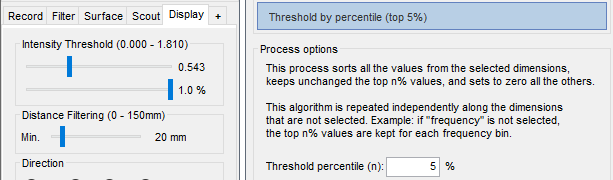
Connectivity graph
It is now possible to apply a threshold based on the percentile of the distribution of the connectivity values. This option is available both interactively and from a new process "Threshold by percentile". More information.
Anatomy
The contralateral registration from FreeSurfer is now supported in Brainstorm and allows the projection of scouts between hemispheres. More information.
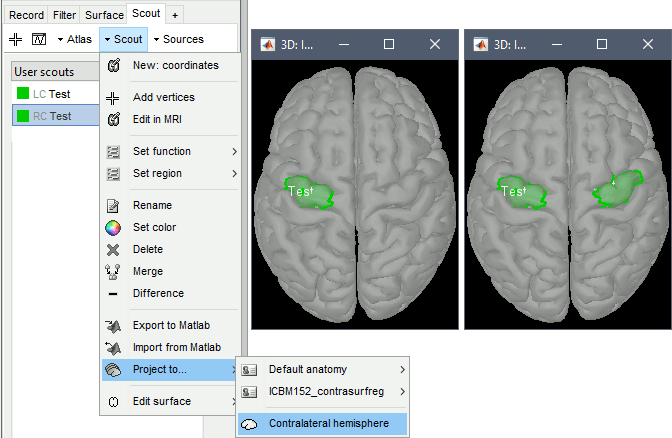
BIDS
Import: Added support for IntendedFor in _coordsystem.json
- Import: Added support for anatomical landmarks (_t1w.json and _coordsystem.json)
- Export _electrodes.tsv, with the choice of the reference MRI
April 2022
Electrophysiology
Major update of the electrophysiology toolbox: new features, many bug fixes, and improved documentation. Explore the new tutorials.
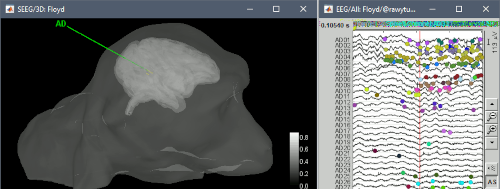
Input / output
- Support for Neuroelectrics EEG with EEGLAB plugin (.easy, .nedf)
- Updated Plexon reader
Export events in AnyWave .mrk format
- Import channel from ASCII files now expect mm instead of cm
February 2022
Connectivity
New tutorial: Functional connectivity (methods)
New tutorial: Corticomuscular coherence (practice with FieldTrip tutorial dataset)
New measures: wPLI and ciPLV
Updated process: Simulate AR signals
- Computation: Removed the p-values thresholding from all connectivity metrics
- Removed JAVA3D/JOGL support and the older connectivity graph display
Anatomy
Extract head with FSL/BET: Right-click on MRI > MRI segmentation.
FreeSurfer: Added support for InfantFS / infant_recon_all
January 2022
SPRiNT: Time-resolved spectral parameterization
New tutorial and process for decomposing and parameterizing spectral components over time:
Spectral Parameterization Resolved in Time (SPRiNT)
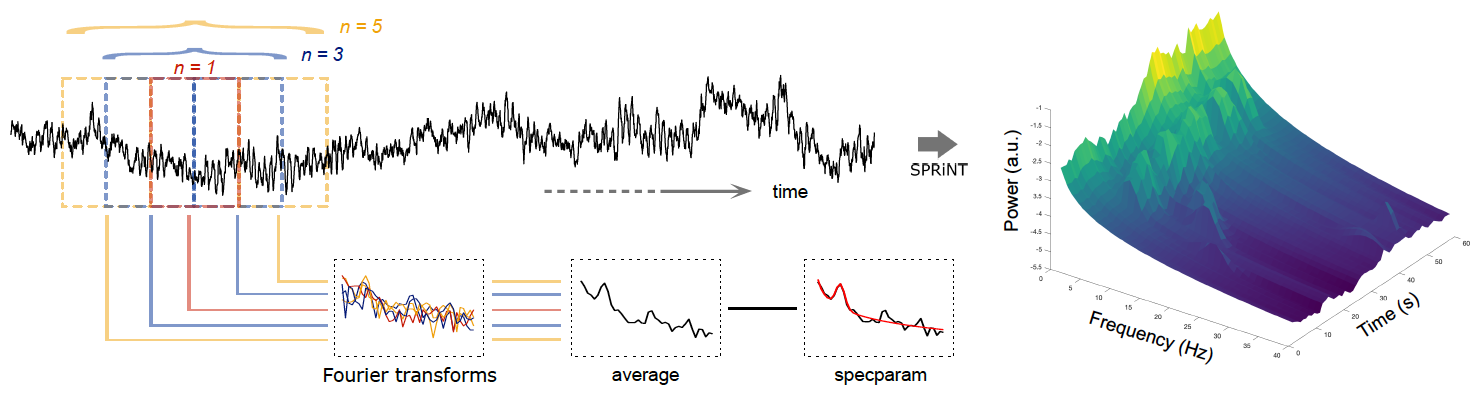
Plugins
SimMEEG: New video tutorials.
New menu: Update > Reproducibility > Export software environment:
Archives the software environment in one zip file, including installed plugins and templates.Track software versions: Save GitHub commit SHA and installation date for each plugin.
Statistics
New process: Conjunction inference
December 2021
Input / output
- Support for Tobii Pro Glasses export .tsv
Support for FieldTrip trialinfo field
- Updated York Instrument MEGSCAN reader (.meghdf5)
November 2021
MRI segmentation
Brainstorm is now fully interfaced with new programs for MRI processing and segmentation:
MRI segmentation with FastSurfer (T1 only)
MRI segmentation with FreeSurfer (T1+T2)
MRI segmentation with BrainSuite (T1 only)
Import BIDS: Add support for CAT12, BrainSuite and BrainVISA
New plugins
MIA: Group statistics on SEEG data, see the website.
DeriveLFP: Part of the e-phys toolbox, available in the compiled distribution.
Download statistics are now accessible for each plugin. See tutorial: Plugins.
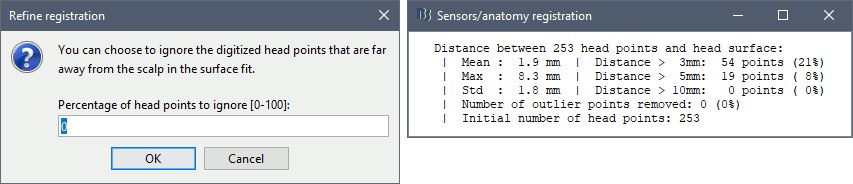
Automatic registration
The automatic MEG/sensors registration using digitized head points has been improved. It now includes an new parameter to exclude outlier digitized points, and a report of the distance between the digitized head points and the scalp surface. See tutorial: MRI registration.
Send reports by email
When running some long computation on a distant server, you can now receive an email when the processing is over. See tutorial: Scripting.
October 2021
Major bug fix: PCA
The computation of the first mode of the PCA from multiple time series has been fixed. It now restores the mean of the signal after the computation of the PCA. See GitHub commit. The output of the following functions and processes are affected by this modification:
- Interactive display of the scouts time series, with scout function PCA.
Process: Extract > Scout time series, with scout function PCA.
Process: Sources > Unconstrained to flat maps, option PCA.
Brainstorm compilation
We updated the deployment and compilation scripts of Brainstorm. Now everybody with the Matlab Compiler toolbox can recompile the Brainstorm standalone package, including additional personal scripts of modified versions of the code. See tutorial: Scripting.
September 2021
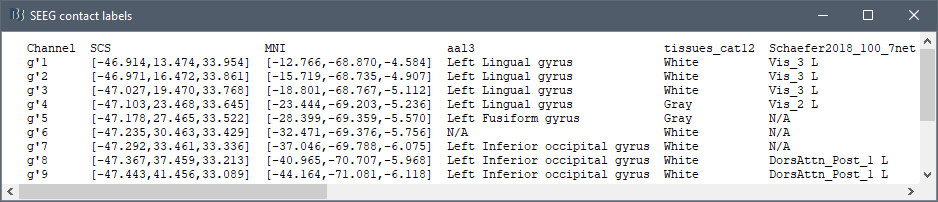
iEEG anatomical labels
New automatic SEEG/ECOG contact labelling from volume and surface parcellations. See tutorials: SEEG epileptogenicity maps and ECoG+sEEG epilepsy.
August 2021
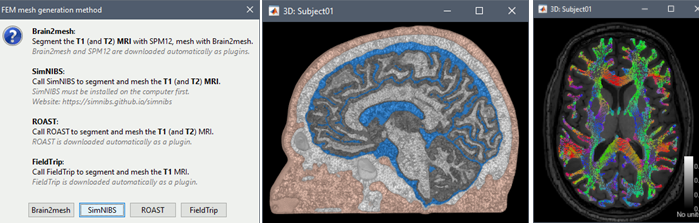
FEM tutorials
The FEM tools in Brainstorm are now documented in new tutorials: FEM mesh generation, FEM tensors estimation, FEM median nerve example, Realistic head model: FEM with DUNEuro.
July 2021
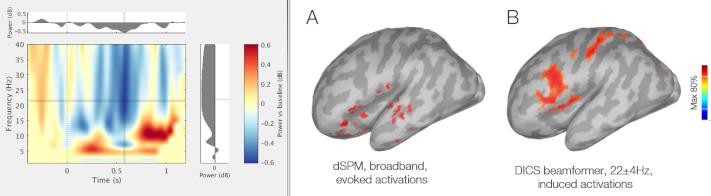
Coherence
The computation of the coherence has been revisited: new formula for the Imaginary Coherence, online averaging of the cross-spectra, estimator window length as a duration. See GitHub PR.

Inverse: FieldTrip DICS
The FieldTrip DICS method is now available as a process function, thanks to the contribution of Vahab Youssof Zadeh: GitHub PR, Tutorial.
June 2021
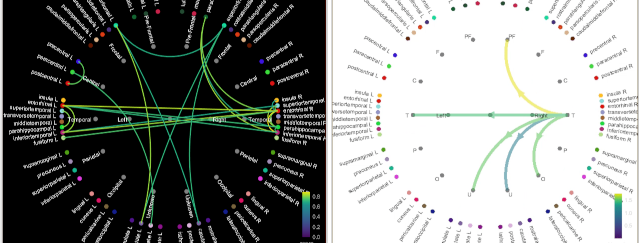
Connectivity graphs
The display of connectivity graphs has been revisited. This new version is now fully written in native Matlab code, not dependent anymore on the JOGL library. See the new tutorial: Connectivity graphs, GitHub PR.
May 2021
Anatomy
- FEM: Reconstruct triangular tissue envelopes based on FEM tetrahedral meshes
- Use imported volume parcellations as volume scouts
- Updated CAT12 to version 12.8 (r1825)
- Fixed O'Reilly infant templates
Input / output
- Support for Micromed .EVT
- Added Blackrock NPMK libary as plugin
- Added support for Axion AxIS recordings (.raw)
March 2021
Compiled distribution
A fresh compiled distribution of Brainstorm is now available, including many of the new Brainstorm plugins. Without a MATLAB license, you can now use functions from plugins: SPM12, FieldTrip, Iso2mesh, Brain2mesh, OpenMEEG, DUNEuro and libSVM. Because the size of the full pacakge binary + sources increased a lot, we now recommend to download either the src or the bin package from the Download page.
February 2021
Plugin manager
A new plugin manager now allows an easy management of all the software packages related with Brainstorm. See the Plugins tutorial.
Input / output
Support for ADInstruments LabChart EEG (.adicht)
- Updated support for Neurodata Without Borders (NWB)
December 2020
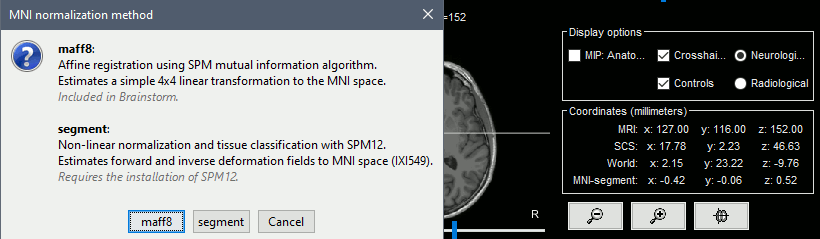
MNI normalization
- MNI non-linear normalization and tissue segmentation with SPM12 Segment
- Handling non-linear MNI registrations in the interface (SPM deformation fields y/iy)
Import and reslices MNI atlases using linear/nonlinear normalization
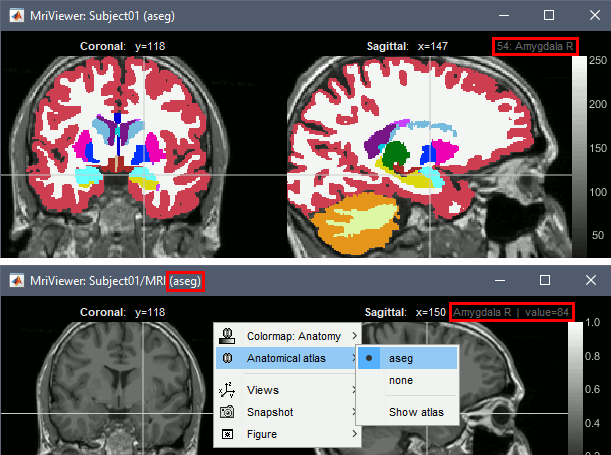
Volume parcellations
Support for volume parcellations in the database and the MRI viewer (volatlas)
Import all volume parcellations from FreeSurfer, CAT, BrainVISA, BrainSuite
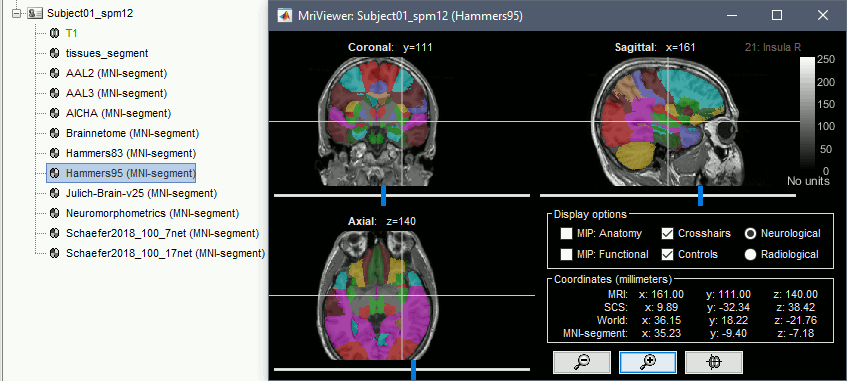
Standard MNI atlases available for download, applicable both with the linear and the non-linear MNI normalization: AAL2, AAL3, AICHA, Brainnetome, Hammers, Neuromorphometrics, Julich-Brain, Schaefer2018.
Anatomy
- Updated CAT12 to version r1733
FreeSurfer import now loads all the .annot files in /label
- Standard EEG caps included with new infant templates
New automated import, with no user interaction: using 15000 vertices, MNI fiducials
November 2020
MNE-Python
- More generic Python initialization and environment definition, using pyenv when available
- New integration with Python and MNE-Python based on CPython
SSS/tSSS process = Wrapper for mne.preprocessing.maxwell_filter
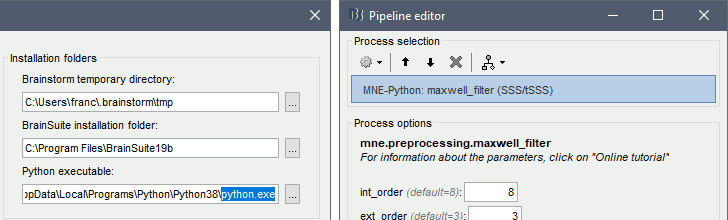
Extensions
Compiled version: Now includes NIRSTORM, Brain2Mesh and Iso2Mesh
- DUNEuro FEM: Added SEEG/ECOG support
- E-phys toolbox: Phase locking circular plots
October 2020
Major change: Power spectrum computation (PSD)
- New option in the PSD (Welch) process for scaling of the power spectrum. The new default displays physical units: (signal units)^2/Hz, whereas the previous scaling (still available) was rather proportional to power "per frequency bin", and depended on the window size. A third option gives normalized frequencies (0 to 2pi).
- Y scale can now be set to log in power or magnitude settings, from the right click or display settings menus, maintaining the physical units instead of dB.
September 2020
Simulations
New tutorial and processes for simulating signals, and simulating EEG/MEG recordings from dipoles. See the Simulations tutorial.
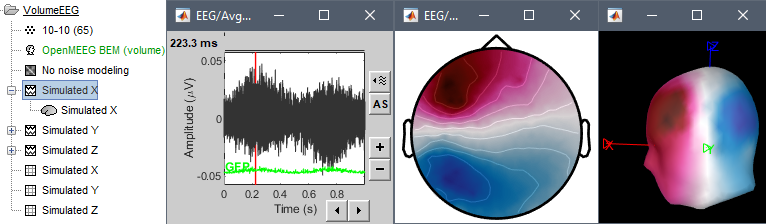
Spectrum analysis
New tutorial and process for modeling and analyzing the power spectra:
Fitting oscillations and one-over-f (FOOOF)
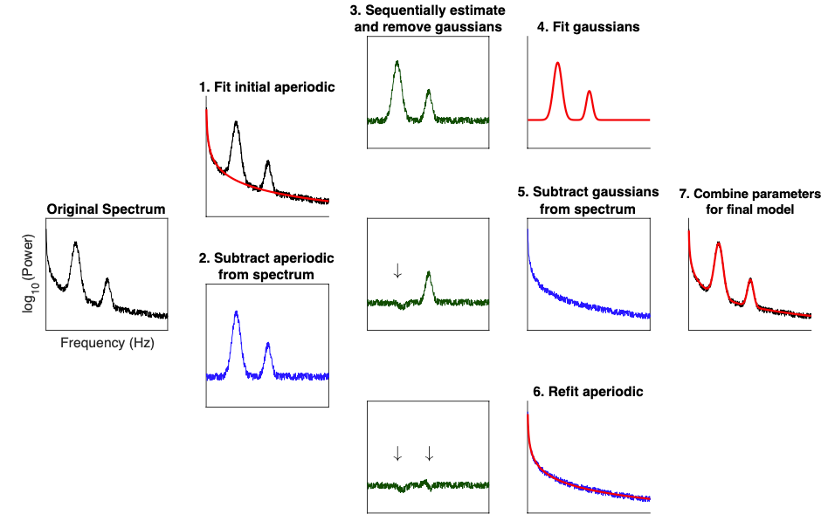
August 2020
2D surface projection
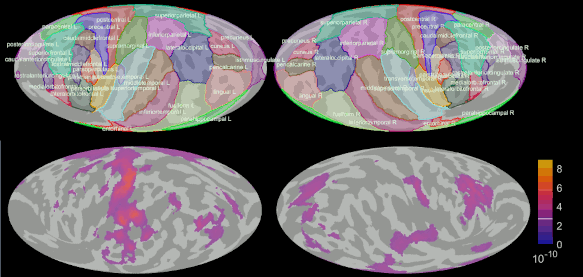
Display cortical surfaces and source maps as flat 2D maps using the Mollweide projection,
The FreeSurfer registered spheres can now be used to compute a flat 2D projection of the cortex surface using the Mollweide projection, as described in this article: (Kang et al. 2019) Hemispherically-Unified Surface Maps of Human Cerebral Cortex: Reliability and Hemispheric Asymmetries. For more information, see the FreeSurfer tutorial.
Connectivity
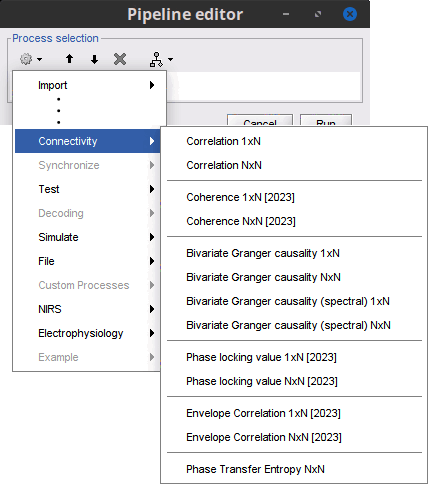
New process Connectivity > Envelope correlation. It will be document as part of the new Connectivity tutorial, which is still under construction for the moment.
Input / output
- Support for Spike2 .smrx files
- Support for Open Ephys flat binary file (.dat/.oebin)
July 2020
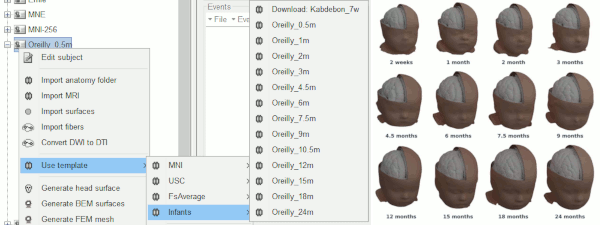
Infant anatomy templates
Thanks to Christian O'Reilly, we can now distribute 13 anatomical models for infants between zero and 24 months of age (O'Reilly et al. 2020), available as FreeSurfer templates.
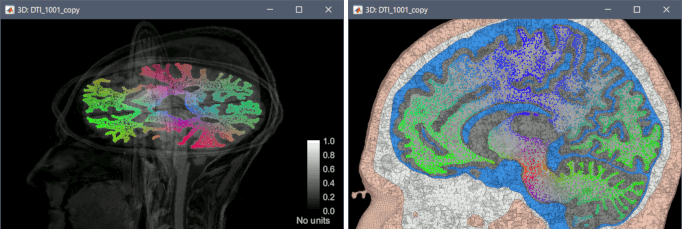
FEM: Use of conductivity tensors
Computation of DTI tensors from DWI images with BrainSuite, display on tetrahedral meshes.
May 2020
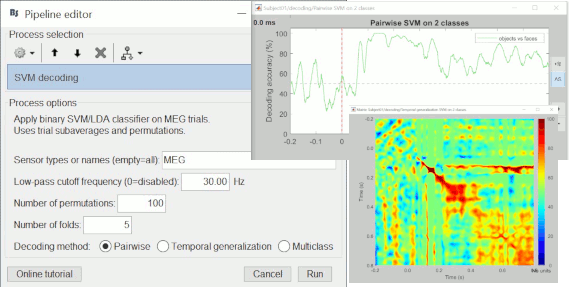
Machine learning / Decoding
The MIT MEG lab presents new processes for MEG/EEG decoding (a type of multivariate pattern analysis / MVPA) using support vector machines (SVM): Machine learning: Decoding / MVPA.
Input / output
Support for fNIRS format .snirf
- Support for The Virtual Brain .h5 EEG recordings (TVB)
April 2020
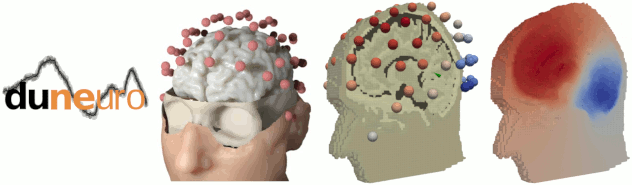
FEM forward modelling with DUNEuro
The state-of-the-art library for FEM forward modelling DUNEuro is now interfaced with Brainstorm. It is based on the DUNE library and its main features include solving the EEG and MEG forward problem and providing simulations for brain stimulation. See the tutorial: Realistic head model: FEM with DUNEuro.
View leadfield vectors
Select one or multiple forward models in the database explorer > View leadfield vectors. Press H for help, use the arrows to navigate between the different sensors.
March 2020
Anatomy
- CAT12: Updated to 12.7 and added import of extra maps (gyrification, sulcal depth)
BEM surface generation with FieldTrip / ft_volumesegment:
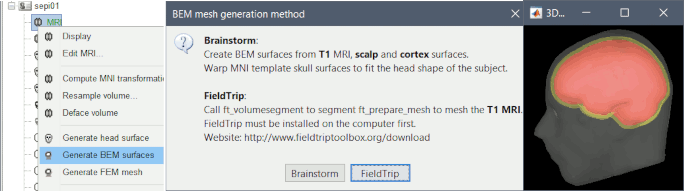
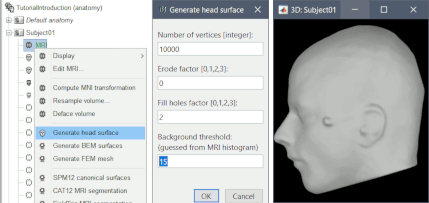
Use custom background threshold for head surface generation: allows fixing some cases where the automatic analysis of the MRI histogram fails.
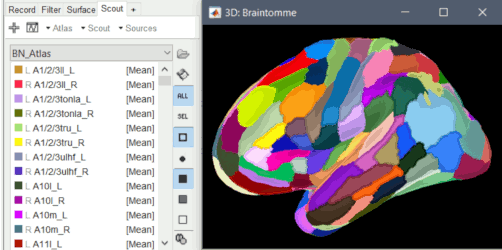
Updated FSAverave template, including the new Braintomme atlas.
Software distribution
- Upgrade compiled version of Brainstorm to Matlab 2020a
Preparation Java/Swing-free Matlab: deprecated JavaFrame and javacomponent
January 2020
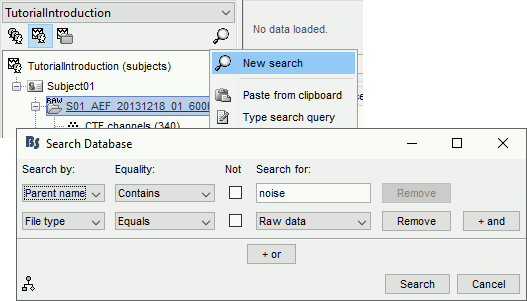
Database search interface
The database explorer now offers advanced and scriptable search tools: see tutorial.
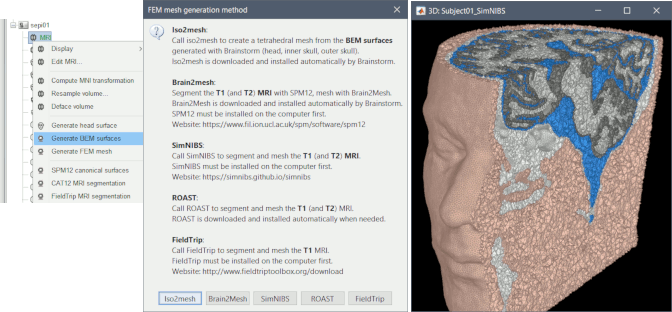
3D mesh generation for FEM
Mesh generation using iso2mesh, Brain2mesh, SimNIBS/headreco, ROAST and FieldTrip.
Optimized 3D tetrahedral mesh display
Input / output
- Support for Wearable Sensing CSV files
- Support for EmotivPRO EDF recordings
- Support for SimNIBS meshes
- Support for time-varying MRI and DWI volumes
- Support for Localite digitizer (.csv)
- Anonymize FIF files
October 2019
Recordings
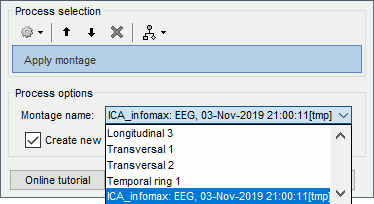
Montage editor: New option to filter each signal in a specific frequency band: see tutorial
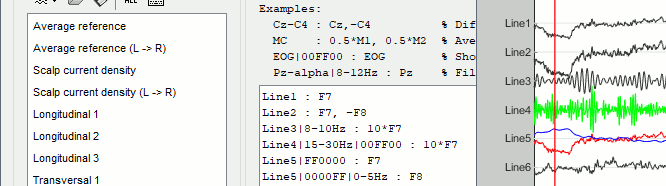
Montage editor: Display and apply SSP/ICA projectors as montages, to export IC time series
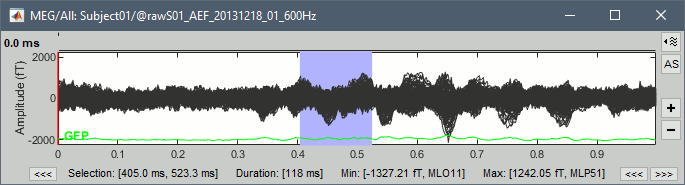
Zoom on selected time/frequency segments with shift+click (time series and spectrum)
Interface
- New colormap: Turbo, Google's JET alternative
- Database: Support for regular expressions when importing events
September 2019
Anatomy
- Import volume atlases as scouts without overlap between ROIs
- Storage and display of FEM meshes
Computation of FEM meshes with ROAST and iso2mesh: see GitHub discussion
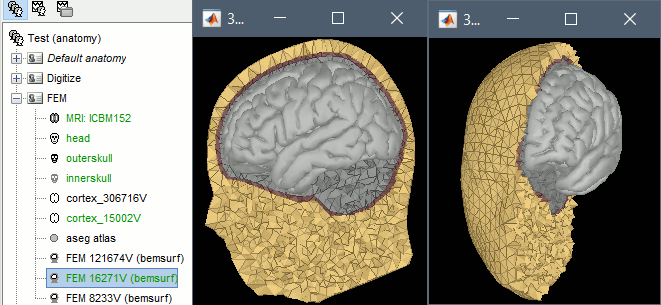
Input / output
Conversion functions from/to MNE-Python objects (Raw, Epoched, Evoked)
- Support for Curry epoched .dat files
- Report viewer can now export snapshots as .PNG images
- Process box: support pasting plain list of files in various formats
July 2019
MRI segmentation with CAT12
CAT is a SPM12 toolbox that is now fully interfaced with Brainstorm. It can replace efficiently FreeSurfer for generating the cortical surface from any T1 MRI. It runs on any OS in about 1 hour, instead of the typical 24hr FreeSurfer recon-all processing. The surfaces are registered to the templates with the FreeSurfer spheres, and include many anatomical and functional atlases.
See new tutorial: T1-MRI Segmentation with SPM12 / CAT12
Averaging: API modification
A new variable Leff has been added to all the files structures, to store the effective number of averages, as defined in the MNE manual. It replaces the nAvg value for weighting files based on different numbers of trials when computing an average or a noise covariance. This modification may change the output when computing weighted averages of subject-level averages.
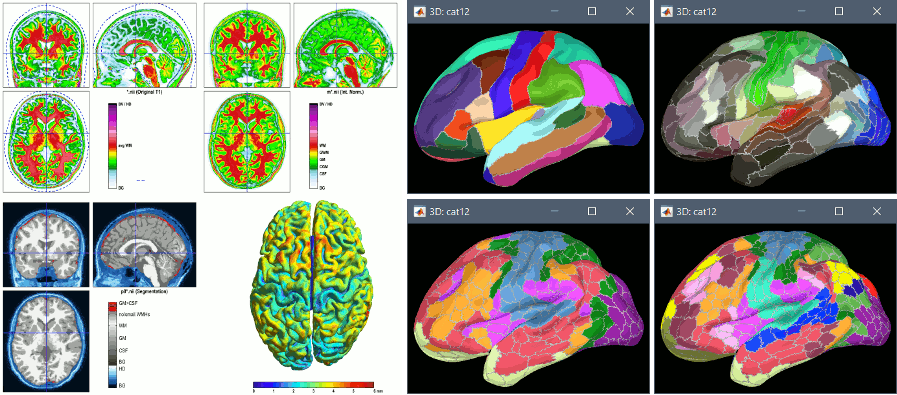
dSPM averages: Scaling with number of averages
Since July 2018, the dSPM maps have been saved without any scaling. As a consequence, the values obtained for averages (ERP/ERF) are much lower than expected. To obtain correctly scaled values, one can now run the process Sources > Scale averaged dSPM.
June 2019
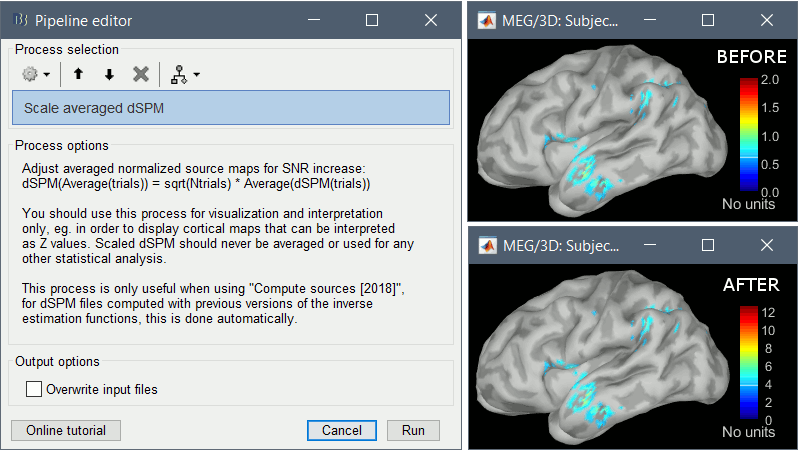
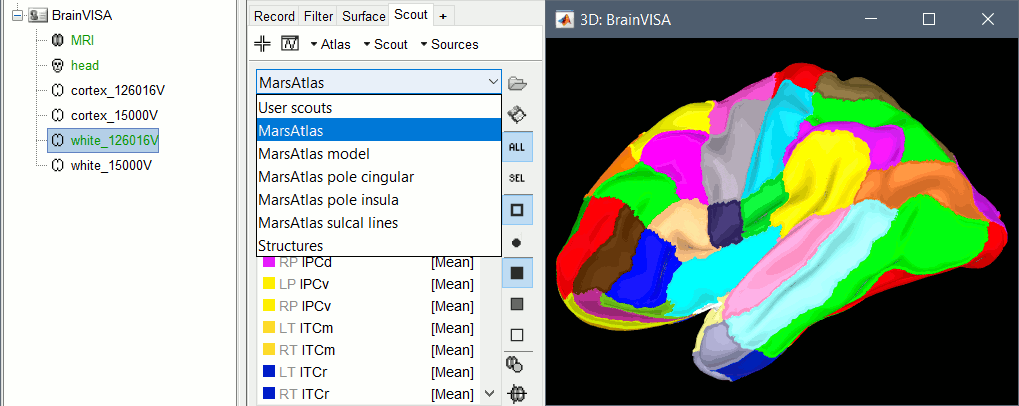
BrainVISA: MarsAtlas parcellation
The default analysis pipeline in BrainVISA implements an automatic parcellation of the cortical surface in anatomical regions. The MarsAtlas model is now imported automatically by Brainstorm.
Input / output
- Support for NWB ECOG (recordings + surfaces)
- Support for MNE-FIF epoched files
- Support for ANT EEProbe .avr files
- Support for Intan RHS+RHD files
- Improved support for discontinuous EDF files (EDF+D)
May 2019
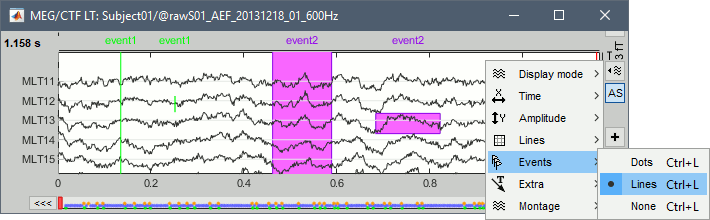
Events display
Three display modes are now available for the event markers: dots, lines or hidden. Select the corresponding entry in the new display options menu, or press CTRL+L multiple times. Selected event groups can be hidden from the display with the new shortcut CTRL+H.
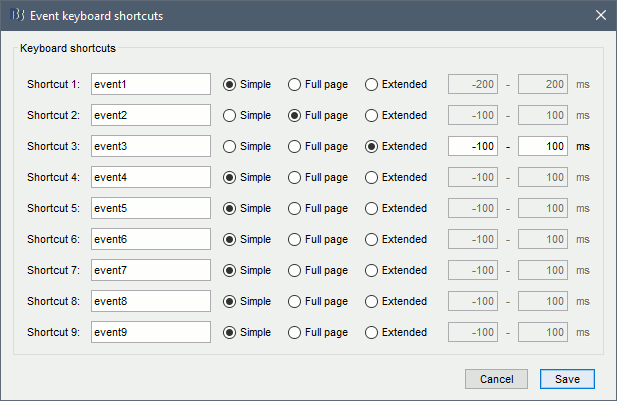
Events shortcuts for sleep staging
New shortcuts have been added to the signals viewer to allow tagging entire pages of recordings: F6 and Shift+F6 scroll in the file without overlap, and the option Full page in the custom shortcuts tags a page and moves to the next one.
Decoding processes for more than 2 conditions
It is now possible to run the decoding processes with more than 2 conditions by grouping your trials in condition folders, and dragging them in the process box rather than the process2 box.
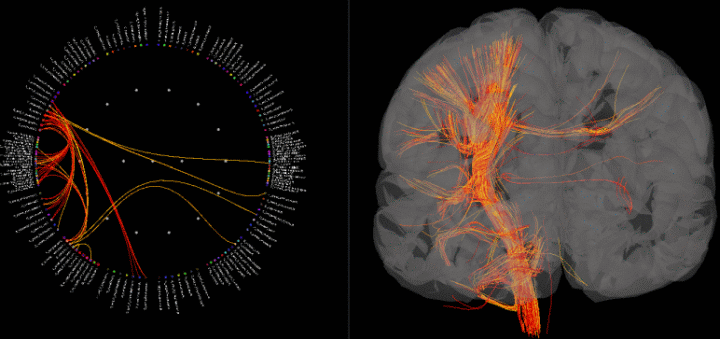
Fiber viewer for connectivity visualization
A new surface type was added to Brainstorm: fibers. One can use this to visualize connectivity results by displaying fibers connecting regions of interest and thresholding by connectivity value.
April 2019
New default colormaps
The colormaps "jet" or "rbw", used by default in Brainstorm for many years, lack important attributes of good colormaps: they dont’t have linear lightness and are not perceptually uniform. This can either cause details in the visualization to be hidden or create features that don’t exist in the underlying data, which results in a distortion of the perceived pattern. New colormaps were added to better represent the data: the three colormaps below were created, and the viridis and magma were taken from mpl colormaps. For more information about these new colormaps: colormap_optimization.pdf

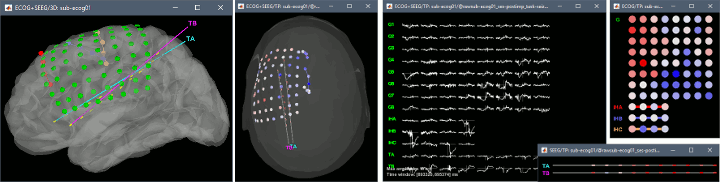
New ECoG/sEEG tutorial
The tutorial ECoG+sEEG epilepsy illustrates all the new features related with SEEG/ECOG: coregistration of MRI and CT scans, placement of depth electrodes in the post-implantation scans, SEEG bipolar montages, display and processing of intracranial recordings, computation of epileptogenicity maps.
Programming: Field "samples" removed
Important for users who edit Brainstorm structures manually:
The field samples was removed from the events structure and the sFile.prop structure. This field was redundant with the field "times", with risks of misinterpretation and divergence between the two. The topic is discussed in this pull request. If you need the corresponding samples information, it can be computed from the field times: samples = round(sFile.event(i).times .* sFile.prop.sfreq).
Additionnally, the fields channels and notes were added to the events structure. More details.
March 2019
Anatomy
Deface MRI with FreeSurfer or SPM.
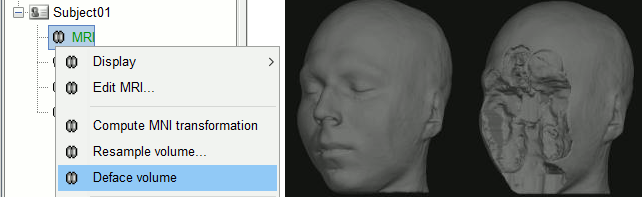
World coordinates now available in the MRI viewer. More details.

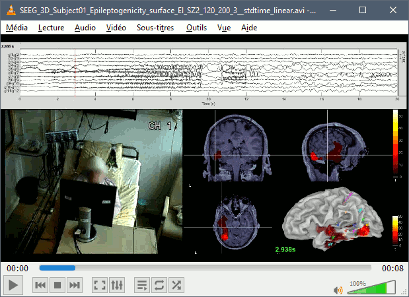
Video-EEG
Support for synchronized EEG-video.
Review of synchronized EEG-video with VLC ActiveX plugin.
EEG and iEEG
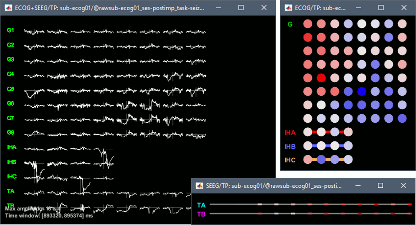
- New default montage: EEG Scalp Current Density (SCD)
New display modes for ECoG and sEEG: 2DLayout and 2DElectrode
February 2019
Processes
New filters: The default frequency filters (band-pass, band-stop, notch) were modified to follow the new Shahabi/Leahy 2019 specifications.
New process: Standardize > Interpolate time
Input / output
- Support for York Instruments MEGSCAN recordings (.hdf5)
- Support for Tucker Davis Technologies recordings (.tdt)
Software distribution
Compiled distribution now includes all SPM and FieldTrip dependencies
Compiled distribution can execute custom scripts (GUI and command line)
- New compiled distribution: Matlab 2019a
- Updating Brainstorm now downloads only the scripts (skips the compiled version)
January 2019
Input / output
- Support for Plexon 2 recordings (.pl2)
- Support for York Instruments MEGSCAN recordings (.hdf5)
- BIDS import/export: Support for EEG and iEEG extensions
Export EEG to BrainVision BrainAmp format (.eeg) for iEEG-BIDS
- Export EEG/SEEG electrode positions in MNI or world coordinates
Bug fix: Fixed coil orientations when reading CTF .ds folders including a .pos file (McGIll MEG system): https://neuroimage.usc.edu/brainstorm/Tutorials/CtfCoilOrientBugFix
Interface
- Added colormap ovun (now default for stat2 files)
- Stat: Colormaps adapted to t-value threshold
- Anatomy: Handle better world coordinates (cs_convert, import/export of channel files)
Management of multiple screens for Matlab>2015b
November 2018
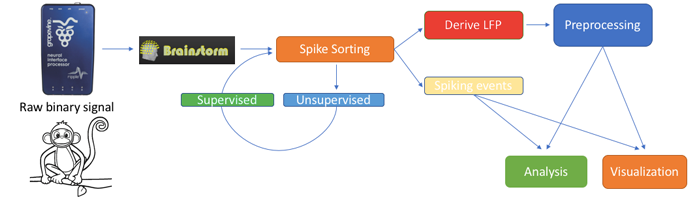
New Invasive Neurophysiology toolbox
A new toolbox featuring basic electrophysiology tools is now included in Brainstorm, and documented in a brand new series of tutorial pages on Invasive Neurophysiology. The toolbox includes both unsupervised and supervised neuron spike sorting, extraction of local field potentials (LFPs), spike field coherence, spike triggered average, noise correlation, and basic visualizations such as tuning curves and raster plots.
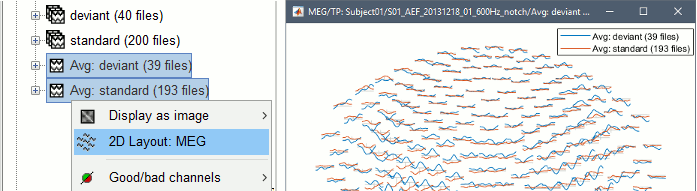
Display multiple files with 2DLayout
Compare easily averages between experimental conditions: Select multiple files in the database explorer, right-click on any of them > 2DLayout.
September 2018
Input / output
- Support for EGI-Philips MFF recordings
- BIDS export process
- Improved download speed
July 2018
Updated dSPM 2018
Major modification in "Compute sources [2018]":
dSPM values are no longer scaled by number of trials. See source tutorial.
Interface
Headless mode: Brainstorm now runs without any rendering capability, allowing the execution of scripts on distant servers and computation clusters that do not have any display attached to them. Adding snapshots to the execution reports is still possible. Start your script with "brainstorm server".
Custom montage shortcuts: In the montage editor, shortcuts can now be associated with any montage in the list.
Stat duration threshold: Added "Minimum duration" threshold in the stat tab.
New montage: "Average reference (L -> R)" for 10-20 EEG caps
Anatomy
BrainSuite: Now reading subcortical structures from the SVReg atlas. See details.
FreeSurfer: Added labels for volume atlases: aparc+aseg, aparc.a2009s, aparc.DKTatlas
- OpenMEEG: Switched to version 2.4, which should increase stability and reliablity.
April 2018
Input / output
- Support for g.tec HDF5 files
- Support for Curry 6-7-8 EEG recordings
- Support for Muse .csv EEG recordings
- Montage can be loaded and saved as CSV files
ICA cleaning
- Added option: Sort components based on correlation with a channel (EOG, ECG...)
Changed reconstruction method: (Winv_good * W_good) => (eye() - Winv_bad * W_bad)
March 2018
SEEG/ECOG
- Updated tools for ECOG (new display options, many bug fixes)
- New 2DLayout display option
- Allow simultaneous display of SEEG and ECOG
- Local average reference: Now available for all the groups of contacts
Bug fixes
- All "Filter" processes were replacing the actual data with the "Std" field
- Added legends to connectivity matrices displayed as images
- Added support for Matlab 2018a
February 2018
Anatomy
FreeSurfer: Import the vox2ras matrix from the .mgz MRI files (ignored until now)
FreeSurfer: Correct registration of additional .nii files with full FreeSurfer anatomy folders
- Export NIfTI: Exporting vox2ras transformation to .nii when volume was not initially .nii
- MRI 3D: New options "Upsample image" to smooth MRI display in 3D figures
- New option to add the ASEG atlas and the white matter surface to any 3D figure
Source modeling 2018
- New version of the linear inverse models: bug fixes and support for mixed head models
- Previous models will remain available as "Compute sources 2016"
Documentation: Source estimation tutorial
January 2018
Input / output
Support for the new RICOH MEG file format
Improved BIDS-MEG importer: derivatives folder (FreeSurfer, MaxFilter), acquisition info (_scans.tsv)
- Fixed EEGLAB reader: Keep events in epochs, allow epochs with multiple event occurrences
Keep acquisition dates in the database (DateOfStudy of the brainstormstudy.at files)
Processes
- Noise covariance: Match automatically noise covariance with subjects by acquisition date
- Average: Added option "median"
November 2017
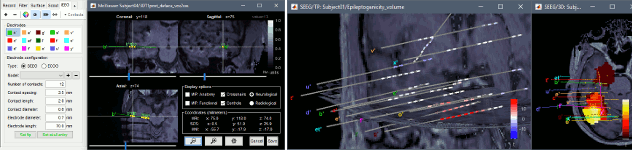
New tutorial: SEEG epileptogenicity maps
The tutorial SEEG epileptogenicity maps illustrates all the new features related with SEEG/ECOG: coregistration of MRI and CT scans, placement of depth electrodes in the post-implantation scans, SEEG bipolar montages, display and processing of intracranial recordings, computation of epileptogenicity maps.
Bug fixes
- Matlab 2017b: A bug in isosurface.m caused irregularities in head surfaces
- More guided protocol creation to avoid user errors and massive deletions of external folders
October 2017
Anatomy
New options for volume coregistration of multiple scans of the same subject (MRI/MRI or MRI/CT)
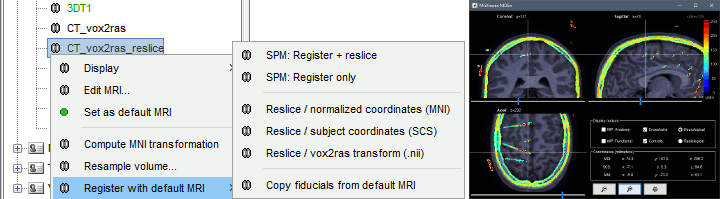
New interface for placing SEEG/ECOG contacts in the MRI Viewer
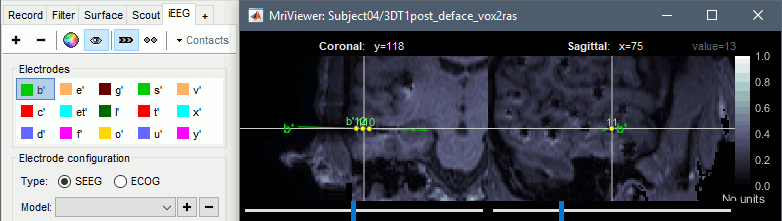
- MRI viewer: Various interface improvements (zoom, shortcuts)
- MRI viewer and MRI-3D: New menu Jump to maximum (shortcut "M")
SEEG/ECOG
- Various improvements for handling SEEG/ECOG data: Import menus, montages, display
Add event markers on the signals when the event names match the channel names
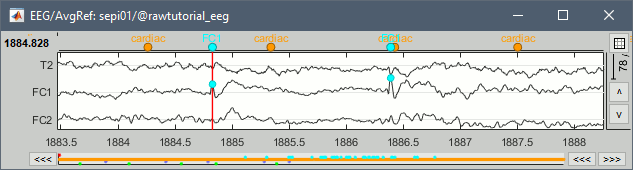
New tab Guidelines for computing epileptogenicity maps
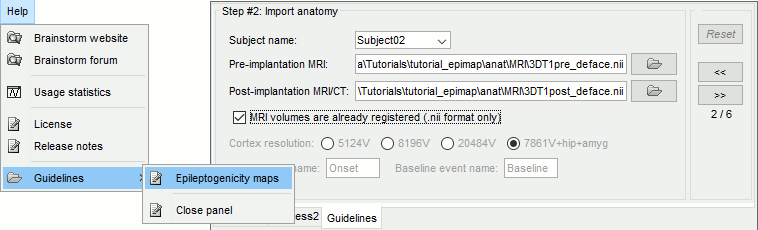
September 2017
Elekta: Major workflow changes
- When importing Elekta .fif files, the default acquisition SSP are not applied by default anymore. These SSP projectors were previously always used in Brainstorm for visualization and source estimation. However, they were meant to be used only to review online recordings from the acquisition computer.
- Now they are still imported but inactive by default. If you want to use them, for instance for backward compatibility purposes, enable them in the window "Select active projectors". If your recordings now look particularly ugly, our advice is to recompute SSP projectors with Brainstorm, rather than using the default acquisition ones.
Input / output
New DICOM converter (SPM-based): Use menu "Import MRI" and select file format "MRI: DICOM"
Export figures to Plotly: See tutorial
- Export volume grids as .nii
Import channel files in .tsv format (IntrAnat)
August 2017
New processes
Epilepsy > Epileptogenicity maps: See tutorial
Frequency > Time-resolved phase amplitude coupling (tPAC): See tutorial
Frequency > FieldTrip: ft_mtmconvol (Multitaper)
Connectivity > Phase transfer entropy
Artifacts > Undo 4D/CTF noise compensation
July 2017
Anatomy
SEEG/ECOG: New option to import sensor positions in subject or MNI coordinates
Computation of SPM canonical surfaces (alternative to FreeSurfer/BrainSuite/BrainVISA)
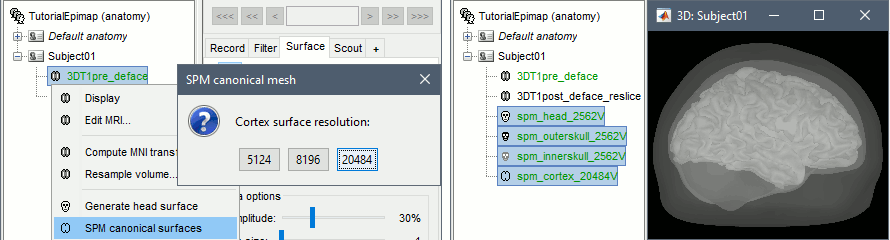
New anatomy template: USCBrain
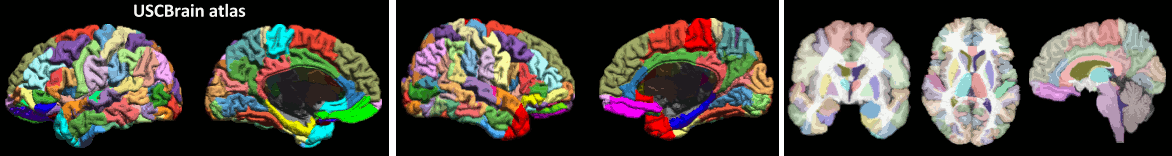
- MRI registration: New option to reslice based on the NIfTI sform/qform transformations
- Surfaces: Load .gii files as surface textures
Recordings viewer
- New buttons for zooming vertically
- New buttons for displaying the vertical and horizontal grids
Resize the display page with the red square at the bottom of the figure
- File viewer made stand alone with the new function bst_autopilot.m
- Optimized reviewing speed: New option "Downsample recordings for faster display"
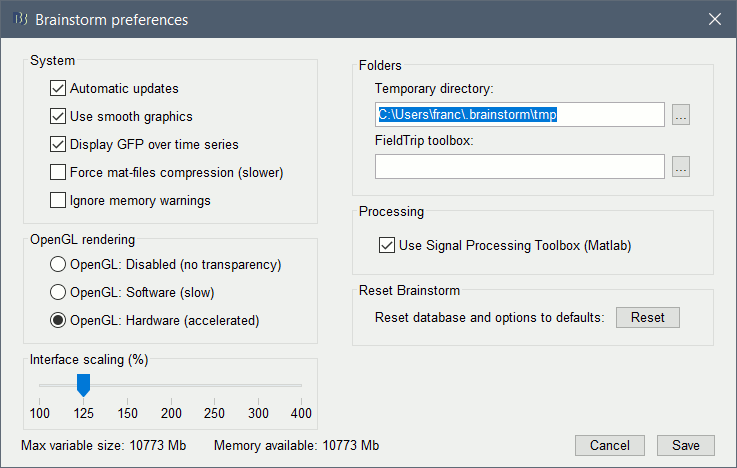
Support for high-resolution screens
If you have a high-resolution screen, the text and icons in the Brainstorm window may not scale properly, leading the interface to be impossible to use. Select the menu File > Edit preferences, the slider at the bottom of the option window lets you increase the ratio of the Brainstorm interface.
May 2017
New file formats
- EEG: Support for Nicolet .e files
- EEG: Support for ANT ASA .msm/.msr files
- EEG: Ugraded ANT EEProbe library for supporting recent files
- EEG: Support for ERPLab files
- Electrophysiology: Support for CED Spike2 .smr
- Electrophysiology: Support for Ripple Trellis NEV/NSx files
- Electrophysiology: Support for Backrock .ns6 files
- Export to EDF: Now Brainstorm can be used to convert any file format to EDF
Major bug fixes
- SSP: When epoching recordings (importing blocks of recordings to the database) from continuous files that were filtered AFTER doing the SSP/ICA cleaning, the SSP/ICA projectors were not properly reported in the channel file of the new imported folder. This was leading later on to a lack of accuracy in the estimation of the sources.
- EEG: Neuroscan .cnt files with low accuracy (int16) were not read correctly
April 2017
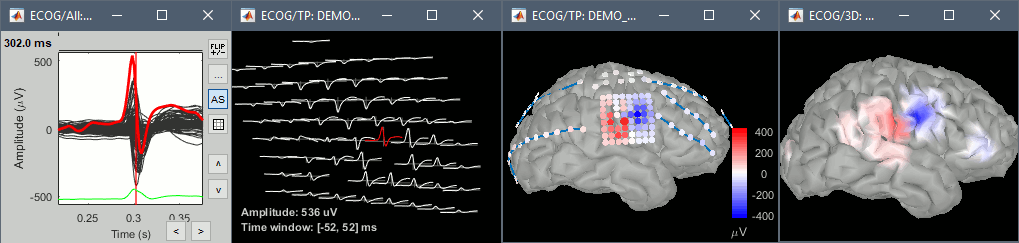
New display for ECoG and sEEG
Many new options are available to render sEEG/ECoG recordings or power: interpolations on the cortex surface, in the MRI volume, or 2D displays of the time series.
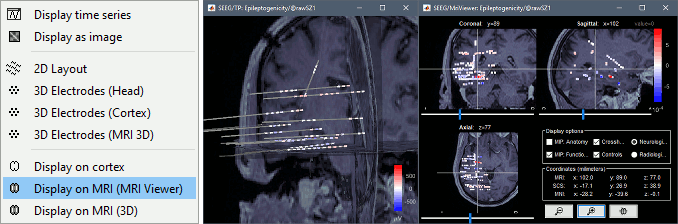
Default fiducials from MNI template
When using the digitized head shape for aligning the MEG/EEG sensors on the MRI, the fiducial points NAS/LPA/RPA do not need to be placed very precisely as they are used only as a first approximation. Computing the MNI transformation for the MRI now sets default positions for the NAS/LPA/RPA fiducial points, you can use these if the accurracy of their location is sufficient. This option allows a fully automatic import of the anatomy+recordings.
March 2017
New file formats
- EEG: Support for Micromed EEG TRC files
- EEG: Support for Nihon Kohden .EEG files (including newer 1200 systems)
- EEG: Export/import continuous recordings with SPM (.mat/.dat)
- SEEG/ECOG: Import contact positions in MNI coordinates
- Events: Support for Presentation .log files
Other improvements
New process: Connectivity > Amplitude envelope correlation
- Anatomy: Apply manual sensor-MRI registration to multiple datasets
- GUI: Added option to filter files by the comment of their parents
January 2017
New tutorial: Human Connectome Project (MEG)
The tutorial Human Connectome Project: Resting-state MEG explains how to download MEG recordings from the Human Connectome Project (HCP) ConnectomeDB database and process them into Brainstorm.
November 2016
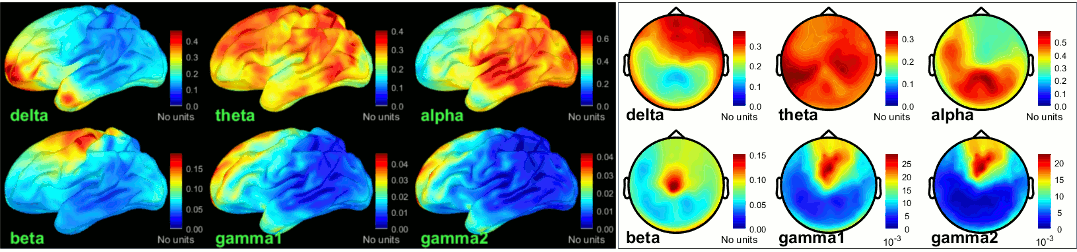
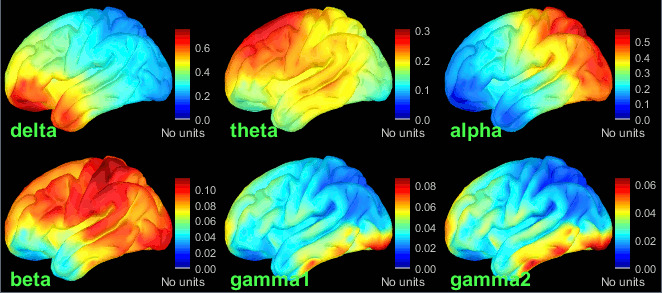
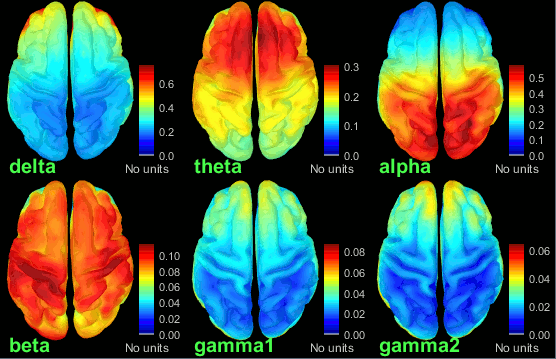
New tutorial: OMEGA database & resting state (MEG)
The Open MEG Archive (OMEGA) offers for download a large database of resting state MEG recordings. The tutorial MEG resting state & OMEGA database describes how to access this database, download a few subjects and process them in Brainstorm.
BIDS specification
The Brain Imaging Data Structure (BIDS) standard for neuroimaging data organization was first established for MRI and fMRI (Gorgolewski, 2016). It is based on simple file formats and folder structures that can readily expand to additional data modalities.
An extension of the BIDS for MEG datasets is currently under review. One objective is that analysis pipelines designed with major analysis tools (such as Brainstorm, FieldTrip, MNE, SPM and others) can be readily applied without requiring software or pipeline redevelopments.
We have implemented functions that load automatically into a Brainstorm database any dataset following the BIDS specification. This prototype is illustrated in the tutorial MEG resting state & OMEGA database.
Volume atlases
Volume atlases in subject space (eg. FreeSurfer's Aseg atlas) can be used in various ways:
Import as scouts for volume source models: See Volume source estimation.
Import as volume: Import the aseg.mgz file as a new MRI in the subject anatomy folder, then display it with the MRI viewer. New colormaps are available for representing labelled atlases.
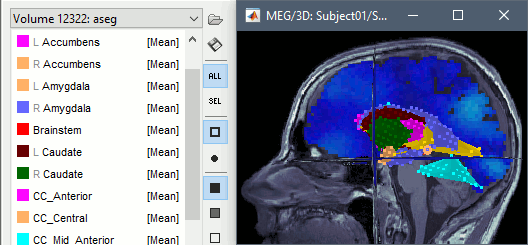
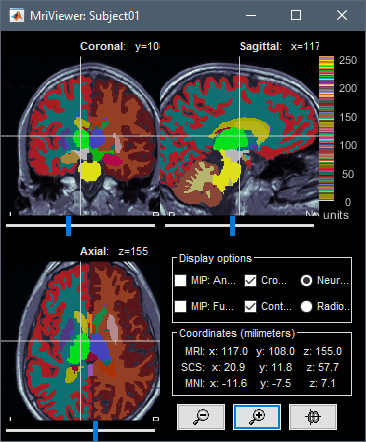
Volumes in MNI space
Volumes in MNI space can be imported and transformed to the subject space. This allows the new features described above also for standard volume atlases, such as the AAL atlas. You just need to make sure you select the file format "Volume mask or atlas (MNI space)" in the import options.
Import MNI volume atlases as scouts for volume source models.
Import MNI volume atlases as a subject volume, for overlaying on the subject structural MRI.
Import MNI volume atlases as surfaces (produces something similar to the "aseg" surface file).
To get the labels of the atlas displayed correctly, a .txt file with the same name should be present in the folder. Each line in this file must include the index of the label ("label_index label_name"), such as the atlases distributed as part of the MRIcron software.
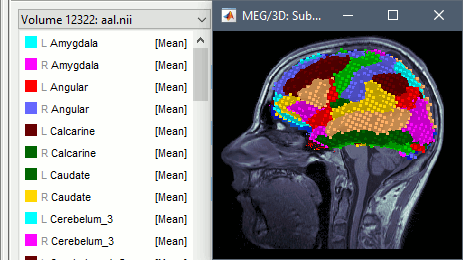
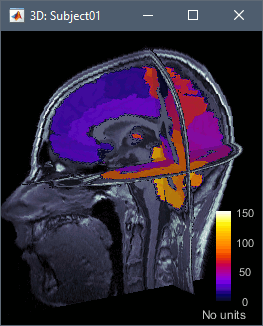
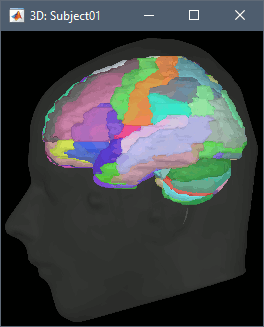
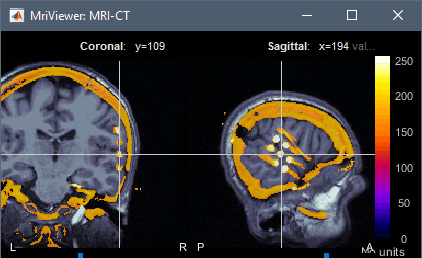
CT-MRI coregistration
The volume registration functions allow to import a CT scan in the anatomy folder of the subject, and then to overlay it with the MRI to mark sEEG or ECoG contacts. This solution computes a linear MNI transformation for both volumes (CT and MR) and applies the two transformations to the new imported CT.
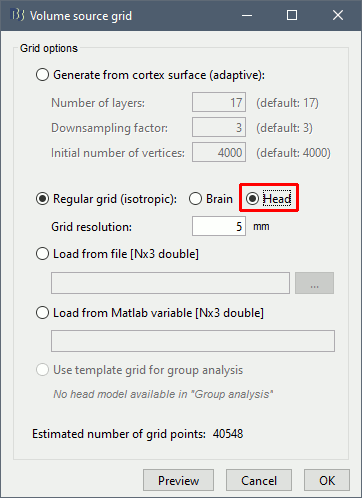
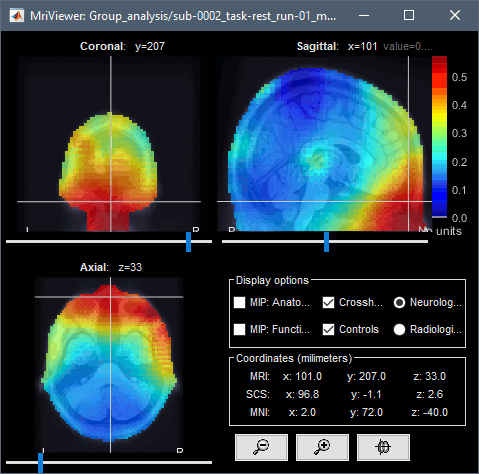
Extended source space
Head grid: The source space can be extended to the full head volume, this can help decide whether some sources are brain signals or artifacts (eg. eye movements or neck muscles, as illustrated below).
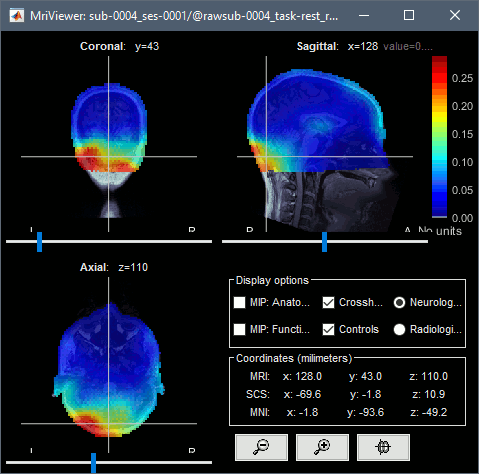
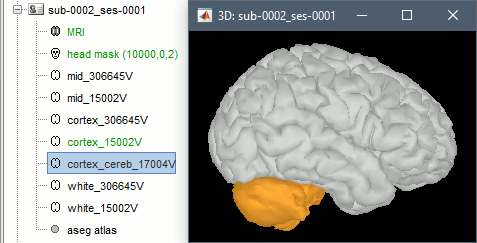
Import FreeSurfer anatomy: Creates an additional surface with the cerebellum by default.
FieldTrip: Improved support
Many new FieldTrip functions are now supported, especially for forward and inverse modeling:
ft_volumesegment: MRI volume segmentation (scalp, skull, brain)
ft_prepare_leadfield: Computation of forward models (calling also ft_prepare_headmodel)
ft_sourceanalysis: Inverse models (beamformers and minimum norm)
Previously made available:
ft_dipolefitting: Dipole fitting for MEG and EEG recordings
ft_channelrepair: Replacement of bad sensor with an average of the neighbors
ft_scalpcurrentdensity: Estimate of the SCD fields
ft_timelockstatistics: Statistics at the sensor level
ft_sourcestatistics: Statistics at the source level
ft_freqstatistics: Statistics in time-frequency space
Most data types can be converted easily between Brainstorm and FieldTrip structures (MRI, surfaces, sensors, recordings, head models, source maps, time-frequency results):
Right-click on any file > File > Export to file > Look for the FieldTrip option in the file formats.
toolbox/io/out_fieldtrip_*.m: Functions to convert from Brainstorm to FieldTrip structures.
toolbox/io/in_*_fieldtrip.m: Functions to load FieldTrip structures as Brainstorm structures.
October 2016
Brainstorm on GitHub
We moved all the code of Brainstorm to a public GitHub repository. You're welcome to join!
https://github.com/brainstorm-tools/brainstorm3
New band-pass filters
The frequency filters have been notably improved. They have been completely redesigned and are now fully documented in the tutorial Frequency filters. From the process "Pre-process > Band-pass filter", you can display the filter response in frequency and time domain, together with all its properties.
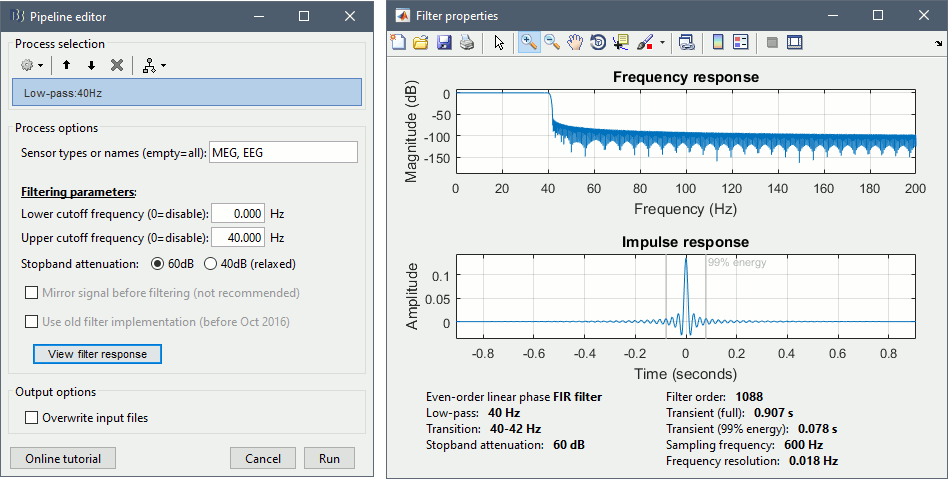
After filtering signals, new events indicate the transient durations to consider. This represents an absolute minimum, if possible always exclude the full filter length, documented in the filter response window.
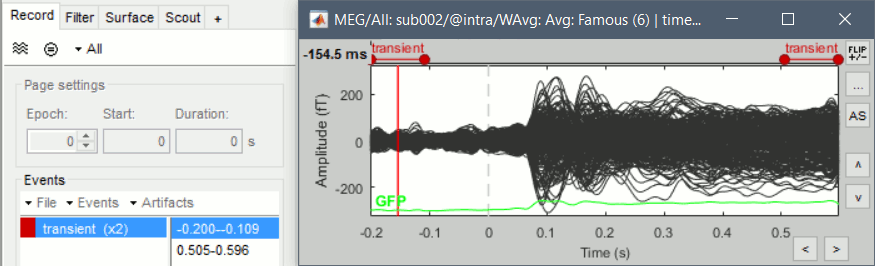
This modification has some impact in many processes using the band-pass filters: the SSP computation, the Hilbert transform, the visualization. These new filters are slightly slower but much more accurate. If reproducibility is a concern, use can select "Use old implementation" in the process options.
Other improvements
New process: Events > Detect cHPI activity (Elekta)
- Volume scouts: Improved rendering and constrained growth now enabled
Volume scouts: Can be exported as volume masks (menu Scouts > Export as MRI mask)
- Snapshots: Frequency contact sheets
- Forum: Added full text search on the forum using Google (available on the home page of the website)
- GUI: Support for high-DPI screens (Retina and other 4K displays)
- GUI: Support for Matlab 2016b
August 2016
2D topography plots
The projection of the EEG/MEG recordings on 2D maps have been updated. The new functions are more stable and the figures will be easier to integrate directly in your publications.
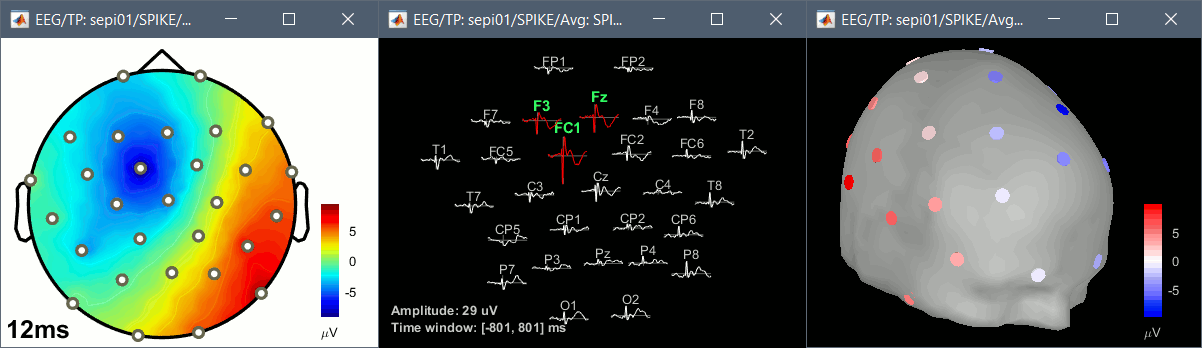
Mixed head models
The basic source reconstruction limits the source space to the left and right hemispheres of the brain. The mixed head models allow to include the cerebellum and various subcortical regions to the estimation and the visualization of the source activity. Their support has been extended to include new features:
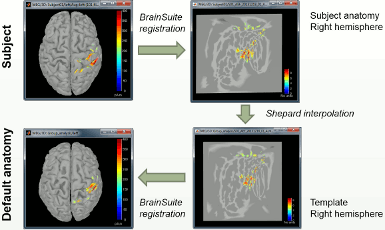
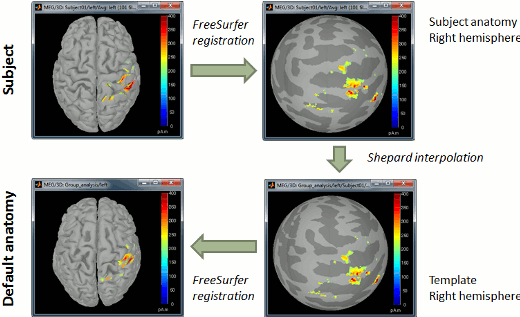
- Projection on an anatomy template for group analysis.
- The regions without orientation constrains are now flattened before the projection on the template (the source orientation is not a meaningful parameter at the group level).
- Additionally, the sources on the template anatomy can now be re-projected in the subject space.
New exploded view to visualize all the subcortical regions on the same view.

MRI coregistration
You may be interested in importing multiple volumes (MRI, fMRI, CT, PET, etc) in the anatomy folder of a subject. You can overlay volumes and compare the results obtained with different imaging modalities or at various stages of a surgical procedure, or use fMRI results in an ROI analysis of you MEG/EEG recordings. However these various volumes usually have different orientations or spatial resolutions, and in order to be displayed in the MRI Viewer in Brainstorm, the files must have the same size and same orientation. We added some functions to help you coregister these volumes.
New menu "Resample MRI" to reslice any MRI volume.
New menu "Register on default MRI" to co-register multiple MRI volumes of the same subject.
These two options are offered when importing additional volumes in the subject anatomy, if their dimensions or resolution do not match the initial structural MRI.
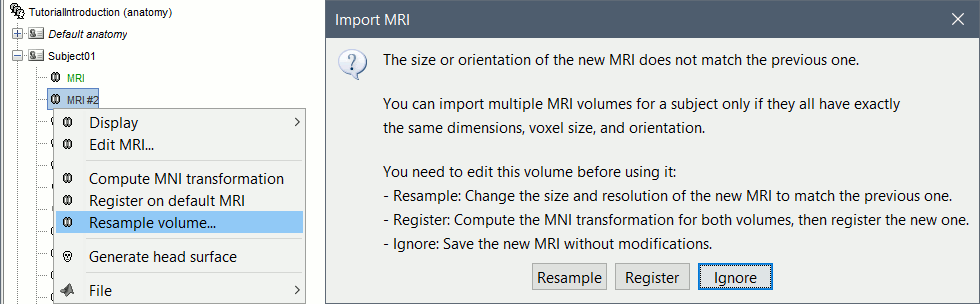
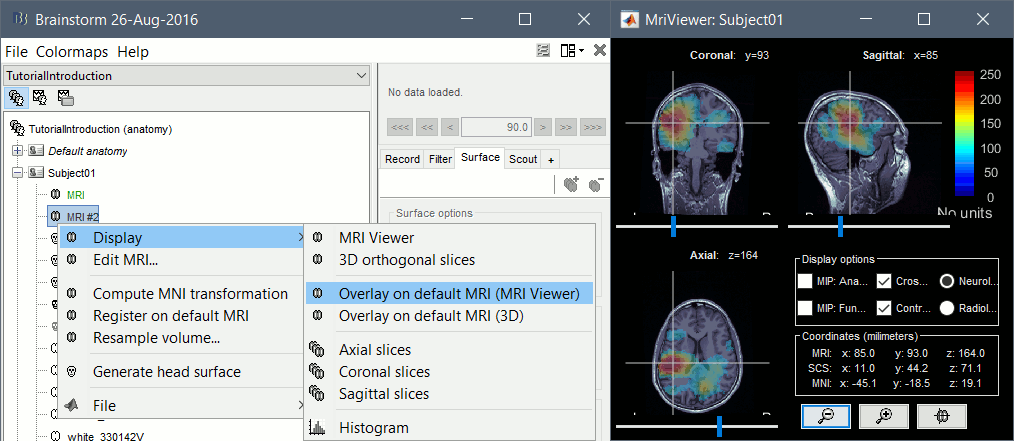
The computation of the MNI transformation is now available for all the MRI volumes (before this was restricted to isotropic volumes with a voxel size of 1x1x1mm). This option first resamples the volume to a 256x256x256 volume with a unit voxel size, and then runs the SPM8-based affine MNI registration.
When you have a secondary volume imported in your subject, you can display it as an overlay on the original MRI and configure the threshold and transparency from the Surface tab.
Updated advanced tutorials
Many of the advanced tutorials were outdated, they have all been updated to include the latest advances in cleaning procedures, workflow management and display.
EEG/Epilepsy: Includes a new section about ICA decomposition.
Elekta-Neuromag median nerve: Reference for processing Elekta MEG recordings.
Yokogawa/KIT median nerve: Reference for processing Yokogawa/KIT/Ricoh MEG recordings.
CTF median nerve tutorial: Dataset used in the previous version of the introduction tutorials.
Other advanced tutorials that have been improved recently:
July 2016
New tutorial: Group analysis
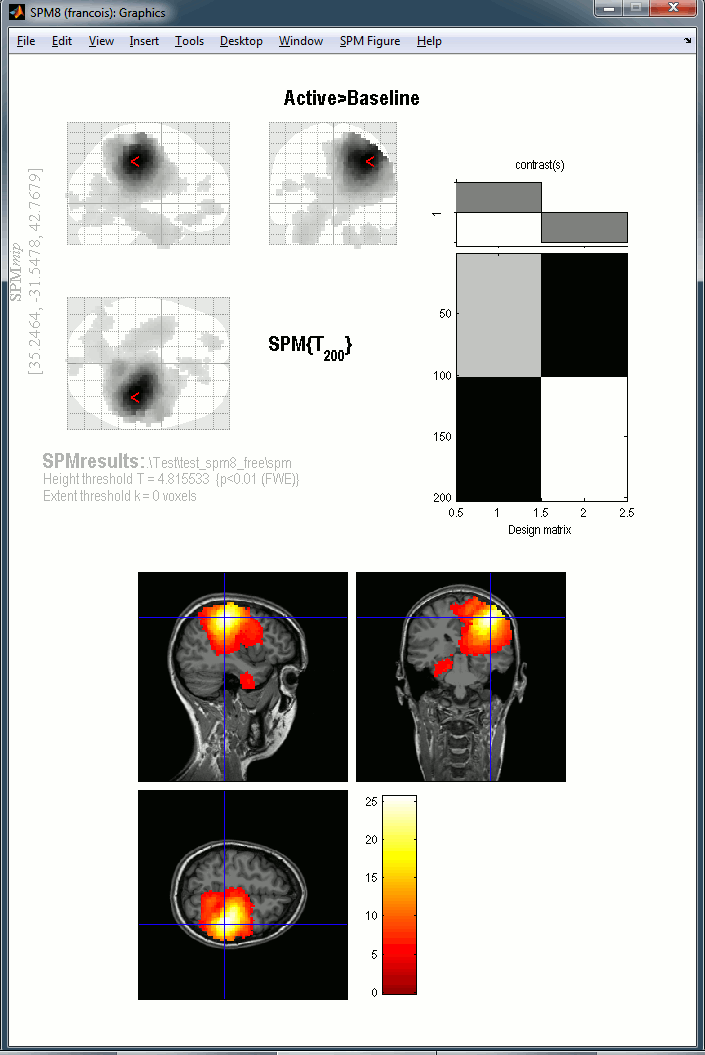
This new tutorial reproduces in the Brainstorm environment the analysis described in the SPM tutorial "Multimodal, Multisubject data fusion". The data processed here consists of simultaneous MEG/EEG recordings from 19 participants performing a simple visual recognition task from presentations of famous, unfamiliar and scrambled faces. The analysis is split in two tutorial pages: the analysis of a single subject and the group analysis.
This is part of a new effort in cross-validating the various academic programs for EEG/MEG analysis (EEGLAB, FieldTrip, SPM, MNE). The results will be presented at a satellite meeting at the Biomag conference, held in Seoul on October 2nd.
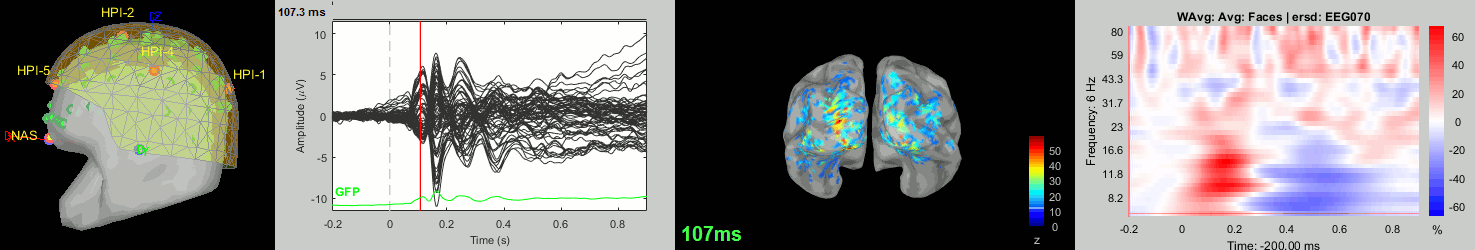
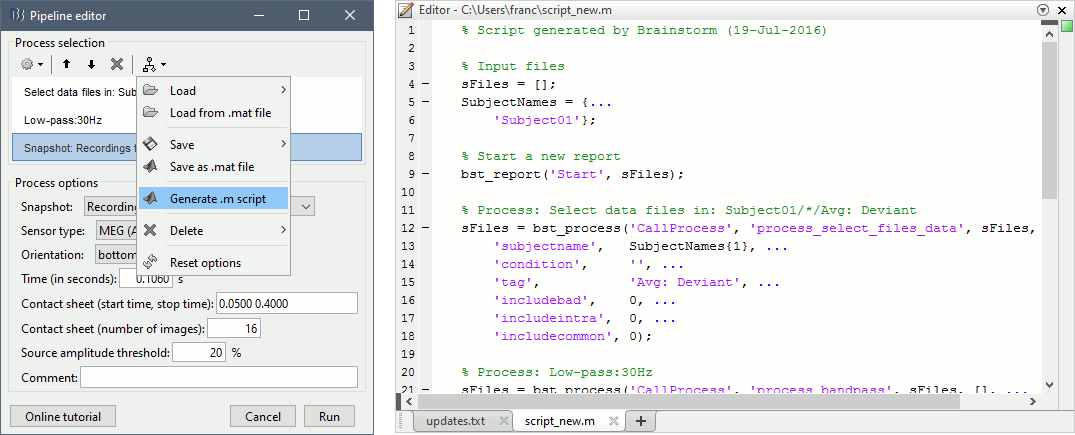
New tutorial: Scripting
Advanced users who want to go beyond the limits of the graphic interface, script large studies or add their own routines to the software now have access to an introduction to the Brainstorm API. The tutorial Scripting explains how to interact with the software from the Matlab command line or a .m script.
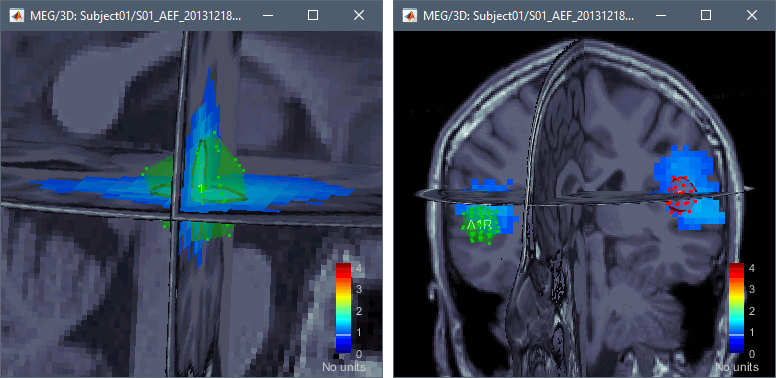
Volume sources
The support for volume source models has been improved and now includes new features:
Implementation of volume scouts: Volume source estimation tutorial.
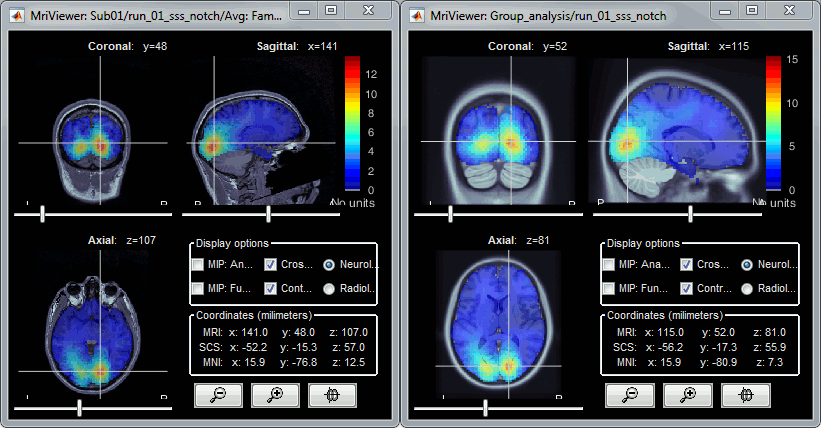
Projection on an anatomy template for group analysis: Subjects coregistration tutorial.
Anatomy: Defining the fiducials
We propose a new pipeline for processing and importing the anatomy for multiple subjects. This is described in this section of the Scripting tutorial: How to process an entire study.
New menu "File > Batch MRI fiducials": Sets the anatomical landmarks for all the subjects.
- MRI Viewer: New popup menu "Save fiducial file".
June 2016
Continuous recordings
New popup menu "File > Copy to database": Copies a native continuous file to the protocol folder, in order to make the database portable between computers.
New menu "File > Export protocol > Copy raw files to database": Does the same as "Copy to database" for all the continuous files available in the current protocol.
Continuous files can now be resampled with the process "Pre-process > Resample".
New menu "File > Load protocol > Change database folder": Changes the folder in which the new protocols are created, but does not have any impact on the existing protocols.
- Process reports: Export the reports to HTML files (including the screen captures).
Statistics
Redesign of all the functions related with statistic testing. See the new Statistics tutorial.
New process(2): Test > Parametric test: Paired/Independent
New process(2): Test > Permutation test: Paired/Independent
New process(1): Test > Parametric test against zero
New process(1): Test > Parametric test against baseline
New process: Extract > Find maximum in time
New process: Extract > Find maximum in frequency
- Allowed the execution of the averaging and statistics processes on the connectivity measures.
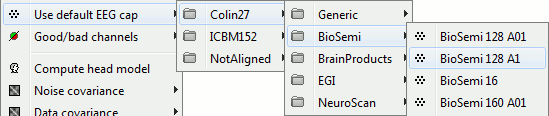
Default EEG caps
Added support for BioSemi caps with electrodes named "A1" instead of "A01".
- Added support for EGI Hydrocel caps with electrodes named "E001" instead of "E1".
Organized the default electrode positions by manufacturer.
May 2016
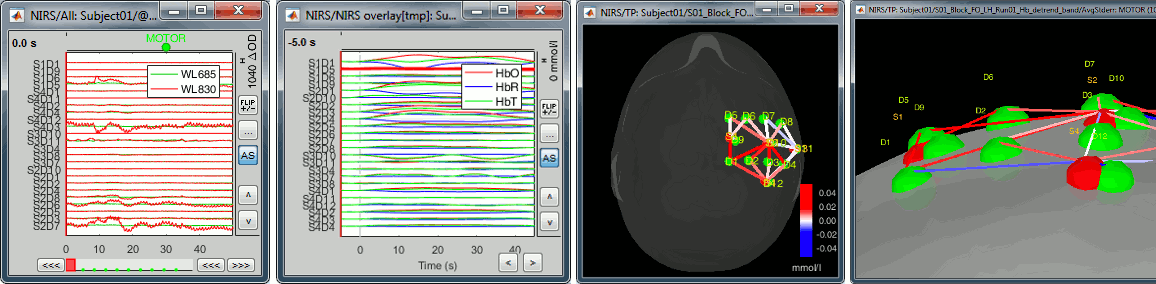
New tutorial: fNIRS
Many new tools are available for importing manipulating fNIRS recordings in Brainstorm. This is explained in a new online tutorial (work in progress): NIRS finger tapping.
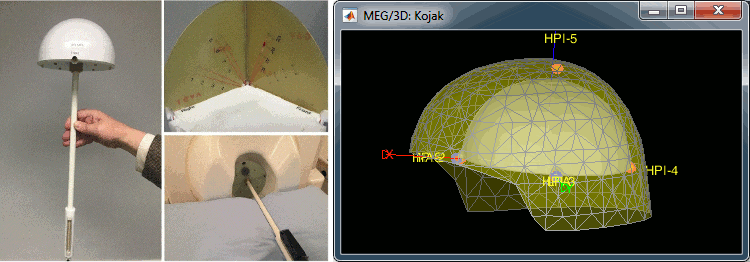
New tutorial: Elekta phantom
A new tutorial explains how to import and process Elekta-Neuromag current phantom recordings. The framework presented on this page is designed to be used for the quality control of an Elekta MEG system (after a maintenance for instance) and for evaluating dipole modeling techniques.
MEG current phantom (Elekta)
Baseline normalization
New process "Standardize > Baseline normalization" that replaces two existing processes: "Z-score normalization" and "Event-related perturbation (ERS/ERD)"
New recommendations for the normalization of source maps (never use an absolute value):
Source estimation tutorial (normalization).- Normalized source maps are now displayed with the same colormap as the original source maps.
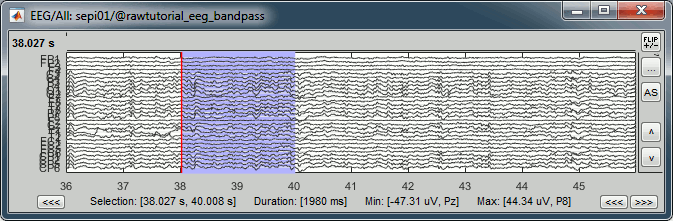
Recordings viewer
Min/max: You can now quickly get the minimum and maximum values in the recordings by simply selecting a time segment. If some channels are selected, it shows the extrema only for them.
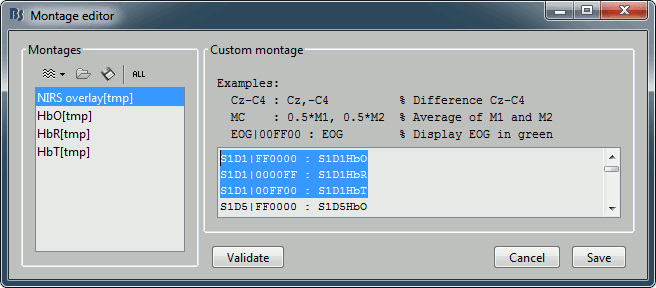
Overlay: The montage editor now supports a syntax for overlaying multiple signals. You just need to set multiple channels with the same display name.
Color: The montage editor now allows the direct definition of a color for specific channels: Use the specific syntax for the display name: "NAME|RRGGBB" (hexadecimal notation).
Make a database portable: New menu "Export protocol > Copy raw files to database".
Make a database portable: New popup menu for raw files "File > Copy to database".
Time-frequency
- Hilbert transform: The new option "power" is now the default (instead of "magnitude" previously)
- Now keeping track of the edge effects when extracting time windows from a larger file.
- PSD: Estimation directly from source files attached to a raw link.
- PLV: Added an option to save the magnitude.
April 2016
Major bug fix: BrainAmp
The gain of the channels in some BrainVision files (BrainAmp or V-Amp) was not read correctly.
The files impacted are the ones for which the gains are specified in the .vhdr/.ahdr files, in section [Channel Infos], field <Resolution in microvolts>.
- To fix the problem: Delete all the imported files and links to continuous files and start over.
Scouts: Sign flip
Major modification in the computation of the scouts time series (constrained orientations only). Previously, the sign flip of sources with opposite directions was applied only if it increased the average scout amplitude. Therefore the procedure was dependent on the data, making it difficult to compare across conditions and subjects. Now the sign flip is always applied.
To disable completely the sign flip: Use the process "Extract > Extract scouts time series".
Source modeling
Major update of the new inverse interface (Compute sources [2016]).
- Bug fix: Dipole scanning orientations were wrong for the unconstrained source maps.
Save whitened recordings: New menu available when right-clicking on source files in the database explorer. It shows how the recordings are actually used in the minimum norm source estimation, after the pre-whitening computed based on the noise-covariance matrix. If this doesn't look correct, you might think about changing the options for the noise covariance regularization (increasing or decreasing the amount of regularization.
- Data covariance computation can now use a separate baseline window for DC removal.
Dipoles
- New dipoles exploration panel (more details to come)
March 2016
Input / output
Support for FieldTrip raw data file (trials)
Convert imported files to continuous files: Right-click > Review as raw
Copy external raw files to the database: Right-click > File > Copy to database
Make the entire protocol portable: File > Export protocol > Copy raw files to database
Software distribution
- Script distribution: Added support for Matlab 2016a
- Compiled distribution: Switched to Matlab 2015b (smaller and faster)
- Compiled distribution: Does not crash anymore when Brainstorm is closed
February 2016
Major bug fix: MEG reference sensors
Bug fix: The geometry of the CTF MEG reference sensors was not imported correctly. The forward model was not as accurate as expected, especially with a very high signal-to-noise ratio.
For more accurate results, re-import the recordings and compute a new forward model.Bug identified: Similarly, the 4D MEG reference gradiometers may not be modeled properly
(waiting for tests from any site with a 4D system)New tutorial to investigate this problem: CTF current phantom
Channel file: New menu "Display sensors > CTF coils (ALL)" to represent the CTF reference sensors.
Similar menus available for 4D and Elekta-Neuromag systems.
Anatomy
New template BCI-DNI_BrainSuite: Description of the atlas | Brainstorm documentation
Replaced the default template Colin27 with ICBM152: Brainstorm documentation
Process "Sources > Spatial smoothing": Replaced with a function from the SurfStat toolbox
Parametric statistics
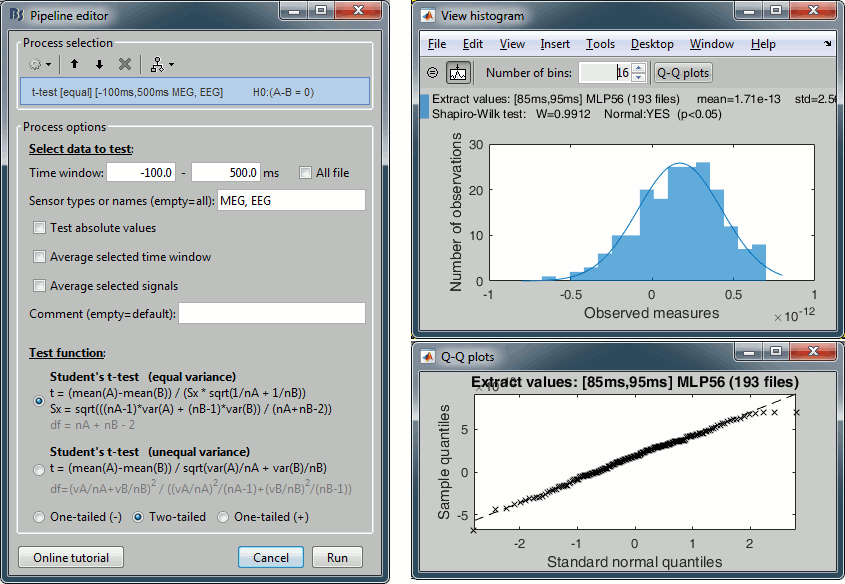
- New functions for parametric tests (one-sample, two-sample, paired)
- Histograms: Added the result of the Shapiro-Wilk test and a "Q-Q plot" button
New Process2: File > Select uniform number of files: Selects the same number of files in A and B
January 2016
Display as image
[Trials x Time] Right-click on a group of trials > Display as image: Raster plot for the values of one sensor across the trials. Change the channel from the Display tab (or up/down arrow key).
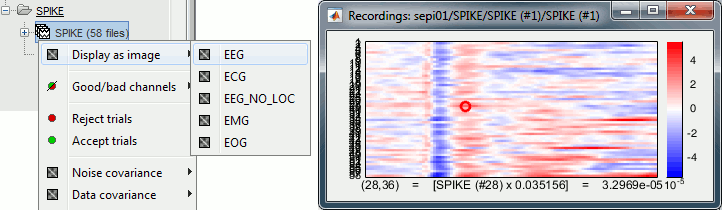
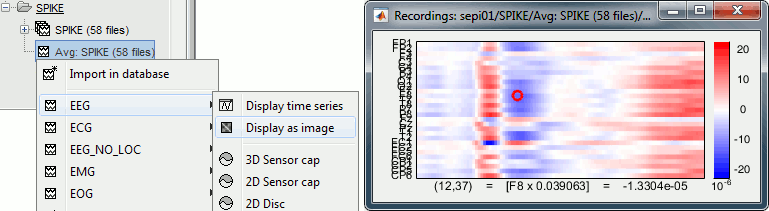
[Channels x Time] Right-click on one single file > MEG/EEG > Display as image:
Shows the same information as the "time series" view, but with colors instead of lines.
Synchronization with an eye tracker
A new tutorial from Martin Völker explains how to import the eye movements detected by an eye tracking device into an EEG/MEG file. The new generic process "Synchronize > Transfer events" can be used for synchronizing the event markers between files with different sampling rates.
See tutorial: Synchronization with eye tracker.
November 2015
Documentation
New introduction tutorials: Difference and Statistics
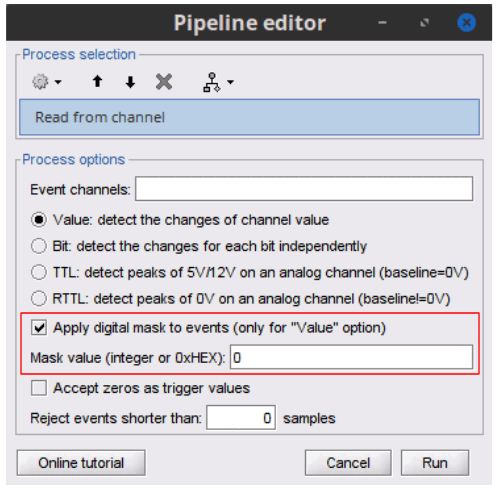
New button [Online tutorial] in the pipeline editor, to jump instantly to the corresponding tutorial.
Input / output
Support for EyeLink eye tracker recordings (.edf)
- Support for g.tec Matlab exports (.mat)
- Support for Brainsight NIRS systems (Homer file format)
- Support for Cartool .mrk events file
- Support for BESA .evt events file
Microstate segmentation
A new external plugin for microstate segmentation developed at John Cacioppo's lab (UChicago) has been made available publicly in Brainstorm. See the CENA website and the corresponding online tutorial.
October 2015
New processes
Standardize > Scalp current density (FieldTrip: ft_scalpcurrentdensity)
Standardize > Concatenate signals
Extract > Extract values: Generic process for extracting and concatenating blocks of files
File > Select uniform number of trials
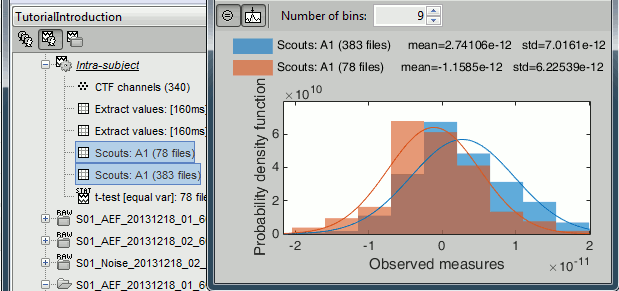
Histograms
Display of value histograms for all the files in the database (right-click > File > View histogram)
September 2015
New introduction tutorials
We completely re-organized the tutorials page. A new set of introduction tutorials is now available for beginners to learn the basics and for expert users to find references and technical details. We added a lot of material to describe the improvements we brought to the software in the past five years.
We encourage advanced user to have a look at them to learn about the new features and the new best practice guidelines we came up with over the years.
Input / output
- Support for ITAB MEG file format
- Support for Neuralynx file format (.ncs)
Support for MEGA NeurOne file format (.bin)
Support for Blackrock NeuroPort file format (.nsX, .nev)
Support for BrainVision channel files (.bvct, .bvef)
Import FieldTrip structures: data_timelocked saved as .mat files
Export FieldTrip structures: source files, time-frequency maps and head models
Source modeling
- Forward model: Generation of regular isotropic source grids
New process: Source > Dipole fitting (FieldTrip: process_ft_dipolefitting)
Anatomical registration with BrainSuite
BrainSuite now provides an accurate registration method to the BrainStorm anatomy templates (ICBM152, Colin27). See the BrainSuite tutorial.
August 2015
FieldTrip statistics
Some FieldTrip functions can be called from the pipeline editor in Brainstorm, this brings a lot of long-awaited tools for statistical analysis. See the Statistics tutorial.
- Permutation-based non-parametric t-tests
- Cluster-based correction for multiple comparisons
This is done with direct calls to FieldTrip functions:
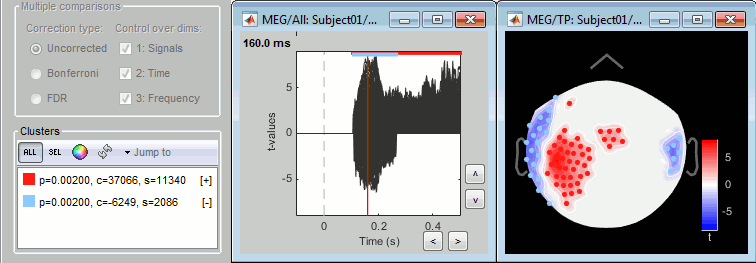
ft_timelockstatistics, ft_sourcestatistics, ft_freqstatisticsNew displays for exploring significant clusters
New anatomy template: One year old infant
Christian O'Reilly proposes a new anatomical atlas for one year old babies, described in this forum post: http://neuroimage.usc.edu/forums/showthread.php?2123-Atlas-for-1-year-old-babies
To use this template: create a new subject using an individual anatomy, right-click on the subject >
Use template > Download: Oreilly_1y.
Project scouts between subjects
A simple yet very powerful tool is now available to project a region of interest between surfaces: from different surfaces of a subject, or between a subject and a template. When using surfaces generated with FreeSurfer, this projection uses the accurate registration described in the coregistration tutorial.
In the Scout tab, this is available in the menu Scout > Project to...
Scouts from MNI coordinates
You can now start a scout from anatomical positions specified in MNI coordinates.
In the Scout tab, this is available in the menu Scout > New: coordinates.
Z-score normalization
Modifications were made to the Z-score process, specially for handling the normalization of sources:
- The static and dynamic versions of the process were merged. Now the dynamic version is only called for the normalization of kernel-based source files.
- Unconstrained sources: Now it first converts the unconstrained sources to flat maps by taking the norm of the three dipole orientations, then computes the Z-score of the norm.
- The dynamic version does not save new shared kernels anymore, because it doesn not make much sense to compute the Z-score of single trials.
July 2015
New inverse functions
New functions are now available for solving the MEG/EEG inverse problems. Not all the options are publicly available yet, but they will be soon. See the new tutorial: Source estimation.
Artifact detection
A new process is available for helping with the detection of non-standard artifacts in MEG/EEG recordings: Events > Detect other artifacts. It is documented in the new tutorial Additional bad segments.
Decoding and MVPA
The recent work from Oliva's lab at MIT on decoding experimental conditions with MEG is now available as a process in Brainstorm. It allows to run support vector machine (SVM) and linear discriminant analysis (LDA) classification on MEG data across time. It is documented in the tutorial Decoding conditions.
Cichy RM, Pantazis D, Oliva A (2014)
Resolving human object recognition in space and time, Nature Neuroscience, 17:455–462.
Custom colormaps
Custom colormaps can be saved in the user preferences, edited and re-used easily. This is documented in the new colormap tutorial.
June 2015
Standard deviation/error
Two new options are available in the proces "Average files", to compute the standard deviant or the standard error together with the average. This additional information is stored in the field Std of the average files, and is displayed as a bounded line around the signals when available.
Synchronized videos
You can attach synchronized videos to the continuous file viewer. Right-click on the "link to raw file"
> File > Add synchronized video. There are many ways to sync EEG and videos, the mechanisms you use are probably not supported yet. Please contact us if you are interested in using this feature.
April 2015
Major bug fix: dSPM
The dSPM values displayed were not scaled properly in the following configuration: Shared dSPM kernel calculated for a folder that contains at the same time single trials and averages of these trials. Values were wrong when displaying the sources for the averaged files using this shared kernel.
The new values are multiplied on the fly by a fixed factor: sqrt(nAvg)
where nAvg is the number epochs that are were averaged to compute the average.
This issue is documented in the source estimation tutorial.
MNI coordinates
It is now possible to use MNI coordinates to navigate in the original individual MRI volumes. We estimate the affine transformation from the subject space to the MNI ICBM152 template using the SPM12 software (spm_maff8 ©J Ashburner). The general approach is described in the following article:
Ashburner J, Friston KJ, Unified segmentation, NeuroImage 2005
To compute the linear transformation matrix between the individual MRI and the ICBM152 template, click on Compute MNI transformation in the MRI Viewer or the popup menu for the MRI.
New functions are also available for converting between coordinate systems: cs_convert and cs_compute.
New processes
FieldTrip > ft_channelrepair
Standardize > Interpolate bad electrodes
Standardize > Normalized difference (A-B)/(A+B)
Standardize > ERDS with two inputs
Events > Detect multiple responses
Events > Detect movement
Test > Student's t-test against zero
Input / output
Export data files to FieldTrip structures (out_fieldtrip_data)
- Export MRI overlay to MRI files (allows to export dipoles density)
- New popup menu "Import data matrix" when right-clicking on a study folder
March 2015
EEG: Artifact cleaning with ICA
Many EEG users are familiar with the Independent Component Analysis (ICA) for identifying and correcting artifacts such as eye movements, muscle activity or 60Hz. We added methods implemented initially in EEGLAB for calculating ICA decompositions of MEG and EEG signals. The methods available for now are Infomax and JADE. We will add soon other methods widely used in the EEG community: SOBI and AMICA.
The ICA decomposition is available for continuous files only and can be found next to the SSP menus, either in the pipeline editor or in the Artifacts menu of the Record tab.
The SSP interface has been extended to select interactively the ICA components and display their spatial topography and time series.
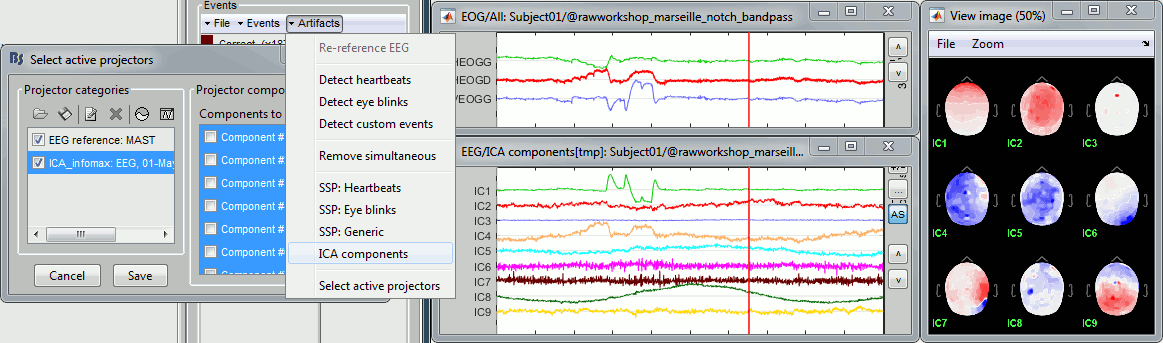
EEG: Montage editor
Improved flexibility of the montage editor, for supporting some common operations in EEG analysis:
- Support for linked EEG references.
Mixing channels from different modalities in the same montage. This is useful for re-referencing the EEG with an electrode that is not classified as "EEG" in the channel file. It allows also to review multiple types of signals in the same figure: EEG, EOG, ECG, EMG, etc.

EEG: Permanent re-referencing
EEG recordings can be re-referenced using projectors, similar to the ICA or SSP projectors. This allows to change permanently the reference without having to create a montage and explicitly apply it to the recordings. This feature is available in two ways:
Process: Standardize > Re-reference EEG
Record tab: menu Artifacts > Re-reference EEG
February 2015
Intra-cranial recordings
New options are available to display and process invasive recordings, sEEG (depth electrodes) and ECoG (grids of strips of subdural electrodes), developed in collaboration with the Cleveland Clinic and the Freiburg University Hospital.
New display mode: 3D Electrodes
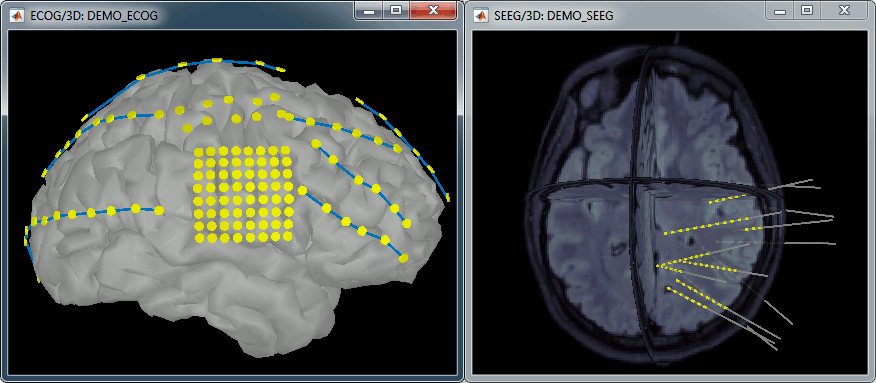
Edit the position of the contact positions using the MRI Viewer.
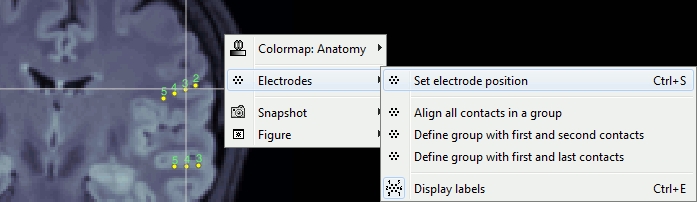
Grouping contacts using the new field "Group" in the channel file. Use the channel file editor to modify these groups if they haven't been identified correctly (right-click > Edit channel file).
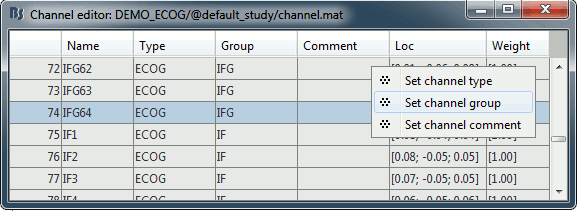
Set of temporary bipolar montages automatically created when loading SEEG/ECOG recordings.

Computation of the forward model for ECOG and SEEG concacts using the OpenMEEG BEM model.
- These tools are still in development, we will document them with a tutorial when they are stable.
January 2015
Major bug fixes
- Error in reading the BDF/EDF channel gains, the signal amplitudes were not scaled correctly
- Error in the calculation of the scouts values with process_extract_scouts in the case of constrained sources and atlases with non-sorted vertices (older atlases loaded with new code)
New supported file formats
- MEG: KRISS MEG .kdf file format
- MEG: BabyMEG system
- MEG: Neuromag Graph event .evl file format
EEG: Compumedics ProFusion Sleep 4 file format
- Import EEGLAB: Added support for categories "latency" and "type"
New processes
Connectivity > Time-resolved correlation and coherence
Events > Convert to simple event
Events > Convert to extended event
December 2014
New processes
Standardize > Spectrum normalization: Corrects for 1/f decrease in power in TF and PSD results.
Pre-process > ARIMA filter: Spectral flattening of a signal (use before TF decomposition).
Average and statistics: All the processes grouping multiple subjects now save their results in a subject "Group analysis" instead of the "Inter-subject" node.
November 2014
Changes in scout processing
We made some important modifications to the way the scouts where handled in the processes. Now it is possible to select the atlas directly in the options of the process, and to process at the same time scouts from different subjects with different list of vertices.
The new process function for extracting scouts is process_extract_scouts (before it was process_extract_cluster). The scout option value in the process is now a cell array, instead of the full scout structure. Example: {'User scouts', {'Scout1', 'Scout2'}}
This will affect all the processes that use scouts: time-frequency decompositions, power spectrum estimates, connectivity and PAC metrics. We tried to keep all the processes compatible with the previous input format, but there might be some modifications to do in personal scripts. If you have processes that use scouts structures in input, you might need to edit them to accept this new format. You can look at how it is done in process_canoltymap.m, or contact us through the user forum for help with updating your scripts.
October 2014
New anatomy template: Infant 7 weeks
The anatomical atlas presented in Kabdebon's article Anatomical correlations of the international 10–20 sensor placement system in infants is made available as a template in Brainstorm. To use this anatomy: create a new subject using an individual anatomy, right-click on the subject > Use template > Download: Infant7w.
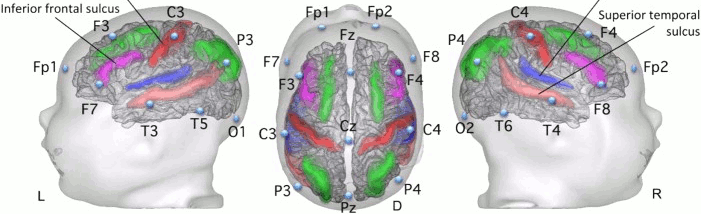
EEG electrodes positions
It is now easier to manage the 3D positions of the EEG electrodes. To add electrodes positions to an existing channel file: right-click on the channel file > Add EEG positions. You can import the points from a file or from one of the default sets of positions available in Brainstorm.
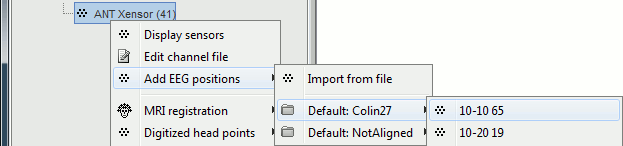
Brain Entropy in space and time
A tutorial is available online to introduce a new Brainstorm plug-in BEst: Brain Entropy in space and time which offers three new source localization approaches for EEG and MEG data, based on the framework of the Maximum Entropy on the Mean (MEM). These methods are particularly dedicated to estimate accurately the spatial extent of EEG/MEG generators with time series stable within parcels, to localize the generators of oscillatory patterns and synchrony.
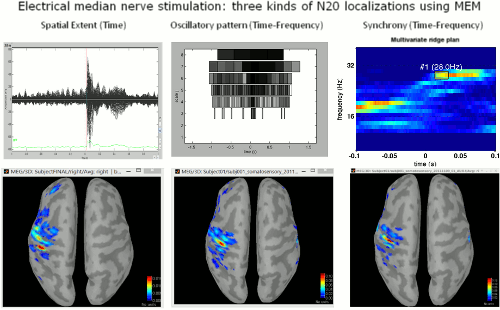
Data export
All the files in the Brainstorm database can now be exported as ASCII, CSV and Excel files: channel positions, recordings, sources, time-frequency maps and connectivity matrices.
September 2014
Filtering continuous files
Major modifications were made to the way the continuous files are handled:
The continuous files saved by Brainstorm when applying a filter to a native file are now saved in a new file format (Brainstorm binary, extension .bst), regardless of the original file format. The values in this file are saved in floating point double precision to avoid the re-conversion to integer 16bit needed to save CTF of FIF files, which cause significant loss of data quality.
- These files are not saved anymore in the same folder as the original data, but in the Brainstorm database itself. Removing the links to the processed files from the Brainstorm database now deletes the actual files.
All the frequency filters can now be applied to all the file formats, EEG and MEG.
To revert to the previous behavior (saving to native CTF or FIF files, in the database folder), use the menu File > Edit preferences > options "Store continuous files in original folder" and "Save continuous files in original format".
- The channel file structure is not copied anymore in the sFile structure used as the descriptor of the continuous files. This information can be read only from the channel file now. Personal scripts might need to be updated to consider this modification.
August 2014
MEG auditory tutorial
A new tutorial proposes a standard pipeline for the complete analysis of a simple oddball paradigm in MEG: MEG auditory tutorial. It presents the workflow in the Brainstorm environment, the equivalent documentation for the FieldTrip environment will be available on the FieldTrip website.
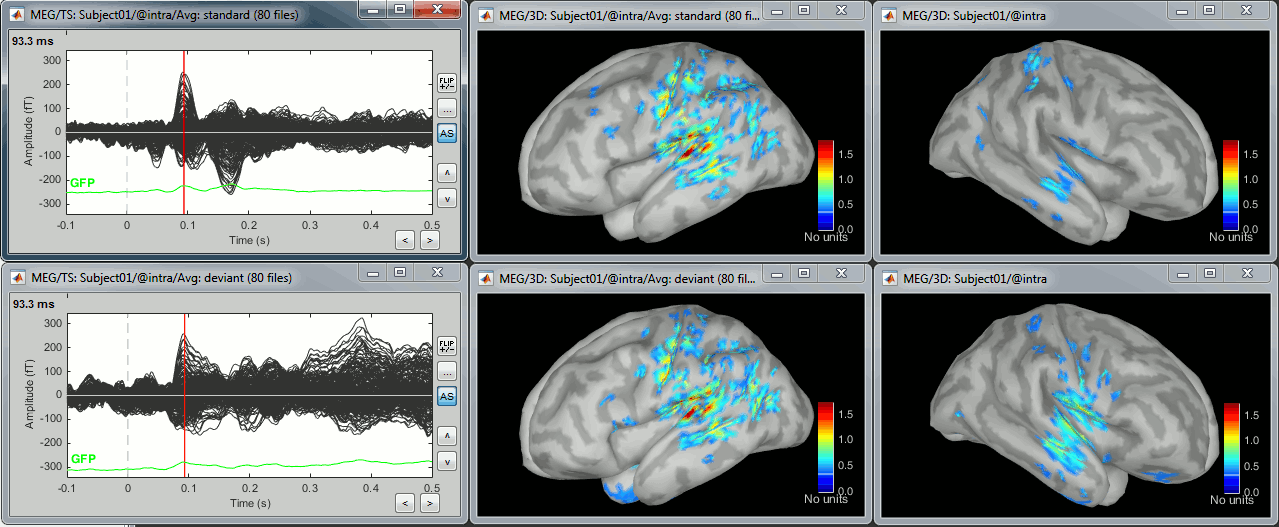
July 2014
New processes
- Notch filter: Offered as an alternative when the sinusoid removal doesn't work well
- Bandstop filter: Same thing for larger artifacts
- Remove linear trend
File > Select files: Get all the files from a specific folder with a process
- LCMV Beamformer (HL Chan implementation)
Interface improvements
- SSP generic: New options "Average", "Use existing SSP projectors" and "Save average artifact"
- Display of the EEG electrodes in the MRI viewer
- Time-frequency figures: New option "smooth display"
New figure popup menu: Snapshots > Open as figure
- Support for the new Matlab2014b graphics system
June 2014
Resting-state MEG and phase-amplitude coupling
Two new tutorial are available online to explore the cross-frequency coupling in MEG resting state recordings: Resting state MEG and Phase-amplitude coupling
They will allow you to reproduce step-by-step the analysis and the results illustrated in this article:
Canolty RT, Edwards E, Dalal SS, Soltani M, Nagarajan SS, Kirsch HE, Berger MS, Barbaro NM, Knight RT,
High gamma power is phase-locked to theta oscillations in human neocortex
Science, 313(5793):1626-8.
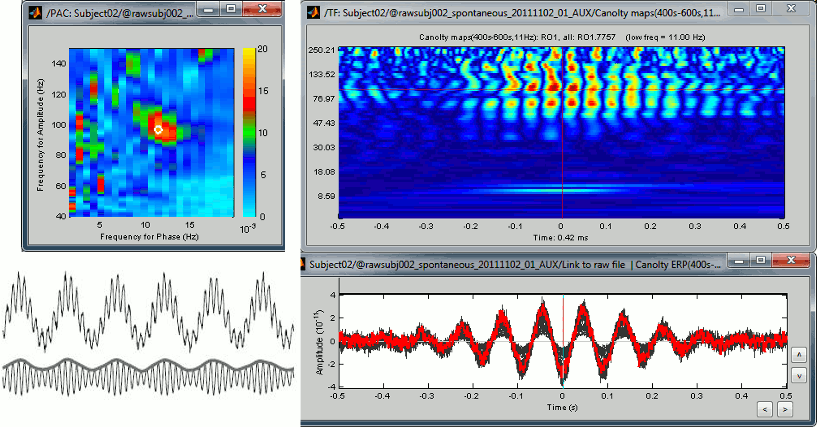
Elekta-Neuromag tutorial
A updated version of the Elekta-Neuromag tutorial is finally available: click here.
May 2014
New processes
- Apply montage: To save the result of a custom EEG electrode montage
- Surface smoothing: Now supportes time-frequency files in input
- Export to SPM: Now supports time-frequency files in input
Other improvements
- Display EEG/ECoG/sEEG electrodes in the MRI viewer
- MRI Viewer: Allow the direct overlay of two MRI volumes
- Spectrum plot: The frequency axis can be displayed using a logarithmic scale
- SSP: Create from one time point of recordings
- EDF file format: Added the support for multiple Annotation channels
April 2014
Mixed head models
New developements allow more flexible configurations of source models. Now, different brain regions (cortical or subcortical) can be modeled using different constraints. From the Scout tab, select the menu Atlas > New atlas > Source model option. All the scouts present in this atlas can be associated with different source constraints (surface/volume, constrained/free orientation). Then, when computing the head model, select the option "Custom source model".
This work will be soon part of a detailed tutorial showing how to use efficiently sub-cortical atlases for estimating sources in the deeper regions of the brain.
3D scouts
Scouts may now be defined on 3D source grids. Display the volume-based source results in 3D, then use the Scout tab exactly the way you would do it for surface maps.
March 2014
Yokogawa tutorial
A new online tutorial teaches Brainstorm users how to import and process Yokogawa MEG recordings:
Yokogawa/KIT tutorial
New processes
- Granger causality spectral
- Native support for scouts in all the connectivity functions
- Simulation of auto-regressive signals
- Simulation of recordings based on scouts time-series
- SSP calculation with two inputs for more flexibility
February 2014
EEG and epilepsy
Many features have been added for displaying and processing clinical EEG and invasive measurement data (ECoG and LFPs). These tools are illustrated in a new online tutorial EEG and epilepsy. It is based on a clinical case from the Epilepsy Center at the University Hospital of Freiburg, Germany. The anonymized dataset can be downloaded directly from the Brainstorm download page.
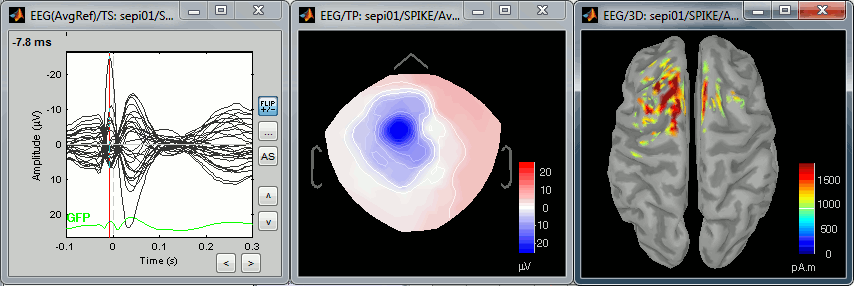
January 2014
Montage interface
The "channel selection" menu and the "EEG average reference" button have been replaced with a more flexible interface to edit montages. Montages can be used to simply select a list of sensors, re-order them in the display or to do more complex operations. The term refers mainly to EEG, to define the referencing system used to display the values recorded on the electrodes: bipolar montages, custom reference electrodes, average reference. See tutorial: Exploring the recordings
Some EEG montages recommended by the American Clinical Neurophysiology Society have been added to the distribution: longitudinal bipolar, transversal bipolar and referential. They are saved in .mon files in the folder brainstorm3/toolbox/sensors/private.
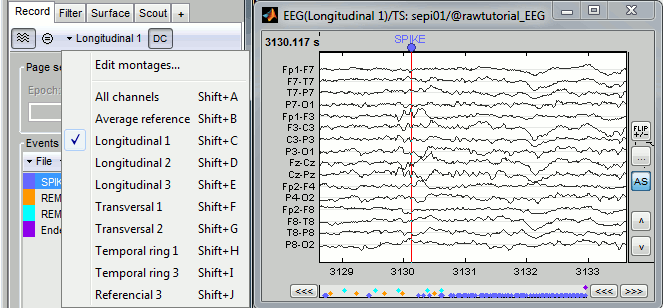
Average reference: SEEG and ECoG
The average reference is now offered in the the list of available montages for EEG, SEEG and ECoG recordings. By default, the average is calculated from all the electrodes and the same value is subtracted everywhere. In some cases, you may be interested in calculating different averages from different subgroups of electrodes: multiple cortical grids or electrodes stripes. To do this, you have two options:
Create a montage manually that defines exactly what are the operations to perform. For instance, if you have 4 electrodes E1..E4 you want to group together, enter four lines, one for each electrode, with this structure: E1: 0.75*E1, -0.25*E2, -0.25*E3, -0.25*E4
The main problem with this approach is that this montage is not used during the source reconstruction procedure. When possible, always consider the second option.Set the comment of the electrodes to define the groups. Right-click on the channel file > Edit channel file. Select the electrodes of group #1, right-click > Set channel comment. The electrodes with the same comment would be averaged together.
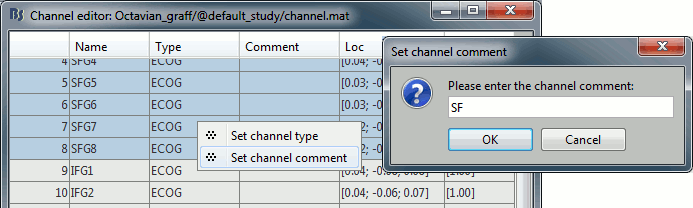
Signal viewer improvements
Flip +/- button: Exchange the direction of the Y axis, useful in clinical EEG
Clone figure: Right-click on the figure > Figure > Clone figure, or open twice the same data file from the popup menu in the database explorer. You can set different montages on the cloned figures, so you can look at the same information in different ways simultaneously.
Shortcut: Go to the next/previous half page (F4 / Shift+F4)
Setting manually time and amplitude resolutions: right-click > Figure > Set axis resolution
- Optimized scrolling time
Desktop management
Dealing with multiple figures can be annoying when you review lots of recordings.
The menu "Window layout options" can help you organize all the opened figures in an efficient way. There are four options for the automatic placement of the figures on the screen, and you can also save your own specific working environment with the new menu User setups.
See tutorial: Exploring the recordings
Colormaps
The management of the colormaps has been revamped to include non-symmetrical colormaps. You can now set independently the minimum and the maximum of the figure colorbar. A white marker has also been added to the colorbar when displaying source maps, to indicate the level of the amplitude threshold.
See tutorial: Exploring the recordings
December 2013
Filter files by name or comment
A new filter text box can help you select easily files in the Process1 and Process2 tabs.
See tutorial: Selecting files in the database.
New supported file formats
- EEG: Deltamed Coherence-Neurofile (.bin/.txt)
Electrophysiology: NeuroScope / Klusters (.eeg/.dat)
Other improvements
- Re-organization of the tabs: now they can be added and removed on demand
- Scouts: Remove a vertex from a scout by holding the shift key while clicking
- OpenMEEG: Help menu to make it easier to install or update
November 2013
New processes
Simulate > Auto-regressive and custom signals
Simulate > Simulate recordings from scouts (transform matrix files into source files)
Extract > Apply statistic threshold (export stat file with a specific static threshold)
Extract > Unconstrained to flat map (convert unconstrained sources to flat cortical maps)
File > Add tag
File > Select files with tag
Events > Detect analog triggers
- Sort event groups by name/by time (Record tab, menu Events)
October 2013
New tutorial: How to write your own plugins
Brainstorm offers a flexible plug-in structure. If you are interested in running your own code from the Brainstorm interface and benefit from the powerful database and visualization systems, the best option is probably for you to create your own process functions.
You would be able to exchange code easily with your collaborators and the methods you develop could immediately reach thousands of users. Once your functions are stable, we can integrate them in the main Brainstorm distribution and maintain the code for you to ensure it stays compatible with the future releases of the software.
Tutorial: How to write your own process
September 2013
Find Brainstorm users next to you
Brainstorm and SPM
It is now very easy to pre-process your recordings and estimate sources in Brainstorm, and do you statistics in SPM8 and SPM12. This powerful combination is illustrated in the following tutorials:
Export volume source maps to SPM8
Export surface source maps to SPM8/SPM12.
August 2013
Using FreeSurfer for the subject coregistration
The FreeSurfer software registers all the subjects to the FSAverage atlas using a spherical representation of the cortex. Each hemisphere of the subject's brain is inflated to a sphere, which is then deformed to match the curvature map of the equivalent sphere in the FSAverage subject.
In the case of a group study, these results are now used by default in Brainstorm to register all the subjects on the same default anatomy. For more information, read the group analysis tutorial.
July 2013
FreeSurfer subcortical atlas and cortical maps
Many news on the FreeSurfer import side. Now Brainstorm imports the subcortical atlas aseg.mgz as a set of surfaces. The cortical thickness results can also be saved in the database and explored easily. More information on the FreeSurfer tutorial page.
New anatomy templates
New models are available to replace the previous Colin27 default anatomy in your Brainstorm protocols. Right-click on (Default anatomy) > Use template. If a package is not available on your system, it will be downloaded from the Brainstorm website and saved in $HOME/.brainstorm/templates. The options available in this menu are:
Colin27: Average of 27 scans of the same head, processed with FreeSurfer 5.3: more information
Colin27_2012: Previous version of the default anatomy distributed with Brainstorm
ICBM152: Non-linear average of 152 subjects, processed with FreeSurfer 5.3: more information
FSAverage: Average of 40 subjects using a spherical averaging described in (Fischl et al. 1999)
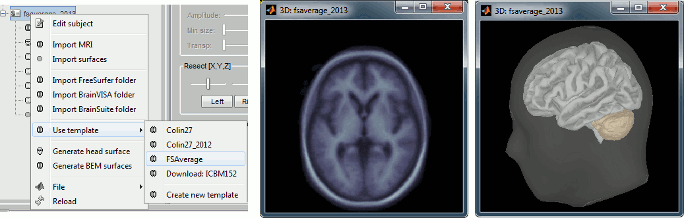
MRI segmentation: CIVET
The results of the CIVET anatomical segmentation pipeline, developed at the MNI, can be automatically imported in Brainstorm (surfaces and cortical thickness maps). Right-click on a subject > Import anatomy folder > Select "CIVET folder".
More information: CIVET website and CIVET tutorial.
Join the Brainstorm user community
A new section of the website dedicated to usage statistics and exchange between users: User community.
Brainstorm movie studio
A new fantastic feature to create movies: the "Snapshop > Movie (time): All figures" that you can find on all the 2D and 3D figures. Instead of capturing one figure only, it captures them all and create a movie showing what you see on the screen. Arrange your figures the way you want and create a movie of all your workspace at once.
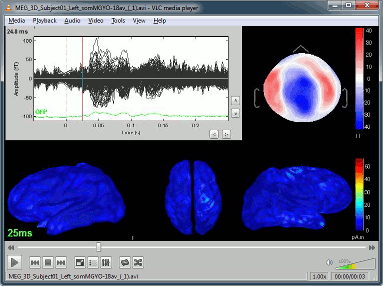
May 2013
MRI segmentation: BrainSuite support
The results of the BrainSuite anatomical segmentation can be automatically imported in Brainstorm (surfaces and atlases). See: BrainSuite website and BrainSuite tutorial.
New processes
Standardize > Concatenate recordings
Test>StudentT-test: Now available on time-frequency maps
Standardize>Uniform row names: Re-order the signals in a set of files to make them compatible.
Average>Average rows: Average the different signals in a time-frequency file.
Pre-process>Band-pass filter: Now available on source maps
Standardize>Event related perturbation
Events > Add time offset
April 2013
Event markers in the imported recordings
All the time markers available in the continuous recordings are now preserved when importing epochs in the database. It is also possible to add new markers to segment of recordings imported in the database, and to generate new epochs based on them.
A new option of the averaging process allows all the events from all the individual epochs to be combined. It can be a quick way to check that an event of interest (for instance the eye blinks) is not time-locked to the stimulus.
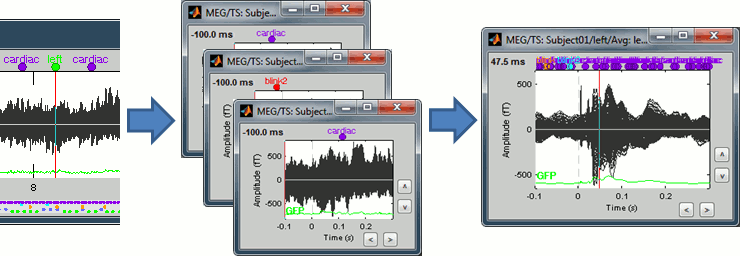
Dynamic Z-score
The Z-score operation, ie. the normalization of the signals by the level of noise calculated over a baseline, now exists in two forms. The process previously called "Compute Z-score" is now available as "Standardize > Z-score (static)".
The new process "Standardize > Z-score (dynamic)" produces the exact same results in terms of display, but calculated in an optimized way. Instead of re-writing the full file with the normalized values, we save only the mean and the standard deviation over the baseline for each signal, and apply the normalization on-the-fly when the file is loaded in memory.
This has almost no impact for the sensor data, but is a great improvement for the source files for which only the inversion kernel is saved. The static Z-score process would rebuild and save the full source matrix [Nsources x Ntime], while the dynamic version can keep the compact formulation of the linear inverse operator [Nsources x Nsensors]. The resulting files are much smaller and faster to open and review.
March 2013
Phase-amplitude coupling
The process "Frequency > Phase-amplitude coupling" offers a new way to explore the interactions between different frequencies. Phase-amplitude coupling (PAC) refers to the modulation of the amplitude of high-frequency oscillations by the phase of low-frequency oscillations. This process identifies in any type of signal the couple of low/high frequencies that maximizes the PAC metric introduced in [Canolty, 2006]:
Canolty RT, Edwards E, Dalal SS, Soltani M, Nagarajan SS, Kirsch HE, Berger MS, Barbaro NM, Knight RT,
"High gamma power is phase-locked to theta oscillations in human neocortex", Science, 313(5793):1626-8.
The process "Frequency > Simulate PAC signals" proposes a model for generating signals with a strong phase-amplitude coupling, for simulation purposes.
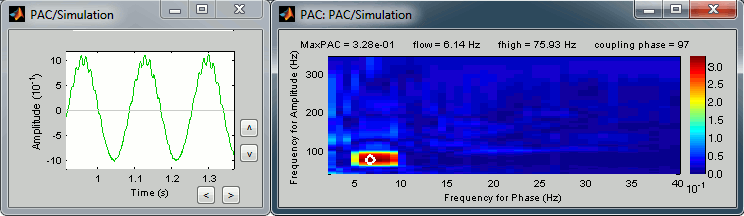
Multiple MRI volumes
Two or more MRI volumes can be imported for the same subject, if they have the same size and are properly co-registered. Only the first volume is used as a reference for placing the fiducial points, the additional ones are there for visualization purpose only. This feature can be useful for grouping different sequences (T1/T2), pre-op and post-op scans, regions of interest from fMRI results, or volume-based anatomical atlases.
Process: Run Matlab command
The process "Pre-process > Run Matlab command" is very simple but very powerful. It loads the files in input and run the signals through a piece of Matlab code that you can edit freely. It can extend a lot the flexibility of the Brainstorm pipeline manager, providing an easy access to any Matlab function or script.
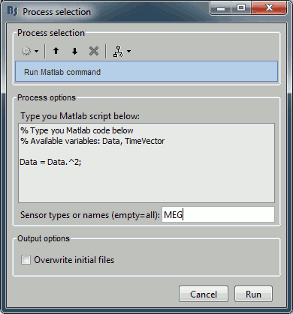
CTF/Neuromag dipole estimation
Dipoles localizations from the CTF DipoleFit and Neuromag Xfit software can be imported in the Brainstorm database and displayed on the same figures as cortical maps. See tutorial: Import and visualize dipoles.
February 2013
Intra-cranial EEG
The last version of the OpenMEEG software includes the ability to calculate forward models for cortical grids and intra-cortical electrodes. We integrated these changes to allow the calculation of sources for these two types of implanted electrodes. To be available in the forward and inverse model options, you need to change manually the type of the channels to respectively "SEEG" and "ECOG" (right-click on the channel file > Edit channel file). More tools are coming to visualize the amplitudes and select the position of the electrodes in the MRI viewer. If you are using these features, please cite both Brainstorm and OpenMEEG in your publications (see How to cite Brainstorm and OpenMEEG BEM head model).
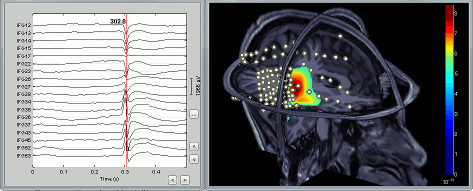
Yokogawa/KIT MEG recordings
The Yokogawa/KIT MEG recordings are now supported. They can be visualized, processed and epoched easily, such as any other file format. The reading function is based on the Yokogawa MEG Reader toolbox in Matlab, version 1.4. Supported extensions: .sqd, .con, .mrk, .ave. Some features will be improved shortly, such as the reading of the channel names, the events and the MEG/MRI registration, but the first prototype is fully working.
The events have to be processed manually: link the continuous file to your subject, then run the process "Events > read from channel" on the "Link to raw file". Select the options TTL or RTTL, and type the names of the channels to use as stim channels (example: " null045, null046").

Figure layouts
We are adding some more flexibility to the way the figures are automatically arranged on the screen. You can explore the different options available in the Layout menu. More features to come, including the ability to save custom layouts for the different types of figures.
Save surface snapshot
You can now create a new surface from the one that is displayed in a 3D figure. The exported surface would include all the current modifications: smoothness and resections. Right click on the figure > Snapshot > Save surface.
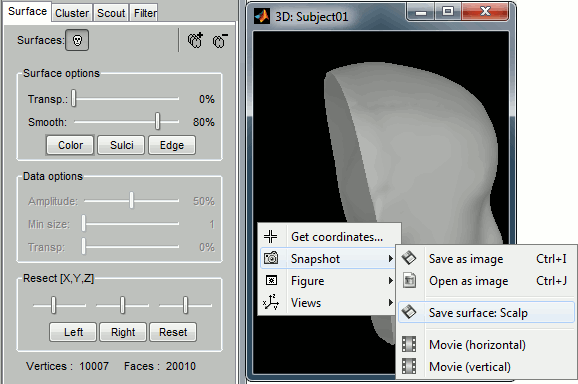
January 2013
Documentation
Many new step-by-step tutorials are now available, covering the processing and epoching of continuous files, the artifact rejection and the graphical scripting:
Report viewer: reload a pipeline
A report is generated each time an operation is executed from the Process1 or Process2 tabs, and automatically saved in the user folder ($HOME/.brainstorm/reports). You can browse through all the previous reports for the current protocol using the Report Viewer (File > Report viewer).
The reports can be reloaded easily: the menu "File > Reload last pipeline" selects again all the files that were selected previously, and opens the Pipeline Editor window and selects all the previous processes. The new button "Reload pipeline" in the toolbar of the Report Viewer allows you to reload any of the previously executed operations.
Processing continuous FIF files
Some important pre-processing operations are now available directly on continuous recordings in FIF format (usually Elekta-Neuromag MEG/EEG recordings). The new available processes include: the band-pass filter, the sinusoid removal (notch filter) and the baseline correction.
To process a continuous FIF file: link it to the Brainstorm database (right-click on the subject > Review RAW file), then drag and drop the "Link to raw file" in the Process1 box. Select a process in the Pre-process category. In output, it would create a new native continuous FIF file in the same folder, and also link it into the Brainstorm database. This feature is also available for CTF recordings, and is one of the key elements of our recommended pre-processing pipeline, as illustrated in the tutorial ?Detect and remove artifacts.
Copy of the noise covariance
As explained in the tutorial ?Noise covariance, the best way to estimate the sensors noise for MEG source reconstruction is to use empty room measurements. This implies that you have to import the noise measurements, calculate the noise covariance matrix, and copy it to the other acquisition runs or subjects.
You can use the popup menus "Copy to other conditions" or "Copy to other subjects", directly copy the file to the target condition using File>Copy/Paste, or use the equivalent keyboard shortcuts Ctrl+C / Ctrl+V. All three operations would automatically re-arrange the noise covariance matrix in case the order/number of channels is different for the noise and the target recordings.
MRI registration: joining interactive and automatic methods
The co-registration of the MEG/Polhemus positions with the anatomy of the subject (MRI+surfaces) is not an easy task. We offer three approaches that should be combined to get the best possible results: a first approximation based on three fiducial points (NAS/LPA/RPA), the automatic fit of the Polhemus head shape on the head surface from the MRI, and an rich interface for refining the results manually. The problem of the automatic method is its strong dependance to the initial conditions.
The new button "Refine using head points" in the edition window allows you to fix iteratively the registration: change the position manually, the run the automatic fit again. This solution can help you getting better registration results while we are working on an improved version of the automatic registration that would be less sensitive to the initial alignment.
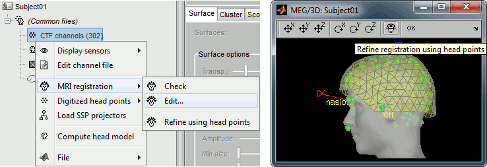
Subdivide scouts
The FreeSurfer atlases are now automatically imported in Brainstorm. They provide a good starting point for subdividing the cortex surface into anatomical regions, but the corresponding scouts can be too large for many functional applications. Averaging the time series from 600 different sources is likely to blur too much our data, we might lose the effects of interest. We are working subdividing efficiently the regions of these atlases. The first functions developed in this direction are the menus Subdivide atlas and Subdivide selected scouts in the Scout tab. They both perform a clustering of the existing scouts based on anatomical information.
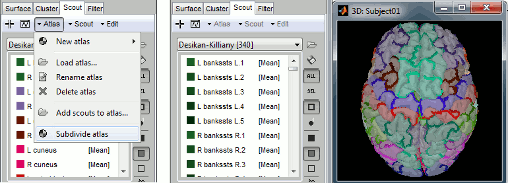
December 2012
FreeSurfer and BrainVISA
Improvements were made for handling FreeSurfer and BrainVISA segmentations. Both software packages generate a tree of folders that contain all the intermediate steps of the reconstruction, the top folder being the subject name or id. It is now possible to import the entire segmentation folders with just one click. Additionally, new menus are available to generate head surfaces from the MRI.
More details: Freesurfer and BrainVISA.
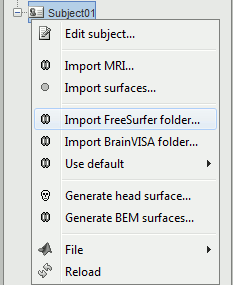
Importing MRI masks
By "MRI masks", we refer to MRI volumes that contain either only 0 and 1 (binary mask) or integer values (labelled atlas). Both type of MRI masks can be imported directly:
as surfaces: Right-click on subject > Import surfaces > Select the file type "Volume mask (all MRI files)". The tesselation is automatically generated by Brainstorm. This works well only in the case of convex envelopes, it is not recommended for complex objects
as scouts: In the Scout tab, menu Atlas > Load atlas > Select the file type "Volume mask (all MRI files)". For each label in the volume, a scout is created, containing all the vertices that are in this region.
New file types supported
- GIfTI .gii surface files, used by default in BrainVISA
- Curry event files (.cef)
November 2012
Updated topography plots
The rendering of the 2D topography plots is slightly improved, with a new option to set the number of contour lines.
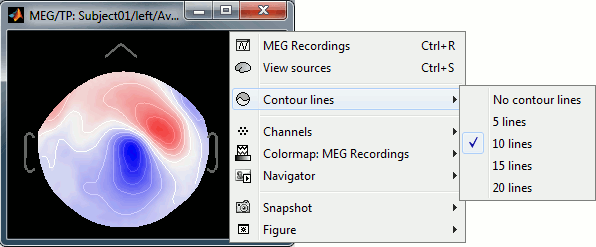
New processes
Standardize > Uniform epoch time: Sets the same time vector for all the input files
Pre-process > Scale values: Multiply the values of one or several sensors by a fixed factor
Average > Average frequency bands: Average values across frequency bands
Extract > Measure from complex values: Apply a function to complex values (typically Morlet or Hilbert coefficients)
Miscelanous
- Processing of time-frequency decompositions of source files is now supported (including for the projection on a default anatomy)
- Copy-paste operations are extended to most of the files available in the database explorer
- All the keyboard shortcuts have been adapted for Mac systems
October 2012
New interface for the scouts
The scouts interface has been redesigned to integrated the logic of atlases, and the classification of each scout in an anatomical region. You are welcome to visit the following tutorials for more information:
?Tutorials/TutScouts
?Tutorials/LabelFreeSurfer
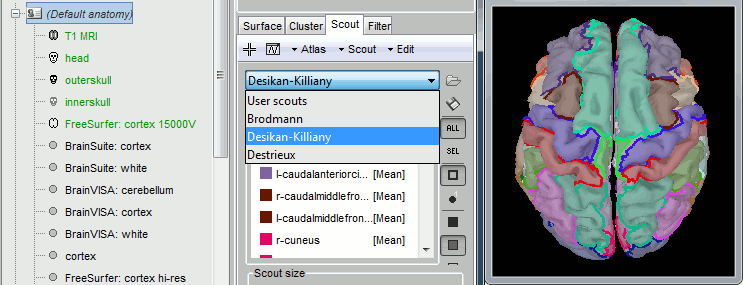
Project sources to an atlas
One of the main problems when dealing with source time series is the size of files. A new process lets you reduce the dimension of your sources by keeping only one value for each scout instead of one value for each vertex: Source > Downsample to atlas. It works well in the case of a set of scouts covering the entire cortex surface, either with a FreeSurfer atlas, or with a random clustering of the surface (menu Atlas>Surface clustering in the Scout tab).
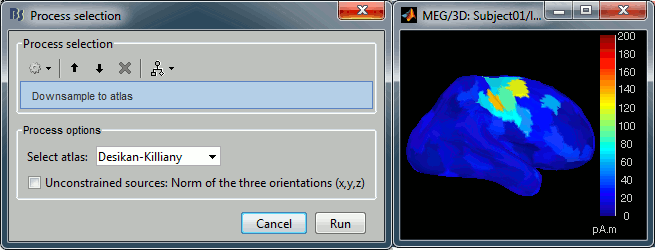
September 2012
Support for Biosemi BDF files
Brainstorm can now import and review native BDF/BDF+ files from Biosemi.
Support for MANSCAN files
Brainstorm can now import and review native MANSCAN EEG files (SAM Technology, Microamp amplifiers): .mbi/.mb2
August 2012
Drive your Polhemus from Brainstorm
For accurate source localization in MEG and EEG with registration to anatomical MRI image volume and efficient head localization in MEG, a 3D digitization solution such as the Polhemus-Fastrak is required. This new, simple Brainstorm feature will help you save time and is efficient for error-checking while digitizing sensor positions and the subject's head shape. See the Digitize tutorial for more information.
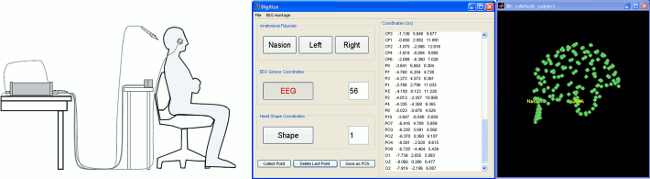
June 2012
Graphical batching interface: final version
All the features available from the Brainstorm interface are now also available as "processes", including the import of the data, the artifact cleaning and the source estimation. A full analysis pipeline, starting from the raw continuous file, can be edited graphically and saved as a Matlab script, easy to read an edit. These scripts can now run on distant servers without any user interaction.
The status of all the computations and the information messages that are generated during the execution of an batch script are now saved in detailed reports that can be reviewed as HTML pages, including screen captures of the data for quality control.
See the two tutorials: ?Processes and Automated processing.
April 2012
Matlab 2012a
The compiled version of Brainstorm is now based on Matlab 2012a. It provides more stability and fixes some issues with the Signal Processing Toolbox. However, if you were previously running the stand-alone version of Brainstorm, you may have to install a newer version of the Matlab Compiler Runtime. See the Installation page.
New processes
Events > Combine stim/response: Create new categories of events to group the markers by pairs (stimulus / response)
Standardize > Z-score with two sets of file in input:
Compute the variance s and average m of the baseline from files A,
Center on m and divide by s all the files B
March 2012
Frequency analysis
The interface for computing and displaying spectrum and time-frequency information has been largely improved. Two new types of windows are available, allowing the display of the time-frequency matrices as [time x amplitude] or [frequency x amplitude] graphs, for one given frequency or time. Four new processes are available, in the Frequency menu:
Fast Fourier Transform (FFT)
Power spectrum density (Welch)
Hilbert transform
Time-frequency (Morlet wavelets)
All these processes can run on any type of time series: recordings, sources, scouts.They all generate complex values, on which we need to apply a function to extract values that can be represented easily. This function can be changed easily after the computation, as a display parameter:
Power: Most common measure to represent the power of a signal for a given frequency bin
Magnitude: Energy of the signal for a frequency bin = sqrt(power)
Log: Convert to decibels (dB) = 10 * log10(power)
Phase: Extract the phase information for a given frequency bin = the angle of the complex value
Improvements made to the continuous viewer
- User-defined keyboard shortcuts to create events (keys 1-9)
- New buttons and keyboard shortcuts to explore the time series
- Bug fix: Automatic positioning of the windows for MacOSX and Linux is now working
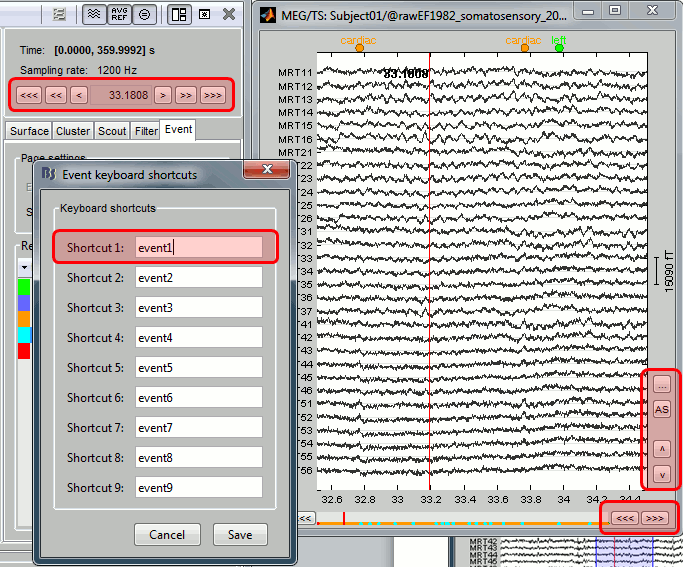
February 2012
Artifact removal with SSP
The SSP projectors are now saved in a more sophisticated way, allowing a dynamic selection of the projectors to apply on the recordings.
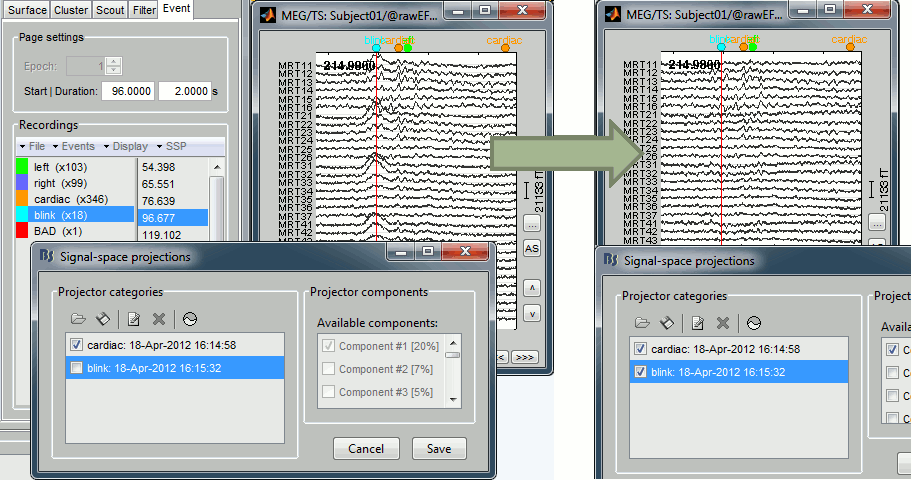
Co-registration of MEG runs
One of the problems related with the MEG analysis is that the subject's head can move with respect with the sensors during the acquisition. Hence, for the same subject, the position of a given sensor is different for two different runs or two different days of acquisition. We added a process (Standardize > Co-register MEG runs) to compensate for these head movements between acquisition runs, and register all the runs on one fixed head position. For now this process is limited to one head position per run, but we will soon work on using the continuous head localization information to compensate for the head movements at each time point.
Another issue with the data acquisition is the sensors selection: depending on the acquisition system or the day of acquisition, one may end up having lists of channels that do not match across runs or subjects. For instance: there were more auxiliary channels recorded for a few files, or different EEG amplifiers listed the electrodes in a different order. In these cases, it is very difficult to average or compare the information at the sensor level. The new process "Standardize > Uniform list of channels" automatically fixes all these issues by matching the sensors by name in the different files, and makes them compatible in terms on channel names and indices.
January 2012
Import FreeSurfer cortical parcellation
FreeSurfer is one of the many great academic software applications available to perform MRI image analysis. Brainstorm users can now use FreeSurfer to generate surface envelopes of head tissues to generate a model of MEG or EEG sources. Brainstorm now features the possibility to import the labels for regions of the cortex generated by FreeSurfer's analysis, which correspond to the regions of the Desikan-Killiany and Destrieux cortical parcellation atlases. The corresponding surface areas can subsequently be treated as scouts in Brainstorm, and therefore be used as anatomical ROIs to guide the exploration of MEG/EEG source maps.
See tutorial: Use FreeSurfer cortical parcellation
Import Nifti-compressed MRI images
We have added the possibility to import compressed Nifti MR images directly into Brainstorm's MRI Viewer and export MR volumes as Nifti.
Improved user interface for time series visualization
We have added new, convenient features to navigate through trials, time samples and files in your Brainstorm database. Buttons have been added to the main Brainstorm panel and also into the figure window that display time series: navigate through complex data structures with just a click! You can also adjust the amplitude gain using buttons now, or readily set the scale of time-series displays. As always, feedback is always welcome through our forum.
December 2011
Brainstorm on Facebook!
'Like Us' on Facebook to stay in touch: 
November 2011
New tools for artifact detection and correction
1) Specific channels can be scanned for automatic detection of ocular and cardiac artifacts from continuous recordings; this feature works best with dedicated control channels (i.e. ECG and EOG),
2) Artifact modeling and attenuation using a signal space projection (SSP) approach.
Stand-alone distribution for Linux and MacOSX
We introduced a new packaging logic for the distribution of Brainstorm executable (i.e., which does require the user to run a Matlab license). The entire set of application files (Matlab scripts and compiled executable) are now integrated in the same unique archive.
The executable is now platform (OS) independent as it has been reduced to a single .jar file created with Matlab's JavaBuilder, that runs on any platform supporting Matlab. To run the stand-alone version of Brainstorm, you just need to install the free, Matlab Compiler Runtime (version 7.16) foryour operating system.
Read more on the Installation page.
Brainstorm course in Montreal
Part of the Brainstorm development team has moved to theMcConnell Brain Imaging Center at McGill University's Montreal Neurological Institute, in Montreal, Canada. At this occasion, a full-day training session with 70 participants from across Canada has been organized for new users, back-to-back with the first MEG training workshop of the Canada MEG Consortium (see pictures here).
For more training opportunities, visit our Training pages.
May 2011
Brainstorm reference article
Publication of a special issue of the journal Computational Intelligence and Neuroscience:Academic Software Applications for Electromagnetic Brain Mapping Using MEG and EEG, co-edited bySylvain Baillet, Karl Friston & Robert Oostenveld. We encourage Brainstorm users to cite this reference in their publications featuring analyses performed using Brainstorm (see How to cite Brainstorm).
Tadel F, Baillet S, Mosher JC, Pantazis D, Leahy RM (2011) Brainstorm: A User-Friendly Application for MEG/EEG Analysis, Computational Intelligence and Neuroscience, vol. 2011, Article ID 879716, 13 pages, 2011. doi:10.1155/2011/879716,pdf]
Improvements in navigating the database
The file manager has been extended to support drag-and-drop and copy-paste operations. Users can now move or copy easily files from a condition or subject entry to another using the mouse, keyboard shortcuts [CTRL+C (copy), CTRL+X (cut) and CTRL+V (paste)], or the File section from the contextual menus from the GUI.
No MRI for your participants?
No worries: Brainstorm can adjust a template anatomy to individual scalp points (MEG) / electrode locations (EEG).
It is now possible to generate pseudo-individual anatomies from a template, for participants for whom the anatomical MRI volume is not available. This operation is possible if digitized head (scalp) points were acquired using a tracking system (e.g., Polhemus Isotrak), which is commonplace in most MEG labs. The default anatomy (volume and surface envelopes) distributed with Brainstorm (MNI/Colin27) can be transformed (warped) to match the subject's scalp points. This is a great cost-saving alternative when individual anatomy is not accessible to investigators (not recommended for accurate, local mapping of the loci of activity though).
See tutorial: Warping default anatomy.
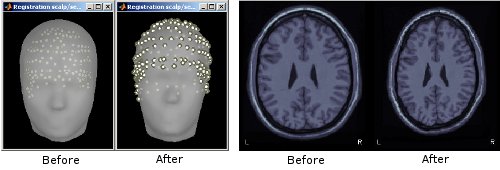
Refine MRI/MEG registration using digitized scalp points
When importing MEG recordings, the registration between the MRI volume and the MEG was formerly based only on three fiducial points only (nasion, left ear, right ear). This approach is known be potentially of poor accuracy. If multiple (>60, typically) scalp points were digitized, the transformation between the MEG's and the MRI's referential can be refined by minimizing the average distance between the scalp surface obtained from MRI and the actual digitized head points.
See tutorial: Review raw recordings.
April 2011
Realistic boundary-element-modeling (BEM) head models for EEG
A new feature has been made available for a more accurate computation of EEG forward models with realistic geometry (strongly recommended). OpenMEEG generates forward models using a symmetric boundary element method that was developed by the INRIA team ATHENA. It considers three realistic layers (scalp, inner skull, outer skull), which can all be generated from the MRI volume using Brainstorm.
See tutorial: BEM head model.

Source estimation in full brain volume
Source models computed in Brainstorm were until now restricted to the cortical surface. To extend the range of analyses made possible with Brainstorm, and to compare Brainstorm's source estimates with these obtained using other software, source models based on dipole grids sampling the entire brain volume can be obtained and reviewed, using the same efficient tools featured in Brainstorm. See tutorial: Volume source estimation.
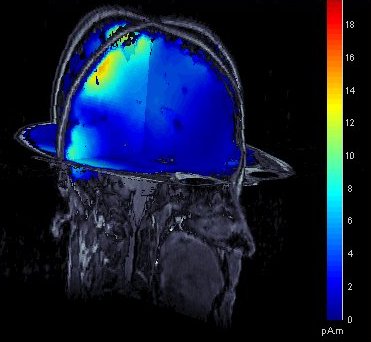
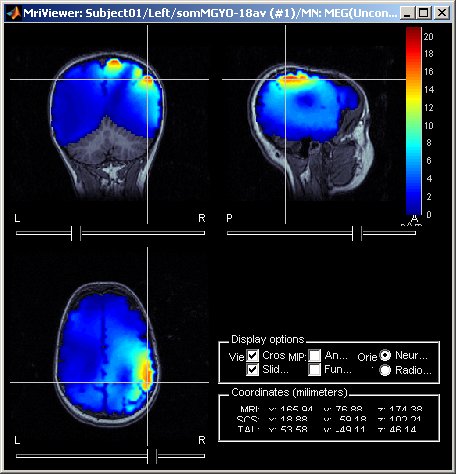
March 2011
Combined MEG/EEG source reconstruction
The approach to combine concurrent MEG and EEG recordings to obtain a joint source model has been substantially improved.
Display of source maps in MRI volume
Added multiple options to review and visualize source maps using contact sheets and volume smoothing of activity maps. 1) source maps are spatially smoothed after interpolation in the MRI voxels. The size of the smoothing kernel can be configured: right-click on the figure > MRI display; 2) the resulting volumes can be used to create contact sheets along all viewing directions, either in time or across the volume.
See tutorial: ?Source estimation.
December 2010
Statistical inference: correction for multiple comparisons
When displaying the outcome of a t-test, there is now a new tab "stat" that is shown in the main window, which allows to adjust dynamically the type of correction applied to multiple comparisons and the p-value of the test.
Graphical batching interface
A brand new version of the "Process" tab is available. Processing pipelines can now be elaborated in a convivial way, in just a few clicks! You can also save and export your pipeline as Matlab scripts. The processes available are now written as plug-ins: you can contribute your own process and it will automatically list to your menu of available tasks that can be performed on data.
See tutorial: ?Processes.
September 2010
Continuous/RAW file viewer and event editor
Brainstorm can now be used to visualize and process full length continuous/raw files in native format. See tutorial: ?Review raw recordings and edit markers.
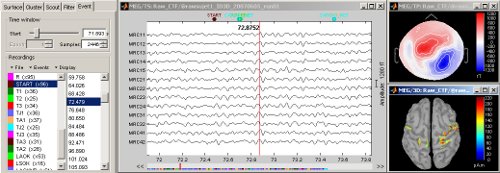
Detection of bad trials / bad channels
New process available to detect bad trials or bad trials based on peak-to-peak values. The peak-to-peak threshold can be set by channel type.
Estimation of the noise covariance from continuous files
The interface that computes the noise covariance matrix can be called on continuous recordings. Right-click on link to raw file > Noise covariance.
Import/export protocols and subjects
New menus to help users exchange data easily. A full protocol can be exported as a single zip file (menu File > Export protocol), and then imported in another database (menu File > Load protocol > Load from .zip file). Same thing for a single subject: right-click on the subject > File > Export subject. The subject .zip file is actually a full protocol zip file with only the exported subject, hence it should be loaded as a protocol.
June 2010
Time-frequency
Computation and visualization of time-frequency decompositions of MEG/EEG and source signals using Morlet wavelets. See tutorial: ?Time-frequency.
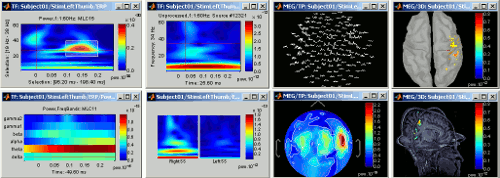
Brodmann and Tzourio-Mazoyer atlases
Two lists of scouts have been created for the default anatomy (MNI/Colin27), inspired from the atlases of Tzourio-Mazoyer and Brodmann. To access these files: display the cortex surface of the default anatomy, then in the "Scout" tab click on the "Load scouts" button, on the right of the scouts list.
Recordings visualization: 2D Layout
The 2D Layout display for recordings has been re-writen to include many tools: sensors selection, interactive gain change, zoom in time and space... See tutorial: ?Exploring the recordings.
March 2010
Support for Xfit dipoles files
Dipoles localizations from the Neuromag Xfit software can be imported in Brainstorm database and displayed on the same figures as Brainstorm cortical maps. See tutorial: ?Import and visualize dipoles.
Minimum norm solution
Integration of a new minimum norm solution, with more and better tuned parameters. This algorithm is now similar to the one implemented in MNE software.
Januray 2010
Project sources on default anatomy
Any source file estimated for a given surface can be re-projected on another surface. This allows to project sources of individual subjects on the default anatomy to compare subjects. See tutorial: Project sources on default anatomy.
Display channels in columns
The time series can be displayed in columns. The displayed sensors can be edited with an interface similar to the selection editor in MNE software.
December 2009
Automatic updates
Brainstorm is now self-updating. It tests at each startup if your version is older than a month; if so, it downloads a new version. The software can also be updated manually very easily (menu Help > Update Brainstorm).